
Int J App Pharm, Vol 18, Issue 1, 2026, 199-205Original Article
STUDY THE EFFECT OF CONJUGATED NOVEL ULTRASHORT ANTIMICROBIAL PEPTIDES IN COMBINATION WITH FARNESYL AGAINST RESISTANCE STRAINS OF PSEUDOMONAS AERUGINOSA
ALI SALAMAa , SANDRELLA MOHAMMED SAIFb*
, SANDRELLA MOHAMMED SAIFb*
aDepartment of Pharmaceutical Chemistry, Faculty of Pharmaceutical Sciences, The Hashemite University, Zarqa-13133, Jordan. bFaculty of Pharmacy, Middle East University, Amman-Jordan
*Corresponding author: Sandrella Mohammed Saif; *Email: sande2001.sf@gmail.com
Received: 16 May 2025, Revised and Accepted: 08 Nov 2025
ABSTRACT
Objective: The objective of this study was to design and evaluate a novel ultrashort antimicrobial peptide, San-112003, structurally conjugated with a farnesyl chain, for its potential antibacterial activity against both antibiotic-susceptible and multi-drug-resistant (MDR) strains of Pseudomonas aeruginosa. Given the rising threat of antimicrobial resistance and the limitations of conventional antibiotics, antimicrobial peptides (AMPs) are explored as a promising alternative. This study aimed to assess the efficacy, toxicity, and potential synergism of San-112003 with conventional antibiotics.
Methods: In this study, we designed and synthesized a novel ultrashort antimicrobial peptide, San-112003, which incorporates repeating tripeptide units (tryptophan–arginine–glycine) chemically conjugated to a farnesyl chain. This structural design enhances membrane targeting and antimicrobial potency. The antimicrobial efficacy of San-112003 was evaluated against both antibiotic-susceptible and MDR strains of P. aeruginosa. Hemolytic toxicity assays were conducted on human red blood cells, and synergy studies were performed in combination with conventional antibiotics, including gentamicin.
Results: San-112003 was successfully synthesized with 99% purity and correct molecular mass confirmation. The peptide showed low cytotoxicity in MTT assays across tested concentrations. It demonstrated potent bactericidal activity against Pseudomonas aeruginosa, with MIC and MBC values of 0.5 µg/ml for the control strain and 2.5 µg/ml for the MDR strain. Hemolysis testing indicated minimal toxicity at concentrations relevant to antimicrobial activity. Synergy testing revealed the greatest enhancement in bacterial inhibition when combined with gentamicin, with FICI values of 0.075 for the control strain and 0.20 for the MDR strain. Additionally, the peptide–antibiotic combinations reduced bacterial biofilm viability more effectively than the peptide alone.
Conclusion: San-112003 is a promising AMP–farnesyl conjugate with potent activity against P. aeruginosa, including MDR strains. Its synergism with gentamicin and low hemolytic toxicity suggest that AMP–antibiotic combinations could serve as a valuable strategy against antibiotic resistance.
Keywords: Pseudomonas aeruginosa, Antimicrobial peptides (AMPs), Multi-drug-resistant, Antibiotics, Bacterial activity
© 2026 The Authors. Published by Innovare Academic Sciences Pvt Ltd. This is an open access article under the CC BY license (https://creativecommons.org/licenses/by/4.0/)
DOI: https://dx.doi.org/10.22159/ijap.2026v18i1.55076 Journal homepage: https://innovareacademics.in/journals/index.php/ijap
INTRODUCTION
The indiscriminate use of conventional antibiotics in recent years for treating human infections has significantly contributed to the emergence of bacterial strains resistant to currently available clinical antibiotics. The substantial global rise in antimicrobial resistance poses a serious threat to the global population and has generated considerable concern among public health authorities and governmental agencies. This escalation in resistant bacterial strains has been accompanied by a notable reduction in the development of new antibiotics for clinical application [1]. Understanding the mechanisms underlying bacterial resistance to specific antibiotics is crucial for developing effective solutions and designing novel antibiotics capable of targeting and eradicating resistant bacterial strains; consequently, the primary mechanisms of antimicrobial resistance are enzymatic degradation of antimicrobial agents, modification of bacterial proteins that serve as antimicrobial targets, and alterations in membrane permeability to antibiotics. Antibiotic resistance can be mediated either through plasmids or by genetic alterations within the bacterial chromosome [2].
Multidrug-resistant (MDR) Pseudomonas aeruginosa remains a prominent nosocomial pathogen of global concern. It is classified among the ESKAPE group of pathogens and designated a “serious threat” by the CDC due to its ability to cause difficult-to-treat infections in hospitalized patients, particularly in ICUs and immunocompromised individuals. Recent epidemiological studies report that P. aeruginosa accounts for approximately 7–10% of all healthcare-associated infections worldwide. Resistance rates vary but can reach up to 24% for β-lactams and over 30% for carbapenems in ICU settings, especially in ventilator-associated pneumonia and catheter-related infections [3, 4]. These statistics highlight the urgent need for new therapeutics specifically targeting this resilient organism.
Ultrashort antimicrobial peptides (USAMPs) have emerged as a promising solution to combat resistant bacteria such as P. aeruginosa. USAMPs consist of fewer than 10 amino acids and are easier and more cost-effective to synthesize than traditional AMPs. Their small size also results in enhanced stability, reduced hemolytic toxicity, and improved selectivity toward bacterial membranes. Unlike larger AMPs, USAMPs generally lack complex disulfide bond structures, making them more suitable for clinical translation. Furthermore, conjugation with hydrophobic moieties—such as farnesyl—can enhance membrane anchoring, improve antimicrobial efficacy, and reduce susceptibility to enzymatic degradation.
Antimicrobial peptides (AMPs) represent a promising class of antimicrobial agents, distinguished by their high potency and efficacy. These peptides offer a potential alternative to conventional antibiotics, which often encounter resistance from microbes, and may also function synergistically with existing antibiotics to combat diverse bacterial strains [5]. Antimicrobial peptides (AMPs) are relatively small, with lengths varying from 10 to 50 amino acids. They possess a net positive charge ranging from +3 to +9 and contain approximately 30% hydrophobic residues, contributing to their amphipathic nature. These structural attributes enable AMPs to form pores in target membranes, leading to cell lysis and subsequent death [6].
The classical mechanism of action of antimicrobial peptides (AMPs) involves the disruption of cell membranes [7]. Although cationic AMPs can electrostatically interact with the surface of the negatively charged cell membrane [8], some AMPs induce membrane damage through specific interactions with membrane constituents, while other compounds target intracellular molecules. AMPs generally demonstrate broad-spectrum activity against pathogenic microorganisms. Due to their non-specific interactions, AMPs typically do not engage specific receptors, which limits the development of resistant phenotypes in microbial targets. Nevertheless, resistance to certain AMPs has been observed. For example, Staphylococcus aureus has developed resistance to dermcidin through the induction of a specific protease, while some Gram-negative bacteria exhibit reduced sensitivity to cationic AMPs by altering their lipopolysaccharide structures via the PhoP/PhoQ regulatory system [9]. Despite these resistance mechanisms, they only modestly impact the minimal inhibitory concentration (MIC) required for AMP effectiveness.
The traditional AMPs have limitations that make the clinical development of these peptides very difficult, including high manufacturing costs, relatively large size with a complex disulfide pattern, high hemolytic activity, and relatively high hydrophobicity, which causes toxicity to human cells [10].
These problems can be overcome by developing ultrashort antimicrobial peptides (USAMPs) by selecting specific amino acids to be involved in the designs. These USAMPs have several advantages over conventional AMPs, including cost-effectiveness as they are not hard to synthesize chemically. USAMPs have relatively low hydrophobicity, so they are supposed to have low toxicity towards human cells and red blood cells, thus low hemolysis action. These peptides have good selectivity toward bacterial cells because of the desirable interactions between the negative head groups on bacterial cell surfaces and the cationic side chains of the peptides [11]. This experiment investigates the effects of a newly designed Ultra Short anti-microbial peptide, composed of Tryptophan-Arginine-Glycine fused with a farnesyl chain, and its effect on Pseudomonas aeruginosa species.
MATERIALS AND METHODS
Bacterial cultures
Methicillin-resistant Staphylococcus aureus (ATCC 33591) as a biofilm-forming bacteria, Pseudomonas aeruginosa (ATCC 9027) as a planktonic strain and Pseudomonas aeruginosa (ATCCBAA-2108) as resistance strain were obtained from the American Type Tissue Culture Collection (ATCC, Manassas, VA, USA).
Peptide synthesis and design
The peptides utilized in this study were synthesized using the solid-phase method and Fmoc chemistry and obtained in a lyophilized state from GL Biochem Ltd., Shanghai, China. San-112003 is an ultrashort antimicrobial peptide consisting of four repeating units of the tripeptide WRG (tryptophan–arginine–glycine), with a single farnesyl chain chemically conjugated to the C-terminus of the full peptide, forming a linear structure. In this design, tryptophan contributes to hydrophobic interaction, arginine facilitates electrostatic binding to bacterial membranes, and the farnesyl moiety enhances membrane anchoring and overall hydrophobicity.
The synthesized peptide was purified using reverse-phase high-performance liquid chromatography (RP-HPLC) on a C18 Intersil® ODS-SP column. The elution was performed using an acetonitrile/H₂O–TFA gradient at a flow rate of 1.0 ml/min. The purity of San-112003 was ≥99%, as confirmed by analytical HPLC (fig. 2). Electrospray ionization mass spectrometry (ESI-MS) was used to verify the molecular weight and identity of the purified peptide (fig. 3).
Biofilm antibiotic used in this study
Levofloxacin, Chloramphenicol, Rifampicin Amoxicillin, Clarithromycin, Doxycycline, Vancomycin and Cefixime were obtained from sigma Aldrich
MTT assay
In this study, cytotoxicity was evaluated using the MTT assay on Vero cells (African green monkey kidney epithelial cells, ATCC CCL-81), a standard model for initial toxicity screening due to their sensitivity to toxic insults and ease of maintenance. Cells were exposed to San-112003 at a range of concentrations (0.5, 1, 2.5, 5, and 10 µg/ml), selected to include the MIC range (0.5–2.5 µg/ml) and higher concentrations to assess dose-response. After 24 h of exposure, MTT reagent was added and formazan crystals were solubilized in DMSO. Absorbance was measured at 550 nm. While Vero cells provide reliable baseline toxicity data, we acknowledge that they are not tissue-specific to P. aeruginosa infections. Future work will expand cytotoxicity testing to include keratinocytes (HaCaT) and human macrophage models (e. g., THP-1-derived) for improved relevance to respiratory and wound infection contexts.
Determination of the minimum inhibitory concentration and minimum bactericidal concentration of SAN-112003 and the individual antibiotics
SAN-112003, along with eight individual antibiotics (levofloxacin, amoxicillin, rifampicin, doxycycline, gentamicin, and cefixime) with minimum inhibitory concentration (MIC) and minimum bactericidal concentration (MBC), each representing different mechanisms of antimicrobial action, were assessed using the micro-broth dilution method as specified by the Clinical and Laboratory Standards Institute [12]. Substantially, Pseudomonas aeruginosa species were grown in Muller-Hinton broth (MHB) medium and diluted to CFU/ml in the same medium before use. Speaking of SAN-112003, other conventional antibiotics were prepared in different concentrations, aliquoted with the bacteria in 96-well plates, and incubated for 18-24 h at 37 °C. Cell growth and minimum inhibitory concentrations (MICs) were quantified using an ELISA reader at an optical density of λ = 570 nm. Minimum bactericidal concentrations (MBCs) were determined by transferring 10 μl from the negative wells onto agar plates, which were then incubated for 24 h at 37 °C. The MBC was defined as the concentration that achieved a 99.9% reduction in viable cells. All experiments were conducted in triplicate.
The determination of the synergistic activity of SAN-112003 in combination with individual antibiotics
According to the broth micro-dilution checkerboard technique, the synergistic activity and the MIC values of SAN-112003, when combined with antibiotics, were tested and determined [13]. But in this essay, each micro-titer well contained a combination of one peptide and one antibiotic at varying concentrations. Specifically, 25 µl** of each peptide concentration and 25 µl** of each antibiotic concentration were added to six wells of a sterile flat-bottomed 96-well plate, which already contained 50 µl** of the diluted bacterial suspension. All experiments were carried out in triplicate.
Determination of the fractional inhibitory concentration
As the fractional inhibitory concentration (FIC) represents the summation of the inhibitory concentration values of each antimicrobial component in the antimicrobial combination divided by the inhibitory concentration of the individual antimicrobial agent [14], the evaluation and analysis of the fractional inhibitory concentration (FIC) index and antimicrobial efficacy of peptide-antibiotic combinations were performed using the broth micro-dilution checkerboard method. It has been demonstrated that an FIC index is synergistic (FIC<0.5), additive (FIC index = 0.5-1), and antagonistic (FIC>1). FICI = (MIC peptide in combination/MIC peptide alone)+(MIC antibiotics in combination/MIC antibiotics)
Hemolytic activity of SAN-112003
The potential of SAN-112003 to induce damage to the membranes of normal mammalian cells and its ability to cause hemolysis of horse erythrocytes was assessed through erythrocyte hemolytic assays [15]. Briefly, 2 ml of human blood was transferred into a 50-mL centrifuge tube and centrifuged at 3000 × g for 5 min. After removing the supernatant, the cell pellet was re-suspended in 48 ml of phosphate-buffered saline (PBS) and centrifuged again at 3000 × g for 5 min; this washing step was repeated three times. The final cell pellet was re-suspended in a sterile tube with 50 ml of PBS to achieve a 4% red blood cell (RBC) suspension. PBS solutions containing various peptide concentrations were prepared, and 1 ml of each peptide concentration was combined with 1 ml of the RBC suspension. Controls were prepared by adding 5 µl** of Triton X-100 to 1 ml of RBC suspension (positive control), while blanks were made by mixing 1 ml of RBC suspension with PBS. The mixtures were incubated for 60 min at 37 °C. Following incubation, the tubes were gently vortexed, and 1 ml of each sample was transferred to sterilized Eppendorf tubes, centrifuged at 3000 × g for 5 min, and 100 µl** of each supernatant was aliquoted into wells of a 96-well plate. Absorbance was measured at λ = 570 nm using an absorbance microplate reader. The percentage of hemolysis was calculated using the following equation
Hemolysis %= ((A − A0)/(A[?] − A0)) × 100
Where A is OD 450 with the peptide solution, A0 is OD 450 of the blank and AX is OD 450 of control (0.1% Triton X-100).
Biofilm activity assay
Biofilm formation of S. aureus (ATCC 33591) was performed as reported in earlier studies employing the Calgary biofilm device (Innovotech, Edmonton, Canada). The bacteria was left to grow in TSB at 37 °C for 20 h. Then, a concentration of 107 CFU/ml was prepared by diluting the cultures in the same medium. An amount of 150 μl of the bacterial culture was added to each peg lid to allow the formation of biofilm on the purpose-designed pegs (96-peg lids), The pegs were then incubated for 20 h at 37 °C under 125 rpm rotation. Blank lanes were prepared by adding 150 µl** TSB to six wells. PBS was used to wash pegs three times to discard planktonic cells after biofilm formation [16].
Statistical analysis
All experiments were performed in triplicate, and data are presented as mean±standard deviation (SD) unless otherwise stated. Statistical significance for MTT assays was determined using one-way ANOVA, followed by Tukey’s multiple comparisons test. For MIC and MBC comparisons, unpaired Student’s t-tests were used. A p-value<0.05 was considered statistically significant. All statistical analyses were conducted using GraphPad Prism (version X. XX).
Fractional inhibitory concentration index (FICI) values were calculated using geometric means of MICs from three independent replicates, as this approach better reflects the multiplicative nature of dilution-based assays and enhances reproducibility.
RESULTS
Design and synthesis of SAN-112003
San-112003 was designed as an ultrashort conjugated tripeptide consisting of alternating subunits of both the amino acids Tryptophan-Arginine-Glycine (WRG) that were conjugated to farnesyl (fig. 1), The peptide’s characterization profile, including its identity and purity, was confirmed using HPLC and ESI-MS, respectively (fig. 2 and 3).
Fig. 1: Structure of san-112003 peptide
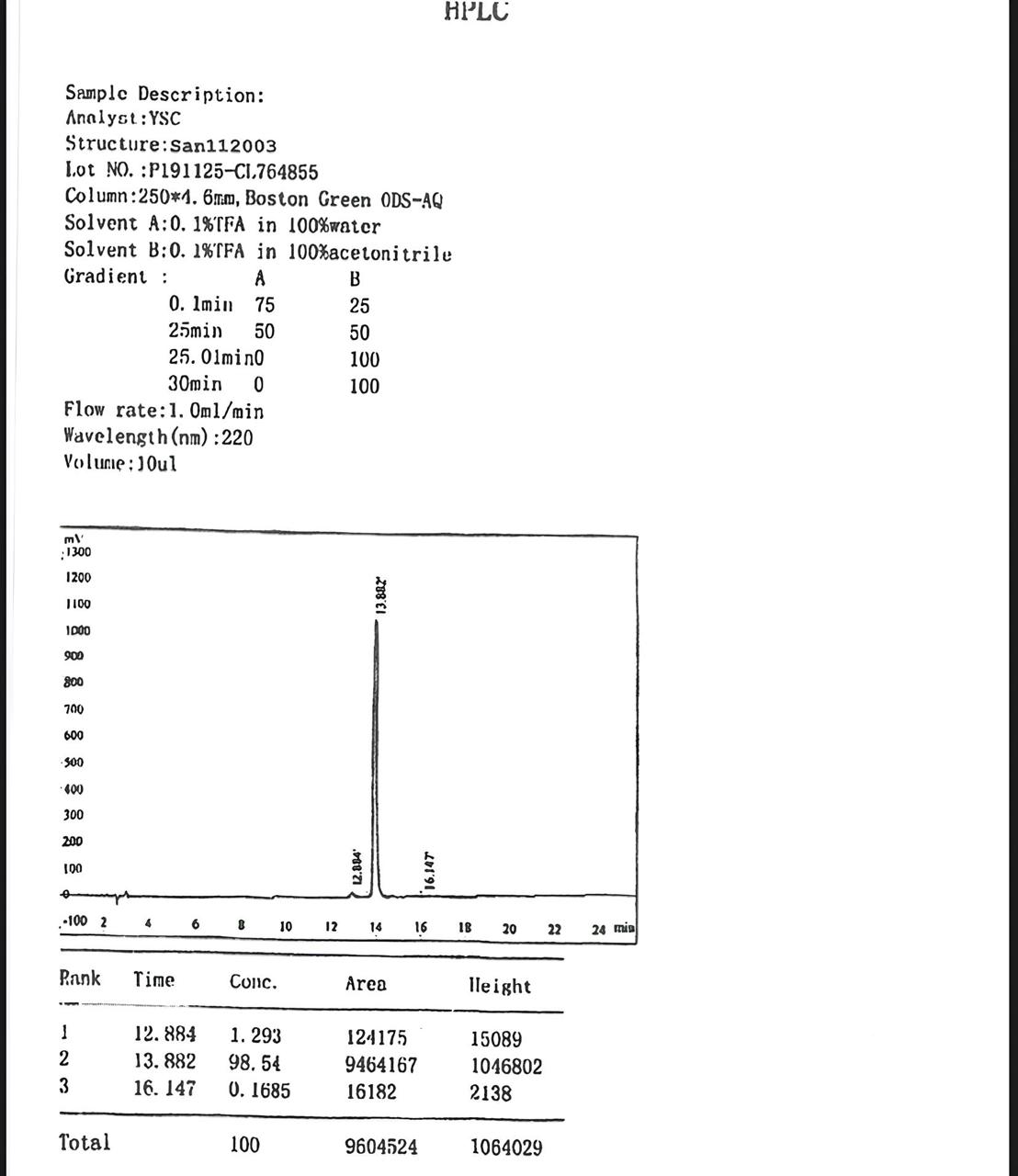
Fig. 2: Reverse phase high-performance liquid chromatography (HPLC) chromatogram of the San112003 indicating 99% purity of the synthesized peptide. The absorbance was at λ = 214 nm
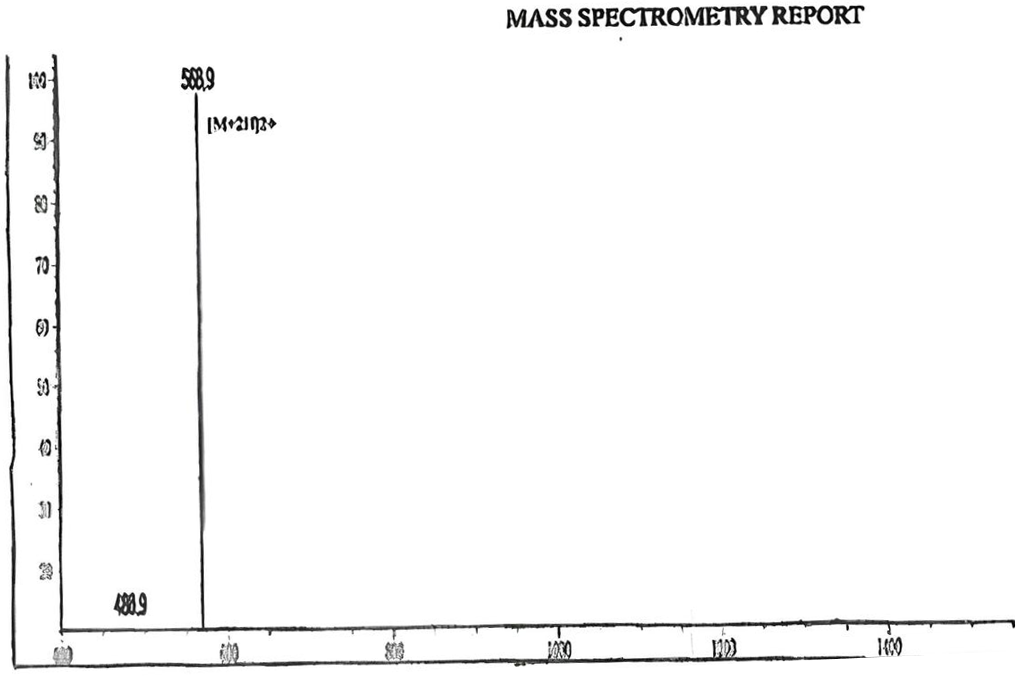
Fig. 3: Positive electrospray ionization mass spectrometric (ESI-MS) analysis of the San112003. The peptide shows major peak in the +2 state of 568.9 Da
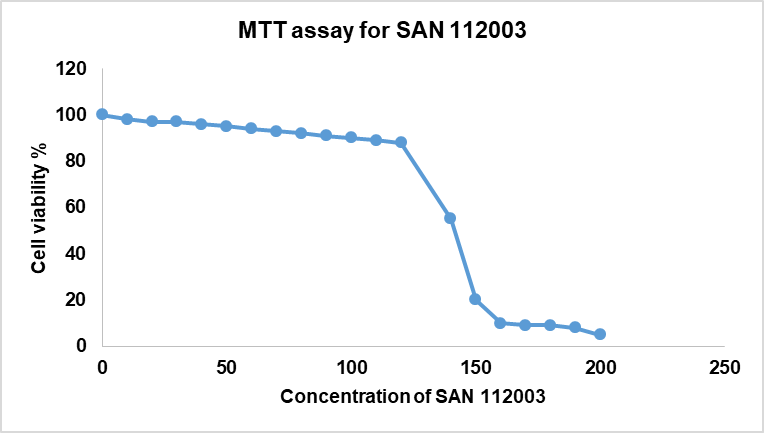
Fig. 4: MTT assay for San112003
Table 1: MIC and MBC values (µg/ml) with SD and p-values
| Compound | Control P. aeruginosa (9027) MIC (MBC) µg/ml | MDR P. aeruginosa (BAA-2108) MIC (MBC) µg/ml | p-value |
| San-112003 | 0.5±0.0 (0.5±0.0) | 2.5±0.0 (2.5±0.0) | <0.001 |
| Levofloxacin | 9.0±1.0 (9.0±1.0) | 30.0±2.0 (30.0±2.0) | <0.001 |
| Amoxicillin | 25.0±1.2 (25.0±1.2) | 200.0±5.0 (200.0±5.0) | <0.001 |
| Doxycycline | 6.0±0.5 (25.0±1.0) | 34.0±2.0 (45.0±3.0) | <0.001 |
| Cefixime | 6.0±0.5 (10.0±1.0) | 82.0±3.0 (110.0±4.0) | <0.001 |
| Gentamicin | 2.0±0.0 (2.0±0.0) | 20.0±0.5 (20.0±0.5) | <0.001 |
MIC: minimum inhibitory concentration; MBC: minimum bactericidal concentration. Value are mean±SD (n = 3). MDR: multi-drug-resistant strain. p<0.05 considered significant.
Table 2: Hemolytic activity of San-112003 against human erythrocytes after 60 min incubation
| Concentration (μM) | Concentration (µM) | Hemolysis % |
| 5 | 4.39 | 0 |
| 10 | 8.78 | 0 |
| 20 | 17.57 | 0 |
| 40 | 35.14 | 0 |
| 60 | 52.70 | 0 |
| 80 | 70.27 | 1 |
| 100 | 87.84 | 1 |
MTT assay
Fig. 4 shows the MTT assay for San112003.
Antibacterial activity of the peptide
Table 1 shows San-112003, 0.5 µg/ml, and 2.5 µg/ml activity against P. aeruginosa strains ATCC 9027 and BAA-2108, respectively. One is a control strain, while the other is MDR. The MBC and MIC values were identical for both bacterial strains. The MIC and MBC values of the eight distinct antibiotics are tabulated in table 1.
Antibacterial activity of SAN-112003 along with conventional antibiotics
The findings in table 3 demonstrate the impact of various peptide San-112003/antibiotic combinations on both bacterial strains. In the combination of San-112003 and gentamicin, the most significant results were achieved against both control and MDR P. aeruginosa strains, with FICI values of, respectively.
The findings in table 3 demonstrate the effect of various San-112003/antibiotic combinations on both bacterial strains. The lowest FICI values, indicating the strongest synergistic effect, were observed in combination with gentamicin (FICI = 0.075 for control and 0.20 for MDR strains).
Biofilm activity
The result in fig. 5 indicates that the combination of the peptide with antibiotics lead to dramatically decrease in the viable cell of the bacteria.
Table 3: The checkerboard results of the effect of combinations of the peptide san-112003 with the antibiotics used in the study against both bacterial strains
| Bacteria strains | Antibiotics | MIC | FICI* |
| Antibiotics | Antibiotics/San-112003 | ||
| Control P. aeruginosa | Levofloxacin | 9 | 1 |
| Rifampicin | 15 | 2 | |
| Amoxicillin | 25 | 10 | |
| Cefixime | 6 | 0.25 | |
| Gentamicin | 2 | 0.05 | |
| MDR P. aeruginosa | Levofloxacin | 30 | 10 |
| Rifampicin | 50 | 20 | |
| Amoxicillin | 200 | 90 | |
| Cefixime | 82 | 40 | |
| Gentamicin | 20 | 2 |
Notably, the MIC and MBC values for San-112003 were identical (0.5 µg/ml for control and 2.5 µg/ml for MDR P. aeruginosa), indicating a bactericidal mode of action. According to accepted criteria, a compound is considered bactericidal when the MBC is ≤4 times the MIC, which is clearly met in this case.
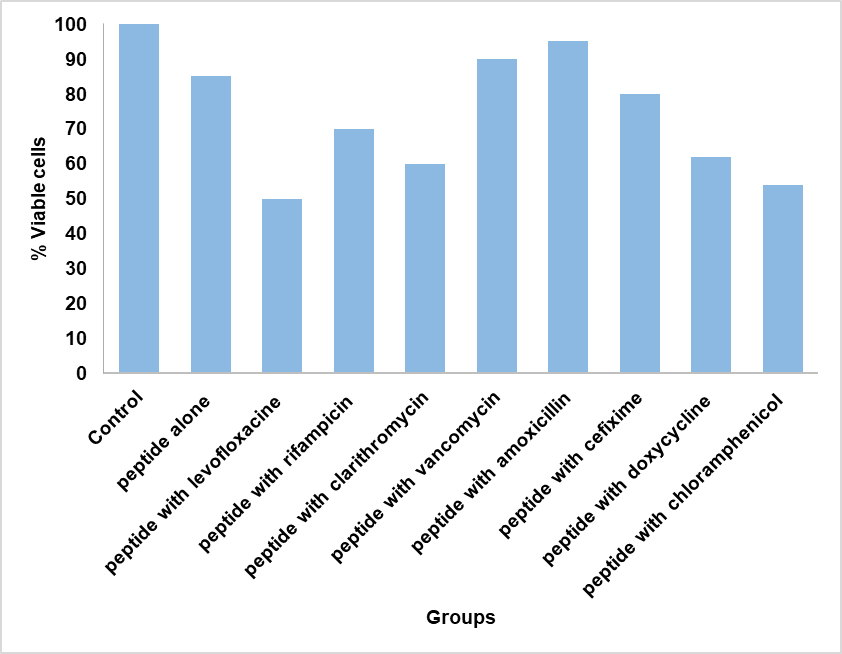
Fig. 5: Percentage of viable cell after the treatment with peptide
DISCUSSION
The rise of antibiotic resistance has reached alarming levels, leading to the emergence of multidrug-resistant bacteria. This crisis underscores the urgent need for novel antimicrobial agents with unique mechanisms of action [17]. Antimicrobial peptides (AMPs) have emerged as a highly promising class of such agents. Unlike traditional antibiotics, AMPs have a lower propensity to induce resistance and are effective against multidrug-resistant strains, making them a critical tool in addressing the challenges posed by resistant pathogens [18].
AMPs primarily exert their antimicrobial effects by targeting bacterial cell membranes. The proposed mechanism involves the electrostatic attraction between the positively charged moieties of AMPs and the negatively charged bacterial cell surfaces. This interaction facilitates peptide entry, leading to pore formation, membrane disruption, and ultimately, bacterial cell death [19]. Our research focused on designing novel peptides with a net cationic charge of approximately+3, which aligns with the established mechanism of AMP action. The designed peptides were structured to balance other physicochemical properties, such as hydrophobicity, to optimize their antimicrobial activity [20].
In our study, the cationic charge of the peptide San-112003 is predominantly attributed to the presence of arginine residues. Our findings are consistent with previous studies, which demonstrated that membrane disruption by AMPs is highly dependent on cationic residues, particularly arginine [21]. The guanidine group of arginine plays a crucial role by forming complexes with the phosphate groups of the membrane phospholipid bilayer, thereby enhancing the binding of the peptide to the bacterial membrane [22].
Hydrophobicity is another critical factor influencing AMP activity. In San-112003, tryptophan serves as the primary hydrophobic moiety [23]. Tryptophan is known for its significant role in enhancing peptide-membrane interactions, which is attributed to the specific affinity between its indole group and the carbonyl groups of phospholipids, leading to membrane disruption [24]. Additionally, the peptide's hydrophobicity is further enhanced by the presence of farnesyl, which disrupts membrane integrity and interferes with intracellular processes by modifying proteins and altering lipid composition [25]. The farnesyl group plays a pivotal mechanistic role in the antimicrobial action of San-112003 by enhancing its ability to anchor into lipid bilayers. This hydrophobic isoprenoid moiety promotes stronger interactions with the lipid-rich domains of bacterial membranes, much like lipid tails in clinically approved antimicrobial agents such as daptomycin. The farnesyl tail improves the peptide’s overall membrane affinity, facilitating insertion and pore formation or membrane disruption, which in turn may accelerate intracellular delivery of co-administered antibiotics like gentamicin.
Despite the high hydrophobicity introduced by tryptophan residues and the farnesyl chain, San-112003 exhibited minimal hemolytic activity toward human erythrocytes. This may be attributed to the balanced charge distribution of the peptide (net charge ~+3), which preferentially targets the negatively charged bacterial membranes over neutral mammalian membranes. Moreover, bacterial membranes are rich in phosphatidylglycerol and cardiolipin, which favor electrostatic binding of cationic peptides, whereas mammalian membranes contain cholesterol and zwitterionic lipids like phosphatidylcholine and sphingomyelin, reducing susceptibility to cationic AMP-induced disruption.
These properties together contribute to the selective toxicity of San-112003 and support its potential for therapeutic development with reduced risk of host cell lysis.
The hemolytic toxicity of San-112003 was evaluated, revealing negligible effects on human red blood cells. The cationic charge of approximately +3 contributed to minimizing hemolytic toxicity, likely due to the electrostatic attraction targeting bacterial cell membranes while sparing erythrocytes [26]. Previous studies suggest that erythrocytes are protected from lysis by cationic peptides due to the neutralizing effect of negatively charged sialic acid moieties on their surface, which is located approximately 80 Å from the cell membrane [27].
San-112003 exhibited potent antibacterial activity against both antibiotic-resistant and susceptible strains of P. aeruginosa. The minimum inhibitory concentration (MIC) of San-112003 was lower than that of several traditional antibiotics across various chemical classes, and the identical MIC and minimum bactericidal concentration (MBC) values indicate a bactericidal effect. The observed MIC value of 2.5 µg/ml for San-112003 against MDR P. aeruginosa falls within the clinically relevant range, as defined by the Clinical and Laboratory Standards Institute (CLSI) for aminoglycosides like gentamicin (≤4 µg/ml for susceptibility) [CLSI M100 guidelines]. This suggests that San-112003 could potentially achieve therapeutic levels in clinical use, pending pharmacokinetic and in vivo efficacy studies.
The results in fig. 5 show that the peptide alone had minimal impact on biofilm viability, but when combined with certain antibiotics—particularly rifampicin, clarithromycin, doxycycline, and chloramphenicol—a marked reduction in viable cells was observed, suggesting a synergistic effect. This supports previous findings that antimicrobial peptides (AMPs) enhance antibiotic penetration by disrupting bacterial membranes and partially destabilizing the biofilm matrix, thereby improving the intracellular access of antibiotics that target protein and RNA synthesis, while combinations with β-lactams and vancomycin were less effective due to the reduced metabolic activity of biofilm-embedded cells. Such results highlight the potential of AMPs as adjuvants in biofilm-related infections, where their ability to weaken biofilm integrity and interfere with quorum sensing can help overcome antibiotic tolerance and re-sensitize persistent bacteria to conventional drugs [28]. Compared to other well-studied USAMPs, San-112003 demonstrates competitive antimicrobial potency. For instance, WLBU2 exhibited MIC values ranging from 4–8 µg/ml against MDR P. aeruginosa, while DP7 showed MICs of 8–16 µg/ml in prior studies [1, 2]. Even colistin, one of the last-resort antibiotics, typically displays MIC values of 1–2 µg/ml against sensitive P. aeruginosa strains. San-112003's MICs (0.5–2.5 µg/ml) are comparable or lower, highlighting its potential as a next-generation antimicrobial.
Furthermore, the equivalence of MIC and MBC values in our findings supports the hypothesis that San-112003 exerts a bactericidal effect. Although not included in the current study, time-kill kinetics assays are recommended for future work to confirm rapid bacterial killing and further elucidate the mode of action.
In combination studies, San-112003 demonstrated additive effects when used with doxycycline, amoxicillin, levofloxacin, rifampicin, and cefixime against P. aeruginosa. Notably, a synergistic effect was observed when San-112003 was combined with gentamicin, suggesting that the peptide enhances gentamicin's intracellular entry by disrupting the bacterial membrane, thereby facilitating rapid target engagement [29]. The pronounced synergistic activity observed between San-112003 and gentamicin is likely attributable to enhanced membrane disruption caused by the peptide, which facilitates gentamicin’s intracellular uptake and access to ribosomal targets. Amino-glycosides are known to depend on energy-dependent transport systems that can be enhanced by prior membrane perturbation, making them more effective in combination with membrane-acting peptides [30].
In contrast, antibiotics like amoxicillin and cefixime, which target bacterial cell wall synthesis, showed additive or no synergistic effects. These antibiotics are often less influenced by membrane-permeabilizing agents, especially in Gram-negative bacteria, where outer membrane porins and β-lactamase activity dominate resistance mechanisms. Therefore, the interaction of San-112003 with these antibiotics may be limited by intrinsic resistance barriers that are not significantly overcome by peptide action.
The observed synergistic and additive effects highlight the potential of combining AMPs with conventional antibiotics. Such combinations could significantly reduce the required concentrations of AMPs to effectively kill P. aeruginosa while minimizing cytotoxicity. This approach offers a promising strategy for developing new antimicrobials, particularly in light of the global challenge of discovering safe and effective antibiotics. The formulation of AMPs like San-112003 with existing antibiotics could represent a critical advance in the fight against antibiotic-resistant P. aeruginosa.
CONCLUSION
In conclusion, San-112003 demonstrates significant potential as an antimicrobial agent against Pseudomonas aeruginosa, particularly multi-drug-resistant strains. Its ability to disrupt bacterial membranes, coupled with minimal hemolytic toxicity, underscores its therapeutic value. The observed synergistic effects with conventional antibiotics, especially gentamicin, suggest that combining AMPs with existing treatments could enhance efficacy and reduce the required dosages. This approach offers a promising avenue for developing new, effective antimicrobials in the face of rising antibiotic resistance.
DATA AVAILABILITY
Data availability Underlying data Fig share: Peptide raw data. https://doi.org/10.6084/m9.figshare.23895903.v1.28. This project contains the following underlying data: supplementary file 2. docx-micS. xlsx combination peptide. xlsx Data are available under the terms of the Creative Commons Attribution 4.0 International license (CC-BY 4.0).
ACKNOWLEDGEMENT
The author is grateful to the Middle East University (MEU), Amman, Jordan, for the financial support granted to cover the publication fee of this research article.
AUTHORS CONTRIBUTIONS
AHS: conceptualization and supervision. AHS and SMS: Investigation, methodology, and writing – review and editing. All authors have read, reviewed, and approved the final manuscript.
CONFLICT OF INTERESTS
The authors declare that they have no competing interests.
REFERENCES
Yin R, Cheng J, Wang J, Li P, Lin J. Treatment of Pseudomonas aeruginosa infectious biofilms: challenges and strategies. Front Microbiol. 2022;13:955286. doi: 10.3389/fmicb.2022.955286, PMID 36090087.
Dolan SK. Current knowledge and future directions in developing strategies to combat Pseudomonas aeruginosa infection. J Mol Biol. 2020;432(20):5509-28. doi: 10.1016/j.jmb.2020.07.021, PMID 32750389.
Langendonk RF, Neill DR, Fothergill JL. The building blocks of antimicrobial resistance in Pseudomonas aeruginosa: implications for current resistance-breaking therapies. Front Cell Infect Microbiol. 2021;11:665759. doi: 10.3389/fcimb.2021.665759, PMID 33937104.
Klubthawee N, Adisakwattana P, Hanpithakpong W, Somsri S, Aunpad R. A novel rationally designed hybrid antimicrobial peptide inspired by cathelicidin and aurein exhibits membrane active mechanisms against Pseudomonas aeruginosa. Sci Rep. 2020;10(1):9117. doi: 10.1038/s41598-020-65688-5, PMID 32499514.
Masihzadeh S, Amin M, Farshadzadeh Z. In vitro and in vivo antibiofilm activity of the synthetic antimicrobial peptide WLBU2 against multiple drug resistant Pseudomonas aeruginosa strains. BMC Microbiol. 2023;23(1):131. doi: 10.1186/s12866-023-02886-x, PMID 37183241.
Yin Q, Wu S, Wu L, Wang Z, Mu Y, Zhang R. A novel in silico antimicrobial peptide DP7 combats MDR Pseudomonas aeruginosa and related biofilm infections. J Antimicrob Chemother. 2020;75(11):3248-59. doi: 10.1093/jac/dkaa308, PMID 32737484.
Jahangiri A, Neshani A, Mirhosseini SA, Ghazvini K, Zare H, Sedighian H. Synergistic effect of two antimicrobial peptides nisin and P10 with conventional antibiotics against extensively drug-resistant acinetobacter baumannii and colistin-resistant Pseudomonas aeruginosa isolates. Microb Pathog. 2021;150:104700. doi: 10.1016/j.micpath.2020.104700, PMID 33346078.
Bussmann RW, Sharon D. Plantas medicinales de los Andes y la Amazonia la flora magica y medicinal del norte del peru. Ethnobot Res Appl. 2018;15(1):1-293. doi: 10.32859/era.15.1.001-293.
Grumaz C, Hoffmann A, Vainshtein Y, Kopp M, Grumaz S, Stevens P. Rapid next-generation sequencing-based diagnostics of bacteremia in septic patients. J Mol Diagn. 2020;22(3):405-18. doi: 10.1016/j.jmoldx.2019.12.006, PMID 32146977.
Almaaytah A, Qaoud MT, Abualhaijaa A, Al Balas Q, Alzoubi KH. Hybridization and antibiotic synergism as a tool for reducing the cytotoxicity of antimicrobial peptides. Infect Drug Resist. 2018 Jun 1;11:835-47. doi: 10.2147/IDR.S166236, PMID 29910626.
Birteksoz Tan AS, Zeybek Z, Hacioglu M, Savage PB, Bozkurt Guzel C. In vitro activities of antimicrobial peptides and ceragenins against Legionella pneumophila. J Antibiot (Tokyo). 2019;72(5):291-7. doi: 10.1038/s41429-019-0148-1, PMID 30755738.
Lee YC, Chen PY, Wang JT, Chang SC. A study on combination of daptomycin with selected antimicrobial agents: in vitro synergistic effect of MIC value of 1 mg/L against MRSA strains. BMC Pharmacol Toxicol. 2019;20(1):25. doi: 10.1186/s40360-019-0305-y, PMID 31060599.
Kowalska Krochmal B, Dudek Wicher R. The minimum inhibitory concentration of antibiotics: methods, interpretation clinical relevance. Pathogens. 2021;10(2):165. doi: 10.3390/pathogens10020165, PMID 33557078.
Patil SV, Hajare AL, Patankar M, Krishnaprasad K. In vitro fractional inhibitory concentration (FIC) study of cefixime and azithromycin fixed-dose combination (FDC) against respiratory clinical isolates. J Clin Diagn Res. 2015;9(2):DC13-5. doi: 10.7860/jcdr/2015/12092.5560, PMID 25859454.
Bidaud AL, Schwarz P, Herbreteau G, Dannaoui E. Techniques for the assessment of in vitro and in vivo antifungal combinations. J Fungi (Basel). 2021;7(2):113. doi: 10.3390/jof7020113, PMID 33557026.
Oh R, Lee MJ, Kim YO, Nam BH, Kong HJ, Kim JW. Myticusin beta antimicrobial peptide from the marine bivalve Mytilus coruscus. Fish Shellfish Immunol. 2020;99:342-52. doi: 10.1016/j.fsi.2020.02.020, PMID 32061872.
Yasir M, Dutta D, Willcox MD. Activity of antimicrobial peptides and ciprofloxacin against Pseudomonas aeruginosa biofilms. Molecules. 2020;25(17):3843. doi: 10.3390/molecules25173843, PMID 32847059.
Kim H, Jang JH, Kim SC, Cho JH. Development of a novel hybrid antimicrobial peptide for targeted killing of Pseudomonas aeruginosa. Eur J Med Chem. 2020;185:111814. doi: 10.1016/j.ejmech.2019.111814, PMID 31678742.
Martins AF, Rabinowitz P. The impact of antimicrobial resistance in the environment on public health. Future Microbiol. 2020;15(9):699-702. doi: 10.2217/fmb-2019-0331, PMID 32530297.
Xu SJ, Chen JH, Chang S, Li HL. The role of miRNAs in T helper cell development activation fate decisions and tumor immunity. Front Immunol. 2024;14:1320305. doi: 10.3389/fimmu.2023.1320305, PMID 38264670.
Zharkova MS, Orlov DS, Golubeva OY, Chakchir OB, Eliseev IE, Grinchuk TM. Application of antimicrobial peptides of the innate immune system in combination with conventional antibiotics a novel way to combat antibiotic resistance. Front Cell Infect Microbiol. 2019;9:128. doi: 10.3389/fcimb.2019.00128, PMID 31114762.
Han X, Kou Z, Jiang F, Sun X, Shang D. Interactions of designed Trp-containing antimicrobial peptides with DNA of multidrug-resistant Pseudomonas aeruginosa. DNA Cell Biol. 2021;40(2):414-24. doi: 10.1089/dna.2019.4874, PMID 32023094.
Wang K, Wang J, Li L, Xu L, Feng N, Wang. Synthesis of a novel anti-freezing non-drying antibacterial hydrogel dressing by one pot method. Chem Eng J. 2019;372:216-25. doi: 10.1016/j.cej.2019.04.107.
Aiemsaard J, Singh R, Borlace GN, Sripanidkulchai B, Tabboon P, Thongkham E. Antibacterial activity of cannabis extract (Cannabis sativa L. subsp. Indica (Lam.)) against canine skin infection bacterium Staphylococcus pseudintermedius. Scienceasia. 2022;48(3):348. doi: 10.2306/scienceasia1513-1874.2022.053.
Yasir M, Willcox MD, Dutta D. Action of antimicrobial peptides against bacterial biofilms. Materials (Basel). 2018;11(12):2468. doi: 10.3390/ma11122468, PMID 30563067.
Porto WF, Irazazabal L, Alves ES, Ribeiro SM, Matos CO, Pires AS. In silico optimization of a guava antimicrobial peptide enables combinatorial exploration for peptide design. Nat Commun. 2018;9(1):1490. doi: 10.1038/s41467-018-03746-3, PMID 29662055.
Patrulea V, Gan BH, Perron K, Cai X, Abdel Sayed P, Sublet E. Synergistic effects of antimicrobial peptide dendrimer-chitosan polymer conjugates against Pseudomonas aeruginosa. Carbohydr Polym. 2022;280:119025. doi: 10.1016/j.carbpol.2021.119025, PMID 35027127.
Huang F, Cai X, Hou X, Zhang Y, Liu J, Yang L. A dynamic covalent polymeric antimicrobial for conquering drug-resistant bacterial infection. Exploration (Beijing). 2022;2(5):20210145. doi: 10.1002/exp.20210145, PMID 37325499.
Parducho KR, Beadell B, Ybarra TK, Bush M, Escalera E, Trejos AT. The antimicrobial peptide human beta-defensin 2 inhibits biofilm production of Pseudomonas aeruginosa without compromising metabolic activity. Front Immunol. 2020;11:805. doi: 10.3389/fimmu.2020.00805, PMID 32457749.
Han X, Zheng J, Lin F, Kuroda K, Chen Z. Interactions between surface-immobilized antimicrobial peptides and model bacterial cell membranes. Langmuir. 2017;34(1):512-20. doi: 10.1021/acs.langmuir.7b03411, PMID 29232144.