
Int J App Pharm, Vol 17, Issue 4, 2025, 10-18Review Article
BIBLIOMETRIC ANALYSIS OF DRUG RESISTANCE BIOMARKERS IN BREAST CANCER (2020-2025)
EVA NURSOLEHA, WAHYU UTAMI*
Faculty of Pharmacy, Universitas Muhammadiyah Surakarta, Sukoharjo-57169, Indonesia
*Corresponding author: Wahyu Utami; *Email: wahyu.utami@ums.ac.id
Received: 17 Mar 2025, Revised and Accepted: 29 May 2025
ABSTRACT
Objective: Biomarkers have become critical tools for predicting treatment response and guiding the development of strategies to overcome resistance. This study aims to examine research trends on drug resistance biomarkers in breast cancer from 2020 to 2025, with a focus on identifying key themes, influential authors, contributing countries, and journals contributing to this topic, thus providing insight into ongoing scientific developments and research directions. The bibliometric analysis was conducted using the Scopus database. The search employed the keywords “drug resistance biomarkers” AND “breast cancer,” filtered for open-access articles published between 2020 and 2025 in English. A total of 252 documents were selected based on inclusion and exclusion criteria. Data analysis and visualization were carried out using the Visualization of Similarities viewer (VOSviewer) version 1.6.20, focusing on publication patterns, keyword co-occurrences, and collaboration networks. Out of 1,400 initial records, 252 publications met the inclusion criteria. The year 2022 recorded the highest number of publications. China and the United States emerged as the leading contributors to research on this field. Major contributing journals included Cancers, Frontiers in Oncology and Breast Cancer Research. Prominent research topics included circular RNAs, microRNAs, and protein. Several biomolecules, such as circUBAP2, miR-300, ASF1B, and exosomal Del-1 protein, were identified as key mediators of resistance to chemotherapeutic agents like cisplatin, adriamycin, and paclitaxel. To conclude, this study highlights the potential of specific biomolecules as predictive biomarkers and therapeutic targets for drug-resistant breast cancer. Future research should integrate multi-omics approaches and international collaboration to enhance the clinical translation of these findings and promote personalized treatment strategies.
Keywords: Breast cancer, Drug resistance, Biomarkers, Bibliometric analysis, Chemotherapy, Scopus
© 2025 The Authors. Published by Innovare Academic Sciences Pvt Ltd. This is an open access article under the CC BY license (https://creativecommons.org/licenses/by/4.0/)
DOI: https://dx.doi.org/10.22159/ijap.2025v17i4.54227 Journal homepage: https://innovareacademics.in/journals/index.php/ijap
INTRODUCTION
Breast cancer is the most common type of cancer in women worldwide. Based on data from the Global Cancer Observatory (GLOBOCAN) 2022, breast cancer ranks first as the cancer with the highest number of new cases in women, with 2,296,840 new cases diagnosed, accounting for 23.8% of all cancer cases in women. In addition, breast cancer is also the fourth leading cause of death from cancer, with a death toll of 666,103 cases. The prevalence of breast cancer in the last five years has reached more than 8.1 million cases, indicating that this cancer has a significant impact on global health [1].
Cancer treatment can be carried out through surgery, chemotherapy, or radiotherapy. Recommended treatments for breast cancer include surgery, such as lumpectomy and mastectomy, which aim to remove tumors and surrounding tissue to prevent cancer spread [2, 3]. Chemotherapy is used to kill or inhibit cancer cell growth, either before surgery (neoadjuvant) to shrink tumors or after surgery (adjuvant) to prevent recurrence, primarily using platinum-based drugs (cisplatin) and taxanes (paclitaxel), and radiotherapy is also often administered post-surgery to eliminate any remaining cancer cells [4-6]. Despite treatment advances, drug resistance, particularly to platinum-based therapies, presents a significant clinical challenge. Approximately 50% of breast cancer patients develop resistance after several treatment cycles, substantially compromising their prognosis [7, 8]. This high resistance rate contributes to the poor survival rate (20-30%) observed in advanced stages [6]. Recent research by [9] has identified several molecular mechanisms underlying this resistance, including altered drug metabolism, enhanced Deoxyribonucleic Acid (DNA) repair capabilities, and dysregulation of apoptotic pathways.
Although research into genetic and epigenetic mechanisms of resistance has advanced, no reliable biomarkers have yet been validated in clinical practice to predict response to platinum-based chemotherapy. Accurate biomarkers are crucial for early identification of resistance, real-time monitoring, and personalization of therapy. They support treatment decisions, improve outcomes, and reduce unnecessary toxicity [10].
Bibliometric analysis represents a powerful methodological approach in fundamental research, providing a statistical framework for conducting systematic, transparent, and replicable reviews of scientific literature through quantitative measurements [11]. A large amount of scientific data was collected and analyzed using bibliometric software, such as Visualization of Similarities viewer (VOSviewer), and scientific databases such as Scopus [12]. Bibliometric analysis enables researchers to visualize and understand the complex landscape of drug resistance biomarker research by identifying emerging research clusters and their interconnections, highlighting the temporal evolution of key concepts and methodologies, and revealing knowledge gaps that require further investigation [13].
Bibliometric analysis is a quantitative approach that evaluates scientific publications to identify trends, collaborations, and research gaps. By mapping keywords, co-authorships, and citations, bibliometrics helps researchers understand how a field evolves. In drug resistance biomarker research, this method allows tracking of technological, molecular, and therapeutic developments across regions and institutions. Recent studies [10] emphasize the importance of integrated-omics approaches in biomarker discovery and resistance profiling. Additionally, advanced machine learning techniques have enhanced bibliometric analyses by identifying subtle patterns in research trajectories that traditional methods might miss [13].
This study reports the findings of a bibliometric analysis of scientific literature from the past five years on biomarkers associated with drug resistance in breast cancer. The objective is to highlight emerging trends and research dynamics with promising potential for future investigation and application. It also identifies key contributors, including authors, studies, countries, and institutions, along with other relevant data. This information can be used to design future research, build collaborative networks, and develop planning strategies to improve the treatment and understanding of cancer [14].
Methods
Research design
A bibliometric method was used to analyze scientific articles indexed in the Scopus database from 2020 to 2025. Scopus was selected for its wide coverage of high-impact journals and its compatibility with bibliometric software [15]. Compared to other databases such as Google Scholar, PubMed, and Web of Science, Scopus offers more comprehensive citation data, indexing over 25,000 peer-reviewed journals, and includes a broader range of disciplines beyond biomedical sciences [16, 17].
Additionally, Scopus facilitates easier export of bibliographic data in formats compatible with bibliometric analysis tools like VOSviewer. The analysis was conducted using VOSviewer version 1.6.20 visualization parameters, where each element (countries, keywords, authors) was represented as a bubble, with size indicating relative weight or frequency in the dataset [18, 19]. Other databases like Google Scholar, Web of Science, and PubMed were excluded due to limitations in export features or access. The study focused on original articles and reviews published in English and open access, as shown in the inclusion and exclusion criteria in table 1.
Search strategy and criteria
The search query used was: TITLE-ABS-KEY ("drug resistance" AND "biomarkers") AND TITLE-ABS-KEY ("breast cancer"*). Boolean operators such as AND were used to ensure that the search results contained all the specified keywords simultaneously, thus increasing the precision and relevance of the articles retrieved. The AND operator narrows the search by retrieving only those articles that include both terms, in this case, ensuring the focus remains on studies discussing both drug resistance and biomarkers specifically within the context of breast cancer. Filters included only open-access articles in English. Duplicate records were removed using the Mendeley Reference Manager to minimize limitations or potential biases in the bibliometric analysis. A schematic of the methods employed in this investigation is shown in fig. 1.
Table 1: Inclusion and exclusion criteria for document selection
| S. No. | Criteria | Inclusion | Exclusion |
| 1. | Publication Type | Original research articles, Review articles | Conference papers, Book chapters, Letters, Editorials, Note, Short survey, Retracted, Conference proceeding, and Erratum |
| 2. | Period | January 2020-February 2025 | Publications before 2020 |
| 3. | Language | English | Non-English publications |
| 4. | Access Type | Open access | Subscription-based access |
| 5. | Research Focus | Studies examining drug resistance biomarkers specifically in breast cancer | Studies not directly related to drug resistance or focused on other cancer types |
| 6. | Methodology | Gene expression analyses Ribonucleic Acid sequencing, microarray, Quantitative Reverse Transcription Polymerase Chain Reaction (qRT-PCR), gene silencing, protein inhibition, and molecular abundance studies about chemotherapy response. | Studies without clear methodological descriptions |
| 7. | Study Models | Human patients, animal models, and cellular models | Computer simulations without biological validation |
| 8. | Document Quality | Peer-reviewed publications | Preprints, non-peer-reviewed content |
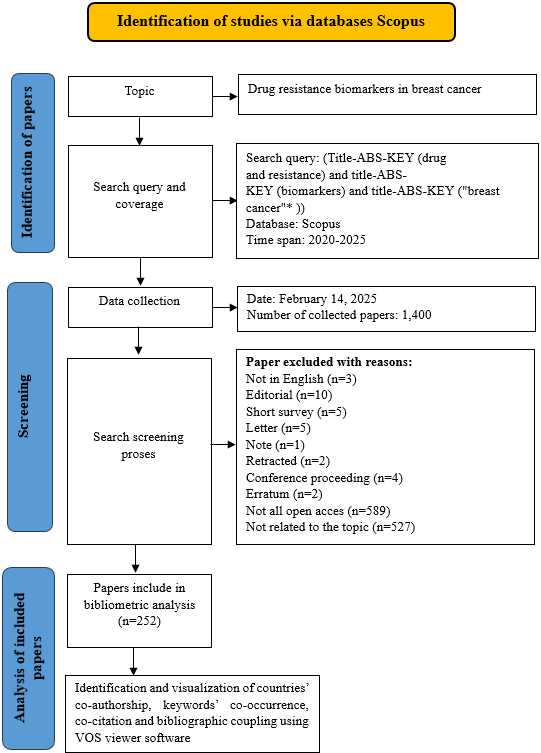
Fig. 1: PRISMA flow diagram of the exclusion and inclusion of literature. The overall workflow of the study includes two main stages: an initial Scopus search for papers produced between 2020 to 2025 related to resistance biomarkers in breast cancer, followed by a screening stage, and finally, bibliometric analysis in VOSviewer. Adapted from references [12, 20]
Data extraction and tools
Citation and bibliographical information, abstracts, and keywords were among the bibliographic data that were obtained for research publications and reviews. CSV files of the data were retrieved from Scopus and exported to Microsoft Excel version 2021. Afterward, data analysis and visualization were conducted using VOSviewer version 1.6.20 and Publish or Perish version 8. These tools were selected for their ability to visualize bibliometric networks and generate cluster-based maps.
Data analysis
All statistical analyses were conducted in two stages. The first stage involved descriptive analysis of bibliographic data, including articles, citations, authors, collaboration, as well as the most frequent journals and keywords. The second phase involved constructing a network to examine bibliographic coupling, co-citation patterns, collaboration, and co-occurrence [21]. Using the PICO model (Patient/Population, Intervention, Comparison, and Outcomes), our search targeted studies involving patients, animal models, and cellular models, as well as research evaluating drug resistance in breast cancer. The methods of interest included gene expression analyses (RNA sequencing, microarray, qRT-PCR), gene silencing, protein inhibition, and molecular abundance about chemotherapy response.
Only published papers that satisfied the inclusion criteria were analyzed. The criteria included original research articles, review articles, publications in English, and studies focused on drug resistance biomarkers in breast cancer. Fig. 1 outlines the process of data collection and study exclusion. Using a bibliometric tool [22] we examined co-authorship among countries, co-occurrence of keywords, and bibliographic coupling of authors, which were visualized as network maps. Each element (countries, keywords, and authors) was represented as a bubble, with its size indicating its relative weight, which is proportional [18, 19, 23]. The bubble’s color signifies the cluster to which the item belongs.
RESULTS AND DISCUSSION
Publication trends
Out of the 1,400 collected published papers on drug resistance biomarkers in breast cancer, only 252 met the inclusion criteria fig. 1 . These selected studies spanned 528 journals and were authored by 376 researchers. The earliest paper, published in 2000 by Amornmarn, R., Bui, Prempree, and Masood, was titled "Molecular Predictive Factors for Local Recurrence and Distant Metastasis of Breast Cancer After Lumpectomy with Postoperative Radiation Therapy" [24]. The most recent study, published in 2025 by Shi, Q., Yang, W., Ouyang, Y., Liu, Y., and Cai in Breast Cancer Research, explored "CXCR4 Promotes Tumor Stemness Maintenance and CDK4/6 Inhibitor Resistance in ER-Positive Breast Cancer" [25].
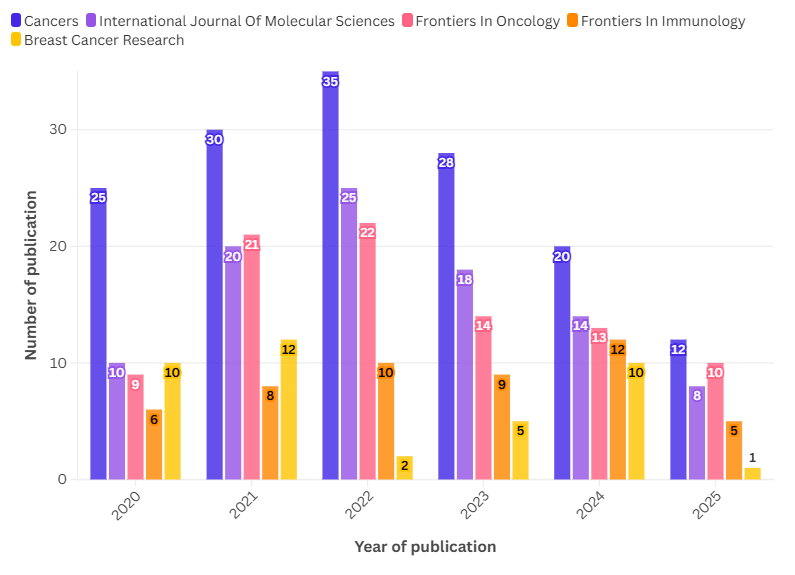
Fig. 2: Publication trends on the topic of drug resistance biomarkers and breast cancer
The number of publications on drug resistance biomarkers in breast cancer has fluctuated over the years, from 2020 to 2025 (fig. 2).. A notable increase in research interest was observed in 2022, when the highest number of publications was recorded, totaling 35 papers. This surge can be attributed to heightened global awareness and funding opportunities following the Coronavirus Disease 2019 (COVID-19) pandemic, which prompted increased collaborative research efforts and policy-driven scientific investments. Notably, 2022 coincided with international campaigns such as Breast Cancer Awareness Month initiatives, renewed commitments by the World Health Organization’s Global Breast Cancer Initiative (GBCI), and several large-scale funding calls from cancer research foundations targeting post-pandemic innovation in diagnostics and personalized medicine. These developments likely created a fertile environment for research on drug resistance biomarkers to flourish. After 2022, the number of publications began to decline. This drop may reflect a natural saturation point in specific research niches, a shift in scientific focus toward other subtypes of cancer or mechanisms of resistance, or the redirection of funding to more translational or clinical implementation projects. Additionally, the tapering of emergency pandemic-related funding and prioritization may have contributed to the observed decline. Across journals, Cancers (blue line) led in publication volume, peaking in 2022 before a notable reduction in subsequent years.
The International Journal of Molecular Sciences (purple line) also saw a significant rise in publications, peaking in 2022 with more than 25 documents. However, similar to Cancers, the number of publications declined in the following years. This trend indicates a concentrated research interest around 2022, followed by a gradual decline. Frontiers in Oncology (pink line) followed a similar trajectory, with a peak in 2022 at approximately 22 publications before experiencing a decline in 2023 and 2024. This trend reflects a growing but temporary interest in the topic within the oncology research community. In contrast, Frontiers in Immunology (orange line) maintained a relatively stable but lower number of publications across the years, suggesting a steady contribution of immunological perspectives to drug resistance research in breast cancer. Unlike the other journals, this journal did not exhibit sharp fluctuations but rather a consistent presence. Breast Cancer Research (yellow line) displayed a different pattern, with the number of publications declining sharply in 2022 before rising again in 2024. This unique trend could indicate shifting research priorities or variations in publication acceptance rates for this journal.
In summary, the publication trend peaked in 2022, likely due to a confluence of post-pandemic research momentum and global health initiatives focused on breast cancer. The subsequent decline across most journals suggests a redistribution of scientific focus, possibly toward translational applications or emerging oncological priorities. Despite this decline, the topic of drug resistance biomarkers in breast cancer continues to hold significant scientific and clinical relevance.
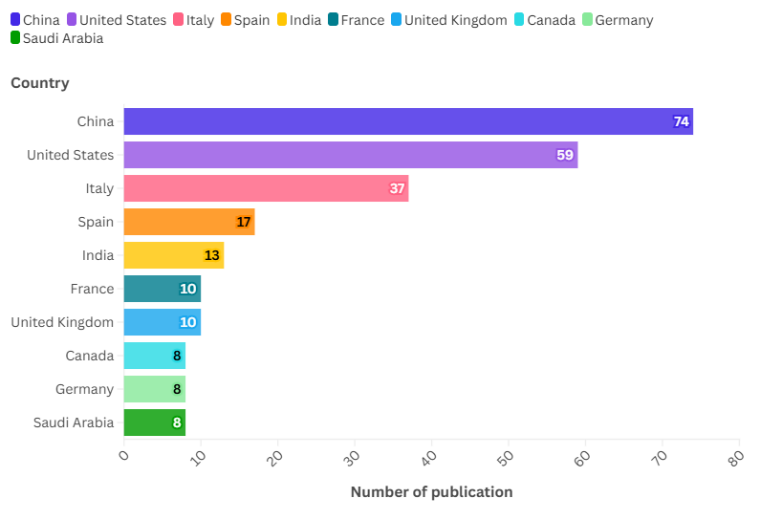
Fig. 3: The most productive countries on the topic of drug resistance biomarkers and breast cancer
Countries, institutions, and main collaboration networks
Fig. 3 highlights the most prolific and engaged countries in the study of drug resistance biomarkers in breast cancer, determined by the affiliated country of the corresponding author. The leading contributor is China, with (74 documents), followed by the United States (59 document), Italy (37 document), Spain (17 document), India (13 document), France (10 document), the United Kingdom (10 document), Canada (8 document), Germany (8 document) and Saudi Arabia (8 document). The dominance of China and the U. S. reflects their substantial investments in biomedical infrastructure, national cancer initiatives, and large-scale funding programs.
These countries benefit from strong governmental and institutional support, including high research budgets, well-established cancer research centers, and access to cutting-edge technology and data systems. China’s rise in scientific output has been driven by major national strategies such as the "Healthy China 2030" plan and extensive government funding through the National Natural Science Foundation of China (NSFC). Similarly, the U. S. has long been a leader in oncology research, with institutions like the National Cancer Institute (NCI) providing continuous support for breast cancer research and biomarker development. Their robust academic ecosystems and international collaborations have positioned both nations as hubs for innovative cancer studies.
However, the concentration of publications from these two countries also raises potential concerns regarding bias in global research contributions. One notable factor is funding inequality wealthier countries tend to dominate high-impact publications due to better access to resources, advanced laboratories, and publication networks. Language barriers may also influence visibility: non-English-speaking researchers, particularly from lower-income regions, may struggle to publish in leading English-language journals, thereby limiting the global diversity of perspectives and findings. These structural imbalances risk marginalizing contributions from underrepresented regions despite local relevance and clinical need.
This indicates that countries with well-established biomedical research infrastructures, substantial funding allocations, and international collaborations are at the forefront of scientific discoveries. The presence of leading research institutions, such as the Chinese Academy of Sciences, the National Cancer Institute (NCI) in the U. S., and major universities in the UK, further reinforces the productivity in this domain. Moreover, the increasing trend in publications from emerging economies suggests a growing global interest in the field [26].
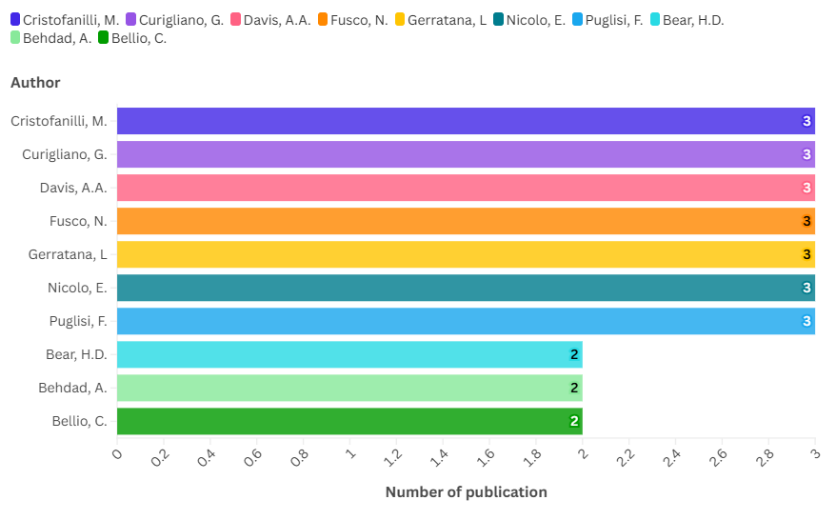
Fig. 4: The top 10 authors on the topic of drug resistance biomarkers and breast cancer
From an authorial perspective, the results reveal that the top 10 authors are responsible for a significant portion of the scientific contributions related to drug resistance biomarkers in breast cancer (fig. 4). Among them, Cristofanilli, M. (3 documents, 38 citations), Curigliano, G. (3 documents, 223 citations), Davis, A. A. (3 documents, 54 citations), Fusco, N. (3 documents, 150 citations), Gerratana, L. (3 documents, 38 citations), Nicolo, E. (3 documents, 30 citations), Puglisi, F. (3 documents, 38 citations), Bear, H. D. (2 documents, 219 citations), Behdad, A. (2 documents, 38 citations), and Bellio, C. (2 documents, 14 citations) have played a key role in advancing knowledge through their extensive publication records.
Table 2: The most productive affiliations on the topic of drug resistance biomarkers and breast cancer
| S. No. | Affiliations | Country | Number of publications | Citations | Brief of studies |
| 1. | Università degli Studi di Milano | Italy | 10 | 397 | Cancer research, drug resistance, biomarkers |
| 2. | Harvard Medical School | United States | 9 | 124 | Oncology, chemotherapy, targeted therapy |
| 3. | Istituto Europeo di Oncologia | Italy | 6 | 336 | Breast cancer, immunotherapy, tumor microenvironment |
| 4. | Dana-Farber Cancer Institute | United States | 5 | 105 | Chemotherapy, tumor immunology, and precision medicine |
| 5. | Sapienza Università di Roma | Italy | 5 | 117 | Drug resistance, molecular oncology, targeted therapy |
| 6. | Università degli Studi di Padova | Italy | 4 | 126 | Cancer metabolism, pharmacogenomics, tumor suppression |
| 7. | Inserm | France | 4 | 11 | Molecular oncology, epigenetics, and drug discovery |
| 8. | Nanjing Medical University | China | 4 | 28 | Cancer biomarkers, targeted therapy, and chemotherapy resistance |
| 9. | Zhengzhou University | China | 4 | 34 | Oncogenic pathways, drug resistance, and chemotherapy |
| 10. | Sun Yat-Sen University | China | 4 | 6 | Immunotherapy, tumor biology, cancer genomics |
Table 2 presents the most productive affiliations in this research area, demonstrating that major cancer research centers are involved. The most productive affiliation was Università degli Studi di Milano, with 10 papers. Among the ten affiliations on the list, Italy had 4 affiliations, the United States had 2 affiliations, China had 3 affiliations, and France had 1 affiliation. All of the countries with the most productive affiliations are located in Europe, North America, and Asia. The highest number of citations among the most productive affiliations was from Università degli Studi di Milano, with 397 citations. This indicates that the papers produced by Università degli Studi di Milano on this topic are more influential than those from other institutions. Their contributions span various aspects of drug resistance mechanisms, biomarker discovery, and therapeutic advancements. Through their strong collaborative networks, they facilitate the dissemination of research findings and foster interdisciplinary efforts, ultimately contributing to more effective breast cancer treatments.
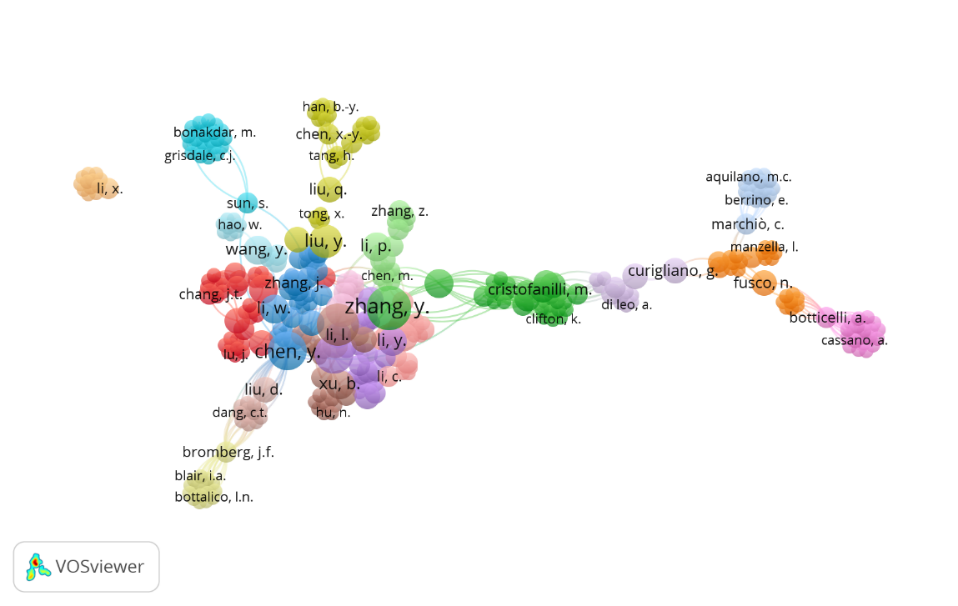
Fig. 5: The network visualization of bibliographic coupling of authors on the topic of drug resistance biomarkers and breast cancer
Fig. 5 visualizes the bibliographic coupling of authors, revealing clusters of interconnected researchers working on drug resistance biomarkers in breast cancer. The different colored clusters represent distinct research groups, with stronger connections indicating a higher degree of shared references. The presence of multiple clusters suggests that research in this field is being conducted by several independent yet interrelated groups. According to [27] bibliographic coupling analysis provides insights into the intellectual structure of a research domain, illustrating how different studies are linked based on shared references.
Notably, certain authors such as Zhang Y., Chen M., and Liu Y. appear as central figures, forming the largest and most interconnected clusters. Their strong bibliographic coupling suggests that their work serves as a foundation for multiple studies in this domain. The green cluster, including Cristofanilli M. and Clifton K., represents another influential group contributing to breast cancer research. Additionally, the rightmost portion of the network highlights authors such as Aquliano M. C. and Botticelli A., who form a separate but still connected research subgroup. This network analysis highlights the importance of collaboration in advancing cancer research. Researchers frequently co-author studies, share methodologies, and build upon each other’s findings.
The most relevant words
The study of drug resistance biomarkers in breast cancer has become a crucial area of research in oncology, particularly in efforts to improve treatment efficacy and clinical outcomes of drug resistance biomarkers and breast cancer. Bibliometric analysis indicates that research in this field remains highly specialized, with over 75% of authors contributing only a single paper, while the 20 most productive authors account for 63% of the analyzed publications. This finding highlights the highly challenging and specialized nature of this research area, requiring deep expertise and strong institutional support. The complexity of understanding drug resistance mechanisms and developing novel biomarkers presents significant challenges in terms of human capital and research resources.
Publications on drug resistance biomarkers in breast cancer are primarily found in high-impact journals such as “Journal of Cancers, International Journal of Molecular Sciences, Frontiers in Oncology, Frontiers in Immunology, Breast Cancer Research, Chinese Journal of Tissue Engineering Research, and International Immunopharmacology”. These journals have maintained a five-year impact factor above 4 and are classified as Q1 in relevant fields, including human study chemotherapy. The prominence of these journals underscores the high relevance of research in this area and the considerable challenge of publishing high-quality studies in such a competitive domain [28].
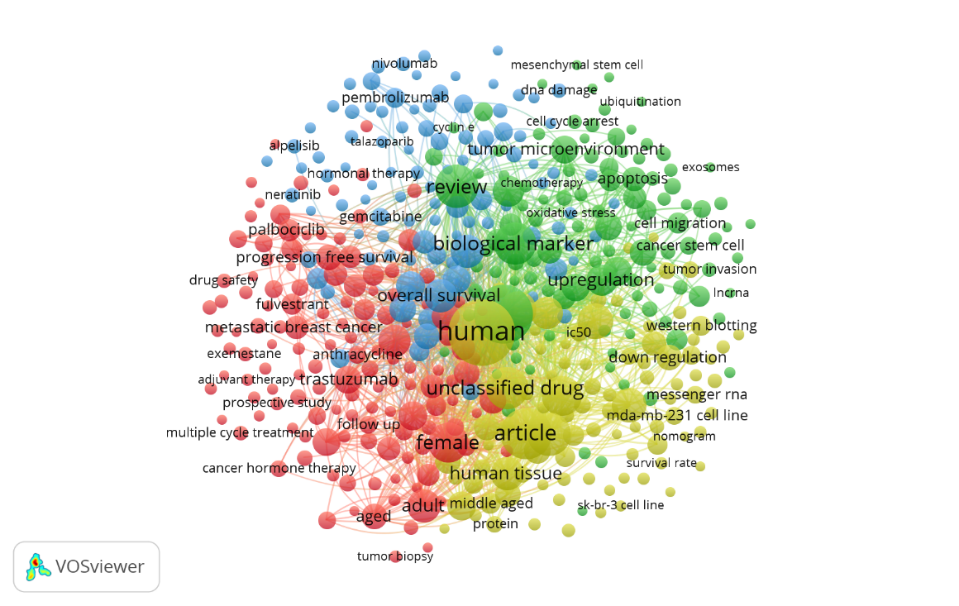
Fig. 6 shows that keyword analysis and thematic mapping reveal four main research clusters in the field of drug resistance biomarkers in breast cancer. The first cluster is centered on fundamental research related to the biology and epidemiology of breast cancer, frequently incorporating keywords such as “breast cancer”, “drug resistance”, “female”, “cancer survival”, and “hormonal therapy”. The second cluster focuses on prognosis and therapy prediction, including terms like “drug response”, “protein”, and “circulating tumor cell”. The third cluster reflects an interdisciplinary approach, emphasizing the role of the tumor microenvironment in drug resistance, with terms such as “tumor microenvironment”, “chemotherapy”, “apoptosis”, and “DNA damage”. The fourth cluster highlights emerging research themes that have the potential to shape future trends, where keywords such as “biomarker”, “human”, “human cell”, “unclassified drug”, “cell line”, and “cell migration” are used.
In summary, the dominance of China and the U. S. in this research domain is rooted in their extensive biomedical infrastructure and funding capacity. While their leadership drives scientific advancements, potential biases related to economic disparity, language accessibility, and institutional visibility must be acknowledged. Future efforts should promote more inclusive and equitable participation in global cancer research to ensure that findings and innovations reflect a broader range of populations and clinical realities.
These findings align with the study by [13], which identified biomarker research in breast cancer as one of the most significant emerging fields. Their study emphasized the importance of drug resistance mechanisms, molecular pathways, and immunological factors in determining therapy effectiveness. The similarities between these results and previous studies indicate that research trends in drug resistance biomarkers are consistent and reproducible across different countries and institutions.
Despite significant advancements in identifying drug resistance biomarkers, several challenges remain [29]. One of the key difficulties is the complexity of tumor heterogeneity, which affects the validity of biomarkers as predictors of treatment response. Additionally, methodological limitations in biomarker validation persist, highlighting the need for integrating multi-omics data to enhance prediction accuracy. Future studies should prioritize the integration of advanced technologies, including Artificial Intelligence (AI), single-cell sequencing, and liquid biopsy methods, to enhance the accuracy and clinical relevance of drug resistance biomarkers in breast cancer.
(a)

(b)
Fig. 6: The co-occurrence network of significant descriptors forms two main clusters (red and yellow) based on frequent descriptors (a). A Word Cloud (b) visualizes keyword frequency, with word size reflecting occurrence in studies on drug resistance biomarkers in breast cancer
Overall, this bibliometric analysis provides a comprehensive overview of the research landscape on drug resistance biomarkers in breast cancer. By identifying leading authors, high-impact journals, and key research trends, this study highlights the trajectory of scientific advancements in this field [30]. Given the growing complexity of breast cancer treatment challenges, continued research into biomarkers and drug resistance mechanisms is essential for developing more personalized therapeutic strategies and improving patient outcomes.
Recommendation for biomolecules with potential use as biomarkers for drug resistance in breast cancer
Breast cancer remains a major health concern globally, necessitating the development of effective biomarkers for diagnosis and treatment resistance. Several biomolecules have been identified as promising candidates for breast cancer management. Our review highlights several biomolecules that demonstrate high potential as predictive indicators and therapeutic targets for drug resistance in breast cancer, as shown in table 3.
Table 3: Biomolecules with potential use as predictive drug resistance biomarkers in breast cancer proposed in this study
| Biomolecule category | Drug response | Breast cancer model | Pathway | Effect | Ref |
| Circular RNA (circUBAP2) | Cisplatin resistance | HCC1937, BT-549, MDA-MB-436 (TNBC cells). | ↓miR-300 → ↑ASF1B → ↑PI3K/AKT/mTOR | Silencing ↑ sensitivity | [9] |
Circular RNA (hsa_circ_0006528) |
Adriamycin resistance | Adriamycin-resistant BC cells | ↓miR-7-5p → ↑Raf1 | Targeting ↑ response | [31] |
| Circular RNA (circKDM4C) | Adriamycin resistance | Adriamycin-resistant BC cells | ↓miR-548p → ↑PBLD | Downregulation ↓ resistance | [31] |
| Circular RNA (circMTO1) | Monastrol resistance | Monastrol-resistant BC cells | ↑TRAF4 → ↑Eg5 | Overexpression ↓ resistance | [31] |
| MicroRNA (miR-300) | Cisplatin resistance | TNBC cells | ↓ASF1B | Downregulation ↑ resistance | [9] |
| MicroRNA (miR-17, miR-20b) | Taxol resistance | Taxol-resistant BC cells | ↓NCOA3 | Targeting ↑ sensitivity | [32] |
| MicroRNA (miR-18a) | Paclitaxel resistance | TNBC patients | ↓Dicer enzyme | Overexpression ↑ resistance | [32] |
| Protein (ASF1B) | Cisplatin resistance | TNBC cells | ↑PI3K/AKT/mTOR | Silencing ↑ sensitivity | [9] |
| Protein (NCOA3) | Taxol resistance | Taxol-resistant BC cells | ↑Drug resistance | Downregulation ↑ response | [32] |
| Exosome (Del-1 protein) | Biomarker for early detection | Circulating exosomes in BC patients | Facilitates tumor communication. | Breast cancer early marker | [32] |
Abbreviations: TNBC: Triple-negative breast cancer, BC: Breast cancer, circRNA: Circular RNA, miRNA: MicroRNA, PI3K: Phosphoinositide 3-kinase, AKT: Protein kinase B, mTOR: Mechanistic target of rapamycin, ASF1 B: Anti-silencing function 1B, PBLD: Phenazine biosynthesis-like domain-containing protein, TRAF4: TNF receptor-associated factor 4, Eg5: Kinesin spindle protein, NCOA3: Nuclear receptor coactivator 3, circUBAP2: Circular ubiquitin-associated protein 2. Arrow notation explanation ↑: Increased expression or activation, ↓: Decreased expression or inhibition, →: leads to/causes/activates.
Circular RNAs (circRNAs) have emerged as critical regulators of drug resistance by acting as competing endogenous RNAs (ceRNAs) that modulate microRNA (miRNA) activity. For example, circular RNA ubiquitin-associated protein 2 (circUBAP2) has been identified as a key player in cisplatin resistance in Triple-Negative Breast Cancer (TNBC). It functions by sponging miR-300, leading to the upregulation of Anti-Silencing Function 1B (ASF1B) histone chaperone, which subsequently activates the PI3K/AKT/mTOR signaling pathway, promoting cell survival and chemoresistance [9]. Similarly, hsa_circ_0006528 and circKDM4C have been associated with adriamycin resistance by regulating the miR-7-5p/Raf1 axis and miR-548p/PBLD axis, respectively, with downregulation of these circRNAs improving chemotherapy efficacy [31].
MiRNAs also play a significant role in drug resistance mechanisms. miR-300 has been identified as a tumor suppressor in cisplatin-resistant TNBC, where its downregulation enhances ASF1B expression, contributing to chemoresistance [9]. Additionally, miR-17 and miR-20b have been linked to Taxol resistance by regulating Nuclear Receptor Coactivator 3 (NCOA3) expression, and targeting these miRNAs has shown potential in reversing Taxol resistance, and miR-18a has also been implicated in paclitaxel resistance, where its overexpression inhibits dicer expression and promotes drug resistance [32].
Proteins such as ASF1B and NCOA3 are key regulators of chemoresistance. ASF1B plays a pivotal role in cisplatin resistance in TNBC by activating the PI3K/AKT/mTOR pathway, and its knockdown has been shown to restore cisplatin sensitivity [9]. Similarly, NCOA3 is involved in taxol resistance by modulating key signaling pathways that promote cell survival and drug resistance [32]. Exosomes serve as carriers of drug resistance-associated biomolecules, facilitating intercellular communication in the tumor microenvironment. Del-1 protein, found in circulating exosomes of breast cancer patients, has been identified as a potential early detection biomarker, with high levels associated with increased tumor progression and chemoresistance [32].
Kinase inhibitors targeting Anaplastic Lymphoma Kinase (ALK), phosphoinositide 3-kinase (PI3K), and Human Epidermal Growth Factor Receptor 2 (HER2) have been widely studied in overcoming resistance to targeted therapies in breast cancer. However, resistance mechanisms such as secondary mutations and pathway compensations continue to pose challenges. Combination strategies involving kinase inhibitors and other targeted agents may enhance therapeutic efficacy and mitigate drug resistance [33, 34].
Based on table 3 presents the biomolecules associated with drug resistance in breast cancer, their corresponding resistance models, and the mechanisms by which they contribute to therapy resistance. Overall, the identification and application of biomolecules, including proteins, circRNAs, and miRNAs, represent a crucial advancement in breast cancer diagnostics and therapeutics. Continued research and clinical validation are essential to translating these molecular markers into effective tools for early detection and treatment resistance management.
Challenges and limitations
This study acknowledges limitations inherent to bibliometric analysis. Exclusive reliance on the Scopus database may result in selection bias due to the underrepresentation of non-English and regional publications. Citation inflation is another concern, particularly in review articles and high-impact journals, which may distort the visibility and influence of original research. To address these issues, future studies should incorporate multiple databases, apply normalized citation metrics, and utilize open-source tools. Standardized protocols for data cleaning and keyword selection are also recommended to enhance methodological rigor and reproducibility.
Future directions
AI and Machine Learning (ML) significantly enhance bibliometric research by automating data analysis, revealing patterns, and predicting future trends. Tools such as Natural Language Processing (NLP) and clustering algorithms help classify and interpret large datasets, while ML models support forecasting and evaluating impact beyond citation metrics. These technologies also assist in identifying new biomarker targets and research priorities. Future biomarker research will benefit from multi-omics integration, high-throughput screening, and advanced bioinformatics, allowing for more accurate detection of resistance mechanisms and guiding personalized therapies. International collaboration is equally vital, as it promotes knowledge sharing, resource accessibility, and inclusive research. Such efforts accelerate the validation and clinical application of biomarker discoveries across diverse populations and health systems.
CONCLUSION
This bibliometric study analyzed 252 publications from 2020 to 2025 focused on drug resistance biomarkers in breast cancer. China led with 74 publications (29.4%), followed by the United States with 59 publications (23.4%), underscoring their dominant roles in global output. Research peaked in 2022 and has since diversified. Major research themes included microRNAs, circular RNAs, and protein-based resistance mechanisms. These findings support further strategic collaboration, prioritization of emerging molecular targets, and development of precision therapy. Researchers, clinicians, and policymakers can use these insights to guide future research, funding allocation, and clinical trial design.
FUNDING
Nil
AUTHORS CONTRIBUTIONS
Wahyu Utami conceptualized the study, contributed to the study design, and provided overall supervision. Eva Nursoleha was responsible for data acquisition, data analysis, and the initial drafting of the manuscript. Both authors contributed to the critical revision and final approval of the manuscript.
CONFLICT OF INTERESTS
Declared none
REFERENCES
The Global Cancer Observatory. Cancer statistics for the year 2020: an overview. Int J Cancer. 2022;149(4):778-89. doi: 10.1002/ijc.33588, PMID 33818764.
Fitria N, Rivaldi RS, Permatasari D. An rinvr scoring evaluation of ondansetron dexamethasone and ranitidine as antiemetics in breast cancer chemotherapy. Int J App Pharm. 2025;17(1):101-6. doi: 10.22159/ijap.2025.v17s1.15.
Ansari AM, Sharma NK, Tripathi K, Bhardwaj G, Chauhan R, Tonk RK. Overcoming multifaceted challenges in cancer treatment: targeting signal transduction pathways and tumor microenvironment for enhanced therapeutic efficacy. J Appl Pharm Sci. 2025;12(3):1-24. doi: 10.7324/JAPS.2025.204012.
Handayani F, Fitria N, Sari YO, Almahdy A. Impact of the combination of doxorubicin cyclophosphamide and docetaxel on ca 15-3 biomarker levels in breast cancer patients: a comparative study between delayed and non-delayed chemotherapy. Int J App Pharm. 2025;17(1):82-8. doi: 10.22159/ijap.2025.v17s1.12.
Wikantyasning ER, Wahyuni AS, Kusumawati IT, Saifudin A, Suhendi A. Antiproliferative properties of tiliroside from guazuma ulmifolia lamk on T47D and MCF7 cancer cell lines. Natl J Physiol Pharm Pharmacol. 2016;6(6):627-33. doi: 10.5455/njppp.2016.6.0617727072016.
Ye S, Chen S, Yang X, Lei X. Drug resistance in breast cancer is based on the mechanism of exocrine non-coding RNA. Discov Oncol. 2024;15(1):138. doi: 10.1007/s12672-024-00993-3, PMID 38691224.
Lohith NC, Ranjini P, Devaraju. Design synthesis anticancer activity and molecular docking studies of lignin pyrrole derivatives as a JAK3 inhibitor. Asian J Pharm Clin Res. 2024;17(12):64-72. doi: 10.22159/ajpcr.2024v17i12.52831.
Chand PN, Komanapalli S, Kuna R, Chowdari B, Devi U, Das B. Clinicopathological study of breast cancer and immunohistochemical expression of BRCA1 and BRCA2 a prospective study. Asian J Pharm Clin Res. 2024;17(1):92-6. doi: 10.22159/ajpcr.2024.v17i1.48713.
Wang L, Yang X, Zhou F, Sun X, Li S. Circular RNA UBAP2 facilitates the cisplatin resistance of triple negative breast cancer via microRNA-300/anti-silencing function 1B histone chaperone/PI3K/AKT/mTOR axis. Bioengineered. 2022;13(3):7197-208. doi: 10.1080/21655979.2022.2036894, PMID 35263216.
Afzal S, Hassan M, Ullah S, Abbas H, Tawakkal F, Khan MA. Breast cancer; discovery of novel diagnostic biomarkers, drug resistance and therapeutic implications. Front Mol Biosci. 2022;9:783450. doi: 10.3389/fmolb.2022.783450, PMID 35265667.
Cardona Galeano W, Yepes AF, Ramirez Malule H. Triazole based click chemistry as strategic for novel therapeutics treatments of cancer: a bibliometric analysis 2007-2024. J Appl Pharm Sci. 2025;15(5):64-74. doi: 10.7324/JAPS.2025.221899.
Donthu N, Kumar S, Mukherjee D, Pandey N, Lim WM. How to conduct a bibliometric analysis: an overview and guidelines. J Bus Res. 2021 Sep;133:285-96. doi: 10.1016/j.jbusres.2021.04.070.
Duan Y, Zhang P, Zhang T, Zhou L, Yin R. Characterization of global research trends and prospects on platinum-resistant ovarian cancer: a bibliometric analysis. Front Oncol. 2023 Jun 5;13:1151871. doi: 10.3389/fonc.2023.1151871, PMID 37342181.
Aria M, Cuccurullo C. Bibliometrix: an R-tool for comprehensive science mapping analysis. Journal of Informetrics. 2017;11(4):959-75. doi: 10.1016/j.joi.2017.08.007.
Effendi DN, Irwandani AW, Anggraini W, Jatmiko A, Rahmayanti H, Ichsan IZ. Bibliometric analysis of scientific literacy using VOS viewer: analysis of science education. J Phys: Conf Ser. 2021;1796(1):12096. doi: 10.1088/1742-6596/1796/1/012096.
Sari AA, Munawaroh R, Sofyanita EN. Bibliometric analysis of antibacterial activity of Centella asiatica: a study based on scopus database. J Appl Pharm Sci. 2023;13(11):1-16. doi: 10.7324/JAPS.2023.139686.
Utami W, Sari AA, Prayitno HJ. Publications affiliated with universitas muhammadiyah surakarta from Indonesia in scopus database. Desidoc Jl Lib Info Technology. 2024;44(4):203-11. doi: 10.14429/djlit.44.4.19429.
Van Eck NJ. VOSviewer manual; 2022. Available from: www.vosviewer.com.
Yeung AW, Tzvetkov NT, Balacheva AA, Georgieva MG, Gan RY, Jozwik A. Lignans: quantitative analysis of the research literature. Front Pharmacol. 2020 Feb 7;11:37. doi: 10.3389/fphar.2020.00037, PMID 32116713.
Rani R, Gahirwar A, Yadav S. Review article medicinal plants from the himalayas: insights into their anticancer effects. Int J Pharm Pharm Sci. 2025;17(2)16-25. doi: 10.22159/ijpps.2025v17i2.52891.
M JC, Lopez Herrera A, Herrera Viedma E, Herrera F. Full-text citation analysis: a new method to enhance scholarly networks. J Am Soc Inf Sci Technol. 2013;64(9):1852-63. doi: 10.1002/asi.22883/abstract.
Ha CT, Thao TT, Trung NT, Huong LT, Dinh NV, Trung T. A bibliometric review of research on STEM education in Asean: science mapping the literature in scopus database, 2000 to 2019. EURASIA J Math Sci Tech Ed. 2020;16(10):em1889. doi: 10.29333/ejmste/8500.
Romero L, Portillo Salido E. Trends in sigma-1 receptor research: a 25 Y bibliometric analysis. Front Pharmacol. 2019 May 24;10:564. doi: 10.3389/fphar.2019.00564, PMID 31178733.
Amornmarn R, Bui MM, Prempree TB, Masood S. Molecular predictive factors for local recurrence and distant metastasis of breast cancer after lumpectomy with postoperative radiation therapy. Ann Clin Lab Sci. 2000;30(1):33-40. PMID 10678581.
Shi Q, Yang W, Ouyang Y, Liu Y, Cai Z. CXCR4 promotes tumor stemness maintenance and CDK4/6 inhibitors resistance in ER-positive breast cancer. Breast Cancer Res. 2025;27(1):15. doi: 10.1186/s13058-025-01965-3, PMID 39875975.
Piepenbrink A, Nurmammadov E. Topics in the literature of transition economies and emerging markets. Scientometrics. 2015;102(3):2107-30. doi: 10.1007/s11192-014-1513-2.
Ferreira FA. Mapping the field of arts-based management: bibliographic coupling and co-citation analyses. J Bus Res. 2018 Apr;85:348-57. doi: 10.1016/j.jbusres.2017.03.026.
Ma S, Ding W, Liu Y, Zhang Y, Ren S, Kong X. Industry 4.0 and cleaner production: a comprehensive review of sustainable and intelligent manufacturing for energy-intensive manufacturing industries. J Clean Prod. 2024 Aug 15;467:142879. doi: 10.1016/j.jclepro.2024.142879.
Matthews H, Hanison J, Nirmalan N. Omics-informed drug and biomarker discovery: opportunities, challenges and future perspectives. Proteomes. 2016;4(3):28. doi: 10.3390/proteomes4030028, PMID 28248238.
Krauss A, Danus L, Sales Pardo M. Early career factors largely determine the future impact of prominent researchers: evidence across eight scientific fields. Sci Rep. 2023;13(1):18794. doi: 10.1038/s41598-023-46050-x, PMID 37914796.
Misir S, Yaman SO, Petrovic N, Sumer C, Hepokur C, Aliyazicioglu Y. Circrnas in drug resistance of breast cancer. Oncol Res. 2022;30(4):157-72. doi: 10.32604/or.2022.027547, PMID 37304411.
Ghafouri Fard S, Khanbabapour Sasi A, Abak A, Shoorei H, Khoshkar A, Taheri M. Contribution of mirnas in the pathogenesis of breast cancer. Front Oncol. 2021 Nov 5;11:768949. doi: 10.3389/fonc.2021.768949, PMID 34804971.
Kota SS, Bandhakavi S. Cost effectiveness of combination anti-cancer drug therapy in the management of HER2-positive breast cancer: a meta-analysis. Asian J Pharm Clin Res. 2023;16(12):20-5. doi: 10.22159/ajpcr.2023.v16i12.48522.
Zhong L, Li Y, Xiong L, Wang W, Wu M, Yuan T. Small molecules in targeted cancer therapy: advances challenges and future perspectives. Signal Transduct Target Ther. 2021;6(1):201. doi: 10.1038/s41392-021-00572-w, PMID 34054126.