
Int J App Pharm, Vol 18, Issue 2, 2026, 1-8Review Article
EARLY LIFE DETERMINANTS AND GENETIC MODIFIERS OF THE HUMAN GUT MICROBIOTA: IMPLICATIONS FOR DYSBIOSIS AND DISEASE
R. DHYANESH, NITHYA SHREE, JEEVANANTHAM I.*, KIRUBA SHALO ALBERT, SUVITHA SRI M., VISHVA PRASATH S.
Department of Pharmacy Practice, JSS College of Pharmacy, JSS Academy of Higher Education and Research, Ooty, Nilgiris-643001, Tamil Nadu, India
*Corresponding author: Jeevanantham I.; *Email: jeevaimayavam@gmail.com
Received: 30 Aug 2025, Revised and Accepted: 17 Dec 2025
ABSTRACT
The gut microbiome comprises trillions of bacteria, viruses, fungi, and archaea. The genes of these microorganisms, collectively referred to as the gut microbiota, are crucial for digestion, xenobiotic metabolism, and the regulation of both innate and adaptive immune responses. This relationship is maintained through a continuous biochemical exchange of proteins, peptides, and metabolites with the host. However, factors such as ageing, chronic stress, poor diet, antibiotics, and underlying illnesses can disrupt this essential balance, leading to gut dysbiosis. Dysbiosis is characterised by reduced diversity and alterations in the abundance of key bacterial taxa. It is not only linked to local digestive symptoms like bloating, diarrhoea, and constipation but also to systemic conditions such as fatigue, immunological imbalance, and metabolic abnormalities. Dysbiosis is now closely associated with the aetiology of inflammatory bowel disease (IBD), irritable bowel syndrome (IBS), obesity, diabetes, cancer, cardiovascular diseases, and neurological disorders. Systemic approaches to address dysbiosis include probiotics, prebiotics, dietary modification, and intestinal microbiome transplantation (IMT), which is particularly helpful for recurrent Clostridium difficile infections.
Keywords: Gut microbiome, Newborn, Bacteria, Microbes, Gut-skin axis, Gut-brain axis
© 2026 The Authors.Published by Innovare Academic Sciences Pvt Ltd. This is an open access article under the CC BY license (https://creativecommons.org/licenses/by/4.0/)
DOI: https://dx.doi.org/10.22159/ijap.2026v18i2.56700 Journal homepage: https://innovareacademics.in/journals/index.php/ijap
INTRODUCTION
The aggregate genomes of bacteria, viruses, protozoa, fungi, and archaea that coexist in various habitats within the human body are referred to as human microbiomes [1, 2]. Up to 3.0 ×10^13 bacteria, or total cellular count in the human body, may be present in a 70 kg adult [3]. Numerous characteristics, including wall type, shape, oxygen demand (anaerobic or aerobic), endospore generation, motility, and metabolism, are used to classify bacteria both morphologically and biochemically. Bacteria are also classified according to the evolutionary richness of different nucleotide sequences of small subunit ribosomal RNA operons, or 16S and 18S RNA genes [1, 2]. The human microbiome project (HMP) has been setting standards for comprehensive and high-quality metagenomic profiling extracted directly from various locations of the body in order to establish the microbial relative abundance of numerous strains and species of various phyla under normal conditions [4-6].
Improvements in computer technologies have enabled many research studies regarding the inflammatory mediators, which are genome-based on the evolutionary clustering and structure of bacterial genomes into taxonomical domains, kingdom, phylum, class, order, family, genus, and species. [7, 8]. Numerous human microbiome datasets were analysed, and the results showed the enormous variety at the individual and population levels during life and evolution [9]. Both permanent and transitory microbial species, along with subspecies of over 17 putative bacterial phyla belonging to the Firmicutes, comprise over 70% of the gut microbiota of healthy individuals. Proteobacteria (<5%), Actinobacteria (<2%), Fusobacteria and Verrucomicrobiome (<1%), Bacteroidetes (>30%), and other phyla. Using 1550 metagenome-assembled genomes (MAGs) for taxonomic profiling and novel bacterial genome assembly, almost 70,000 bacterial and archaeal genomes, as well as new species, are being thoroughly studied [10]. The most common organisms found in human stools include Clostridium species, such as Coprococcus, Ruminococcus, Eubacterium, Bacteroides and fragilis, and Alistipes fine goldii and onderdonkii [11]. The ability to cluster species-level phylotypes into major human enterotypes has been made possible by the bacterial species' extremely broad taxonomic range, microbial history, and metabolism with regard to caloric load and nutritional absorption. High Bacteroides and low Bacteroides but high Prevotella levels are characteristics of the typical human enterotype type 1. They are linked to those who eat a lot of animal protein or carbohydrate type 2 [12].
The enteric nervous system is a unique neurological system found in the gastrointestinal (GI) tract. Through the vagus nerve, neuromodulators and neurotransmitters of the sympathetic and parasympathetic branches of the autonomic nervous system interact with the central nervous system [13]. The GI microbiome bacterial diversity and richness are essential to the tissues' and organs' regular immunological and metabolic processes [14]. Here, we will have an insight into research on the various ways in which immune cells function, how inflammatory disorders develop, and what is the impact caused by microbiomes on metabolic diseases. It is proven that even small compounds, probiotics, prebiotics, and faecal transplantation may aid in the treatment and modulation of different strains, phylum types and encourage the restoration of microbiomes that cause metabolic disorders. One such prebiotic is lactic acid bacteria (LAB). LAB produce lactic acid as the major end-product during the fermentation of carbohydrates. These are abundant in nature. The microorganism is found in milk, meat, green plants, grains and fermenting vegetables. LAB has been isolated from the mucosal surface of animals, sourdoughs, vacuum-packaged refrigerated beef and traditional Indian fermented foods such as appam batter and vegetable pickle. LAB includes a diverse genus of organisms; those are Lactobacillus, Pediococcus, Lactococcus, Carnobacterium, Enterococcus, Lactosphaera, Leuconostoc, Melissococcus, Oenococcus, Streptococcus, Tetragenococcus, Vagococcus, Weissella, and Bifidobacterium. Some of the ideal characteristics of a probiotic bacterium are illustrated below.
Factors influencing the gut microbiome's development in early childhood
Parturition
A small percentage of species that make it to the gastrointestinal tract can colonise, and gut microbiome colonisation varies greatly in the early stages of infancy [15]. The gut microbiome is a topic of much debate. On the other hand, there is proof that organisms that are present in the infant's bowel during mid-pregnancy [17] may start colonising the GIT for the first time as early as in utero [16]. Around the age of three, the gut microbiome develops into its mature, stable form [18]. It was believed till recent times that intestinal colonisation did not start until delivery, when the baby was first exposed to the mother's faecal and vaginal microbes, and that the foetus formed in a sterile environment. Nevertheless, microbes have recently been found in the meconium [22], placenta [19], amniotic fluid [20], and umbilical cord [21]. Despite these results, it is still debatable whether the published study adequately controlled for contamination. Though the significant impact of such early colonisation is still unknown, the growing body of evidence has led to the universal acceptance that the gut microbiome starts to seed in pregnancy.
An infant's intestinal microbiome is extremely vulnerable during this period, and environmental exposures can readily affect colonisation patterns [23]. In order to promote normal development and better clinical outcomes into adulthood, the ontogeny of the gut microbiome at this time is crucial for increased flexibility, which is of great interest [24]. The infant's initial gut colonisation is greatly influenced by the birth method (vaginal or caesarean). The next significant transitional periods occur postnatally, around the start of breastfeeding and the end of formula or breast milk feeding.
The primary source of an infant's gut microbe colonisation in the immediate postpartum period is the mother's vaginal and faecal microbes. The newborn’s gut microbiome changes quickly after parturition. Bare skin contact can transmit organisms commonly found on the body of newborns [26], but the main cause is the delivery of breast milk. A newborn’s first milk, colostrum, is thought to be essential for the early growth of the gut microbiota due to its high concentration of antigen-specific and non-antigen-specific antimicrobials, such as secretory IgA and lactoferrin. human milk oligosaccharides (HMOs) are a particular type of food for bacteria and are also found in large quantities in breast milk [27]. Bifidobacterium, which are well adapted to ingest HMOs, dominate the gut microbiome during the early phase of life, when most of the babies are exclusively fed milk [28]. The gut microbiome composition becomes more "adult-like" when solid meals are introduced, and a milk-based diet is stopped [25, 29]. An adult's gut microbiome is thought to be extremely stable with very slight variations unless there are significant dietary changes, antibiotic use, or medical conditions [30, 31].
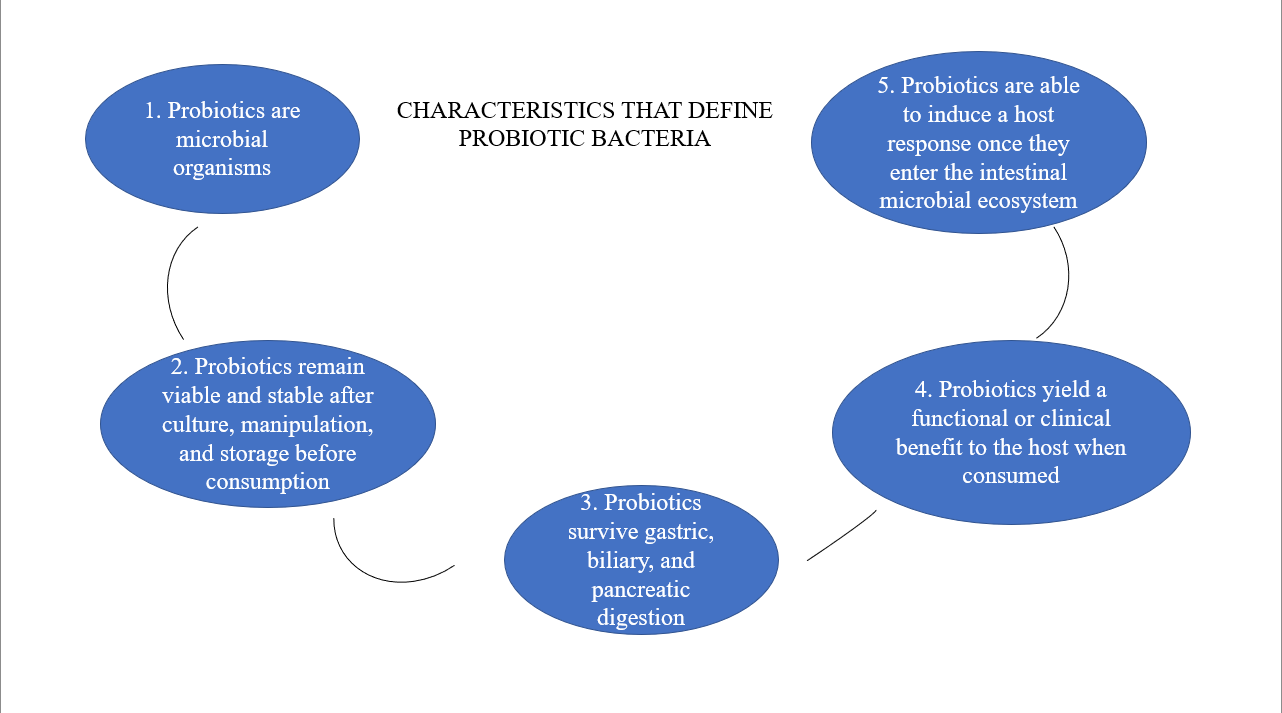
Fig. 1: The ideal characteristics of a probiotic bacterium
Early postnatal life
There was a longitudinal study conducted on 39 Finnish infants between the ages of 2 and 36 mo, collecting stool samples every month to shed light on the early infant gut microbiome's development [32]. Significant levels (average abundance) of the Enterobacteriaceae (25%), Bifidobacteriaceae (15%), and Clostridiaceae (8%) families were discovered in the vast majority of newborns in their study. By the time they were 18 mo old, the relative average abundances of these families had dropped to 1%, 3%, and 2.5%, respectively [32]. To break down the starch for a more complex diet, the gut microbes undergo a change to a predominant overflow of species from the Bacteroides genus once solid food is introduced [33]. The gut microbiome of individuals aged 18 to 65 and children under 3 has been the primary focus of large population investigations. As a result, little is known about kindergarten and early childhood [34]. Children taking over by Bifidobacterium showed fewer overall spectrum and bacterial gene counts than children overshadowed by Bacteroides, or Prevotella, with a similar survey was carried out on 281 early school-aged children [35]. The gut microbiomes of school-age children have high levels of Bifidobacterium, which is similar to what is found in adults. Remarkably, they also discovered that the group that was dominated by Bifidobacterium had a shorter duration of breastfeeding overall [35].
In this survey, they found that children dominated by Bifidobacterium had lower bacterial gene counts and overall diversities than children dominated by Bacteroides or Prevotella [35]. School-age children's gut microbiota has more significant amounts of Bifidobacterium, which is comparable to adult levels. Surprisingly, it was also found that the Bifidobacterium-dominated group breastfed for a small period of time overall [35]. A comprehensive review to close the knowledge gap on school-age children's gut microbiomes. In the preadolescent gut, they found that Bacteroidetes and Firmicutes predominate [36].
Genetics
According to recent research, host genetics can affect the human gut microbiome's composition, which may impact the host metabolism. Interactions between the carrier immune system and the microbiota are linked to the development of the newborn gut microbiome. Significant genome-wide relationships for total microbial variation have been discovered by a number of investigations, including genome-wide association studies [37-39]. The single-nucleotide polymorphism (SNP) heritability of 1475 Chinese individuals was examined, where they discovered that the heritability estimates of Desulfovibrionaceae and Odoribacter were considerable, at 0.456 and 0.476, respectively [40]. Individual genetic variation can affect the composition of the microbiome, and twin studies allow us to evaluate the relative contributions of environment and genes. A twin study was once conducted to evaluate the genetics of the gut microbiome; monozygotic twins have greater gut microbiome similarities than dizygotic twins, with operational taxonomic unit (OTU) hereditary tendency ranging from 0.2 to 0.4 [42].
To categorise closely related species, OTUs-clusters of comparable sequence variants-are utilised. Additionally, they discovered that host genetics affected the abundances of numerous microbial taxa, with the Christensenellaceae family being the most heritable [42].
Even among healthy people, the gut flora is highly individualised as a result of extensive research [43]. Genetic heterogeneity between human populations and geographical areas is well documented. It has also been demonstrated that different human populations have different gut bacterial communities [15]. In a recent study, faeces samples were collected and analysed from more than 2,000 adults from Amsterdam's six biggest ethnic groups using 16S sequencing. They discovered that ethnic differences from ethnic groups residing in the same geographic area explained a higher percentage of the variation in gut microbiome than other important factors, like food [44]. These results imply that gastrointestinal colonization patterns are influenced by environmental factors, dietary patterns, and host-intrinsic genetic diversity where they concluded that 21 microbial groups were shared by all 2084 individual across all ethnicity [44], suggesting that individuals from the same ethnic group tend to share more gut microbiome compositions than those from different [45], even though the composition of gut microbiomes varies among ethnic groups. This lends credence to the idea that certain species are common throughout populations and vital to human health, even though gut microbiome compositions vary greatly between individuals. Understanding the variables, such as genetics, that affect the composition and development of the gut microbiome is necessary to appreciate the role that the microbiome plays in the course of disease.
Infant feeding
Breastfeeding has a major impact on the early gut microbiome’s composition. Breastfed babies often have gut flora rich in Lactobacillus, Staphylococcus, and Bifidobacterium, particularly species like Bifidobacterium breve, Bifidobacterium bifidum, and Bifidobacterium longum, which are highly specialised at breaking down HMOs. Formula-fed babies, however, tend to have gut flora like Roseburia, Clostridium, and Anaerostipes.
Solid food initiation
When solid foods are initiated during the weaning phase, the infant's gut microbiome undergoes its next significant shift. The initiation of solid food causes the composition of the microbes to shift from being dominated by Bifidobacterium to being dominated by species of Firmicutes and Bacteroides [53]. A shift from bacterial genes used for lactate digestion to genes more suitable for carbohydrate digestion is made possible by this shift in species abundance [18]. These alterations continue until the child is three years old, and they help the transition to a more varied and adult-like gut microbiota.
Except for prolonged dietary changes, disease-induced dysbiosis, or antibiotic exposure, the gut microbiome is largely stable after this. It's interesting to note that early solid food introduction (before three months of age) has been connected to immune system issues, oxidative stress, and childhood obesity. By increasing butyrate concentrations and diversity, it has also been connected to alterations in the infant's gut microbiota composition [54].
Delivery method
One of the most significant factors influencing the formation of a newborn’s gut microbiome is the species that first colonises their gut microbiome; this is significantly influenced by the delivery method (vaginal or caesarean) [55]. By altering the type and reducing the diversity of the early colonising bacteria, cesarean births have been shown to have a detrimental effect on the development of the baby's gut microbiome and, subsequently, immune system. Among the immune and allergic disorders for which caesarean section is more likely to occur are asthma, arthritis, IBD, and immunological deficiencies [56]. Perhaps, it is understood that C-section can worsen the Vaginal seeding, which has begun to be practised in some hospitals, is one of the methods studied to reinstate the newborn’s gut microbiome after C-section [57]. Infant delivery is the time of first contact for young newborns with the multitude of microorganisms in the environment, a few of which are the bacteria that colonise the mother’s vaginal canal. A C-section delivery may be necessary in some situations, and the baby gets a new set of microbes. The World Health Organisation (WHO) suggests that the percentage of caesarean deliveries stay below 15%. Nonetheless, the number of caesarean deliveries has been rising in wealthy nations. Between 1990 and 2014, the percentage of caesarean deliveries increased from 6.7% to 19.1%, according to data from 150 countries [58]. While vaginal deliveries are linked to gut microbiome signatures in babies that are similar to the mother's vaginal microbiome, as identified by early stool samples, caesarean births are associated with species that more closely match skin taxa [59]. During the first week following delivery, caesarean newborns typically have higher concentrations of skin microbiome-related microorganisms, such as Staphylococcus, Streptococcus, and Propionibacterium, and lower levels of Bacteroides, Lactobacillus, and Bifidobacterium [60]. After the first week of life, caesarean section babies have decreased levels of Bifidobacterium and greater levels of Klebsiella, Haemophilus, and Veillonella[61]. Differences in delivery methods no longer appear to be significant, despite the fact that small variances in Lactobacillus, Bacteroides, and Bifidobacterium can still be found beyond the first month of life [62]. By six months of age, the colonisation patterns of babies born vaginally and those born via caesarean section are almost the same. Nonetheless, infants born vaginally continue to have higher levels of Bacteroides and Parabacteroides, while infants born via caesarean section have higher levels of Clostridium species [55].
The aseptic environment of the operating theatre can encourage colonisation of flora, which is associated with a higher risk of respiratory diseases [63]. Caesarean deliveries also frequently result in delayed interaction with mothers and delayed breastfeeding initiation. Antibiotics are routinely given to the mother during delivery (via drip) in many hospitals, which exposes the newborn to antibiotics during birth. According to a 2018 pilot study, the use of maternal antibiotics during delivery had a greater effect on the newborn’s gut microbiome than the manner of delivery, notably lowering Bifidobacterium colonisation [64].
It's interesting to note that the gut microbiome profiles of infants are different after elective and emergency caesarean sections. The gut microbiota profiles of caesarean sections performed before the commencement of labour are really more closely linked to vaginal births than to elective caesarean sections. Infants who have elective surgery shield themselves from the inflammatory cytokines that are released into the uterus as a result of the immunological response during delivery. These cytokines have been associated with the development of the infant's immune system [65].
Use of antibiotics
Antibiotic use is linked to immediate and long-term adverse health effects, including a higher risk of asthma and autoimmune illnesses. Antibiotics cause alterations in the quantity and diversity of some gut microbial species, notably Bifidobacterium, leading to a loss in resistance to opportunistic infections and an increase in antibiotic resistance. The effects vary depending on the antibiotic class, mode of ingestion, dosage, duration, and range of activity. Understanding how antibiotics impact the gut microbiome's makeup can help minimise the harm that antibiotic treatment causes by customising probiotic use and antibiotic treatment, which is already being done in adults. More advanced sequencing methods are required to analyse antibiotic resistance genes in the gut microbiome to better inform antibiotic therapy, particularly in early life when the gut microbiome is more susceptible to the effects of antibiotics. Currently, taxa within the gut microbiome can be identified using sequencing data.
Miscellaneous factors
Early contact with siblings or pets can help prevent allergic illness, according to a number of observational studies [75-77]. This is occasionally coupled with the "hygiene hypothesis," which maintains that limiting early exposure to a range of bacteria might not be the greatest strategy for immune development. Young children's gut microbiota is believed to be more diverse when they interact with siblings and household pets, which is known to prevent atopy [76]. Ruminococcin and Oscillopsia, which are linked to childhood obesity and atopy, were shown to be more prevalent in infants exposed to dogs both during pregnancy and after delivery [78]. Therefore, there is controversy around the body of research on how domestic furry pets affect the intestinal microbiome of newborns. These investigations, however, provide credence to the idea that the gut microbiome of infants is extremely malleable.
Very preterm newborn death rates have significantly decreased in recent decades. Nevertheless, there is no correlation between these lower morbidity and mortality rates. Along with their immature immune response, preterm altered gut microbiome causes both pro-and counter-inflammatory reactions [79]. Staphylococcus and Enterobacteriaceae are more common in preterm newborns, while Bifidobacterium colonisation is often delayed [80]. Pro-inflammatory responses in preterm newborns are linked have higher levels of Enterobacter, Enterococcus, and Lactobacillus [81]. Preterm neonates (born before 7 mo) had higher levels of Lactobacillus in their meconium, which predominates in the mother's vaginal microbiome, than extremely preterm neonates (delivered after 7 mo), regardless of the type of delivery [82]. This lends credence to the idea that the maternal microbiome affects the GI tract's first seeding. In addition to having distinct dietary needs and a higher likelihood of being admitted to intensive care units, preterm children are also more likely to be exposed to the aseptic conditions of the hospital and, in certain situations, antibiotics [83, 84]. Preterm babies born with a very low birth weight (VLBW) are prone to experience intestinal microbial dysbiosis, which is defined by a low diversity of the gut microbiome, a decrease in helpful microorganisms, and an increase in opportunistic pathogens [85]. It has been hypothesised to be brought on by the disruption of intrauterine development and may be connected to increased intestinal inflammation. VLBW newborns are often taken from their moms and placed in incubators to help them control their body temperature.
Signs and symptoms
Digestive symptoms
Gut dysbiosis is the condition of having an imbalance of the bacterial composition of the gut, where harmful or dangerous bacteria outnumber those that are beneficial. Disrupting the normal digestive process. Symptoms such as bloating, diarrhoea, constipation, and flatulence can occur when there is disruption of the body's natural digestive cycle. When pathogenic microflora ferments partially digested food, it can lead to gas and bloating in the body [86]. Additionally, dysbiosis may affect the gut's capacity to absorb nutrients and control bowel movements, leading to constipation or diarrhoea. The changed microbial habitat also probably triggers inflammatory reactions in the gut lining, aggravating digestive problems and creating a vicious circle of discomfort [87, 88].
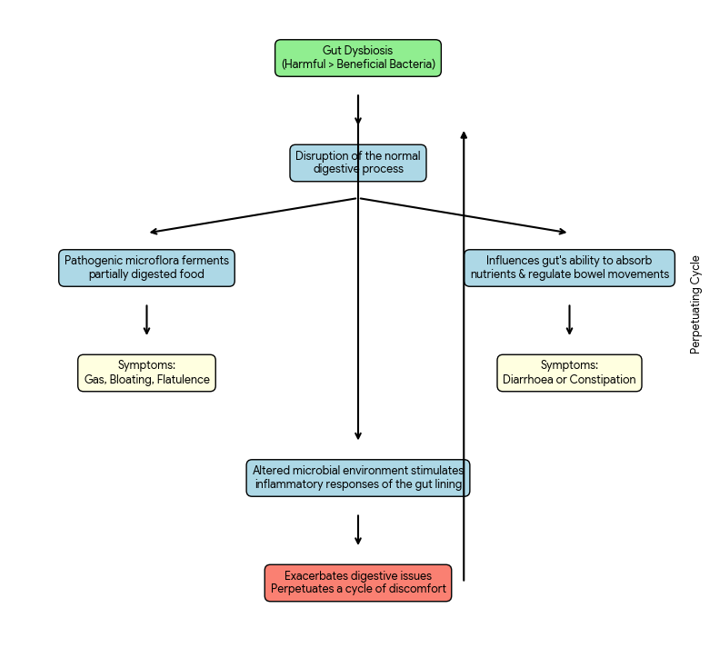
Fig. 3: Shows how the digestive system is affected due to gut imbalance
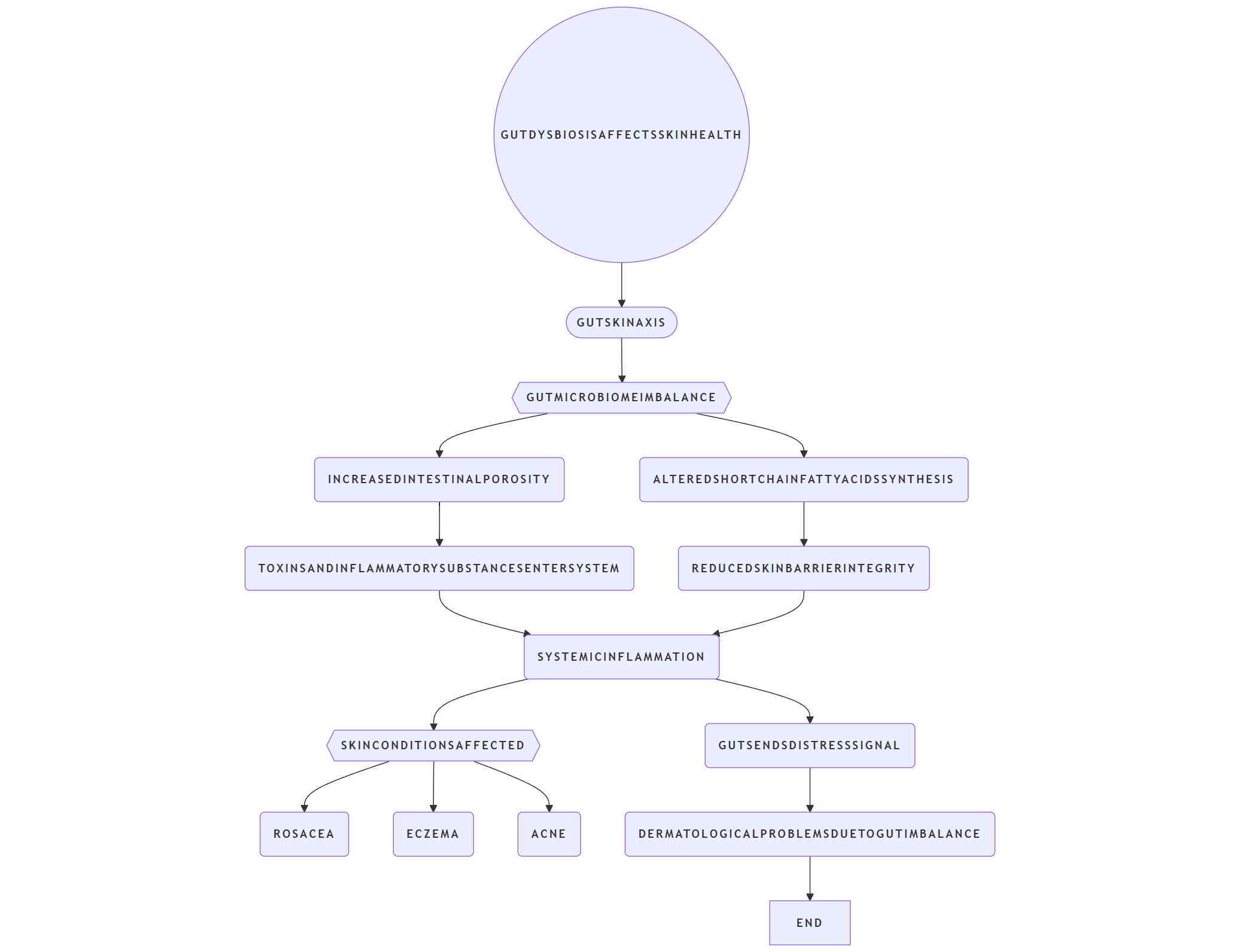
Fig. 4: How gut dysbiosis can lead to skin problems
Skin issues
Gut dysbiosis and implications on skin health via the gut–skin axis Gut dysbiosis can affect skin health through the gut-skin axis, when an imbalance in the gut microbiome influences inflammation and immunity [89]. Increased intestinal permeability brought on by the expropriation of harmful microbes allows toxins and inflammatory chemicals to enter the human body. Skin conditions like rosacea, eczema and acne can be caused by or exacerbated by this systemic inflammation. Dysbiosis can also alter the synthesis of short-chain fatty acids (SCFA), which are important for maintaining skin barrier integrity and levels of inflammation. In essence, the body receives a distress signal from the injured gut. [90]. It also includes skin, resulting in several dermatological problems caused due to gut imbalance.
Fatigue and low energy
An unhealthy gut microbiome can have a direct effect on energy levels, which can cause chronic fatigue and brain fog in rare cases.
By interfering with the synthesis of neurotransmitters and the absorption of minerals, dysbiosis can lead to exhaustion and mental fog. An unbalanced gut flora may impair the generation of B vitamins and amino acids, which are essential for the production of energy [93, 94].
Food sensitivities and intolerances
The digestive system’s ability to digest certain foods, such as lactose, is impaired when the gut is overrun by unhelpful bacteria, leading to intolerances [95]. Furthermore, dysbiosis may be linked to food sensitivities and increased intestinal permeability, which permits undigested food particles to enter the bloodstream and trigger immunological reactions. Digestive enzymes may now also not be synthesised in balance, then things like foods may cause even more chaotic responses than now, divorced from the input [96].
Immune system imbalance
Immunity can indeed be influenced by an imbalanced gut microbiome. A critical region for immune function is the Gut-Associated Lymphoid Tissue (GALT). A disturbed microbiome can lead to an oversensitive or ambivalent immune response. The immune system may fail to adequately defend the body against infection or incorrectly attack cells in the body (autoimmunity) as a result of this imbalance. Increased permeability of the intestine that is induced by dysbiosis could ultimately allow toxins to access the circulation and inflammation [97].
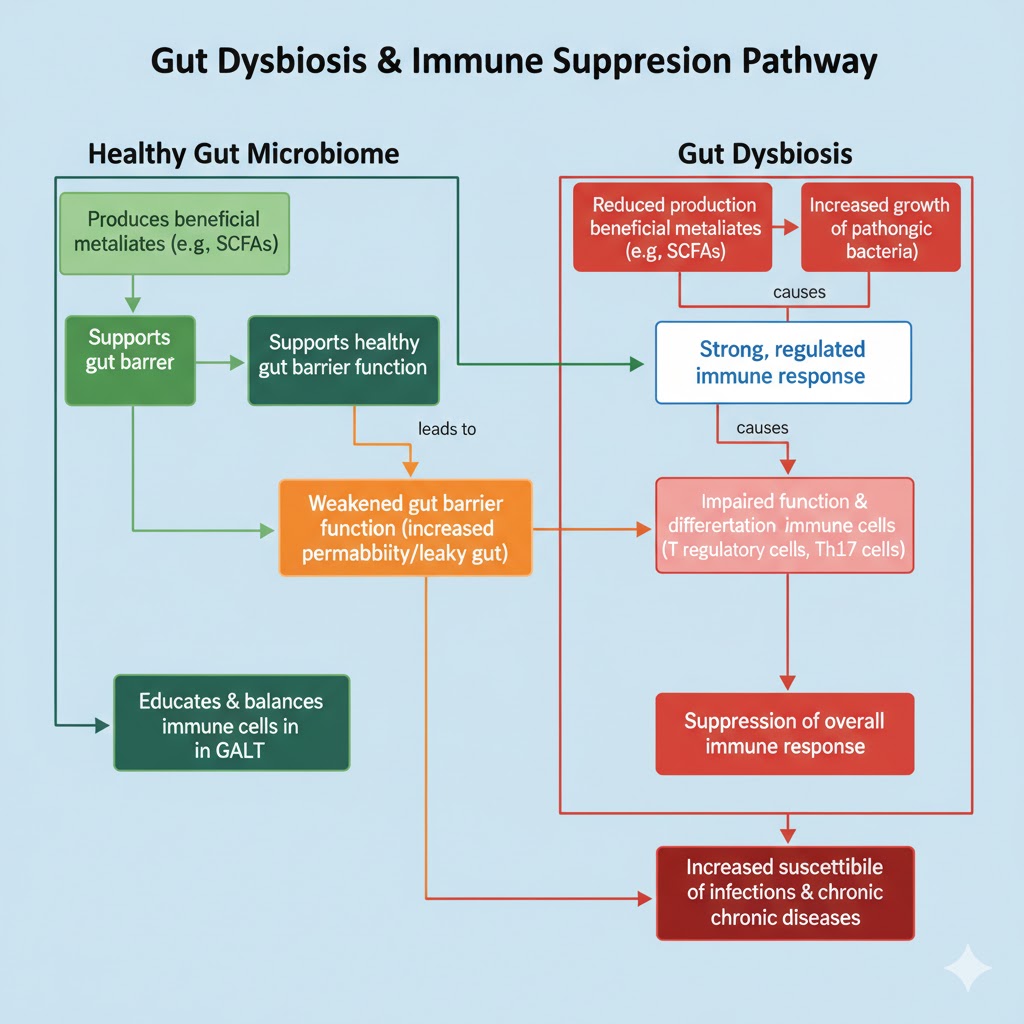
Fig. 5: Flowchart explaining immune intolerance against gut disease
Mental health symptoms
Through the gut-brain axis, this illness may have significant impacts on both the immune system and mental health. By influencing the production of neurotransmitters like serotonin, which is essential for mood regulation, an imbalance in the composition of the microbiome can cause anxiety and sadness. Dysbiosis can also increase inflammation, which may alter brain function and result in mood swings and cognitive issues [98, 99]. The gut flora also creates SCFAs, which are neuroprotective. These imbalances can affect emotional stability and mental alterations.
Weight changes
Unexpected fluctuations in weight can occur when intestinal dysbiosis interferes with metabolism. Both how the body uses food to produce energy and how nutrients are absorbed and maintained are impacted by an unbalanced microbiota. Certain gut bacteria digest dietary fibres to create SCFAs, which regulate the appetite and fat storage. Since dysbiosis reduces the synthesis of SCFAs, it may increase appetite and cause weight gain. It may also lead to inflammation, which can exacerbate insulin resistance and make controlling weight more difficult. It has a major impact on the gut-brain axis, affecting hormones like ghrelin and leptin that control hunger and fullness. In essence, an unhealthy stomach results in a biochemical imbalance that produces weight fluctuations, much like a broken thermostat.
Poor sleep
The gut microbiome is involved in producing neurotransmitters like melatonin, which regulates the sleep cycle. Dysbiosis can disrupt these pathways, leading to insomnia or poor sleep quality.
CONCLUSION
To conclude, gut dysbiosis is a significant disruption to the composition and activity of the intestinal microbiome that has widespread effects on overall health. It is now closely linked to a range of immunological, neurobehavioral, metabolic, and gastrointestinal disorders. Targeted intervention is made possible by an understanding of the major causes of dysbiosis, including genetics, delivery mode, newborn nutrition, and antibiotic use. The need for a holistic strategy is highlighted by the mechanistic understanding of dysbiosis, specifically its impacts through the gut-brain and gut-skin axis. To treat dysbiosis and lessen its long-term effects on human health, more research and a customised approach to gut health are crucial.
FUNDING
The authors have no information on funding resources to declare.
AUTHORS CONTRIBUTIONS
Conceptualisation: JI, MIA. Literature’s collection and writing: JBJ, PK Manuscript design and final draft preparation: SMO, DR.
All the authors have read and approved the final manuscript.
CONFLICT OF INTERESTS
The authors declare no competing interests.
REFERENCES
Backhed F, Fraser CM, Ringel Y, Sanders ME, Sartor RB, Sherman PM. Defining a healthy human gut microbiome: current concepts future directions and clinical applications. Cell Host Microbe. 2012;12(5):611-22. doi: 10.1016/j.chom.2012.10.012, PMID 23159051.
Belizario JE, Napolitano M. Human microbiomes and their roles in dysbiosis common diseases and novel therapeutic approaches. Front Microbiol. 2015;6:1050. doi: 10.3389/fmicb.2015.01050, PMID 26500616.
Sender R, Fuchs S, Milo R. Revised estimates for the number of human and bacteria cells in the body. PLOS Biol. 2016;14(8):e1002533. doi: 10.1371/journal.pbio.1002533, PMID 27541692.
Qin J, Li R, Raes J, Arumugam M, Burgdorf KS, Manichanh C. A human gut microbial gene catalogue established by metagenomic sequencing. Nature. 2010;464(7285):59-65. doi: 10.1038/nature08821, PMID 20203603.
Li J, Jia H, Cai X, Zhong H, Feng Q, Sunagawa S. An integrated catalog of reference genes in the human gut microbiome. Nat Biotechnol. 2014;32(8):834-41. doi: 10.1038/nbt.2942, PMID 24997786.
Lloyd Price J, Abu-Ali G, Huttenhower C. The healthy human microbiome. Genome Med. 2016;8(1):51. doi: 10.1186/s13073-016-0307-y, PMID 27122046.
Lloyd Price J, Mahurkar A, Rahnavard G, Crabtree J, Orvis J, Hall AB. Strains functions and dynamics in the expanded human microbiome project. Nature. 2017;550(7674):61-6. doi: 10.1038/nature23889, PMID 28953883.
Parks DH, Rinke C, Chuvochina M, Chaumeil PA, Woodcroft BJ, Evans PN. Recovery of nearly 8,000 metagenome-assembled genomes substantially expands the tree of life. Nat Microbiol. 2017;2(11):1533-42. doi: 10.1038/s41564-017-0012-7, PMID 28894102.
Arumugam M, Raes J, Pelletier E, Le Paslier D, Yamada T, Mende DR. Enterotypes of the human gut microbiome. Nature. 2011;473(7346):174-80. doi: 10.1038/nature09944, PMID 21508958.
Koren O, Knights D, Gonzalez A, Waldron L, Segata N, Knight R. A guide to enterotypes across the human body: meta-analysis of microbial community structures in human microbiome datasets. PLOS Comput Biol. 2013;9(1):e1002863. doi: 10.1371/journal.pcbi.1002863, PMID 23326225.
Le Chatelier E, Nielsen T, Qin J, Prifti E, Hildebrand F, Falony G. Richness of human gut microbiome correlates with metabolic markers. Nature. 2013;500(7464):541-6. doi: 10.1038/nature12506, PMID 23985870.
Moeller AH, Li Y, Mpoudi Ngole E, Ahuka Mundeke S, Lonsdorf EV, Pusey AE. Rapid changes in the gut microbiome during human evolution. Proc Natl Acad Sci USA. 2014;111(46):16431-5. doi: 10.1073/pnas.1419136111, PMID 25368157.
Qin J, Li R, Raes J, Arumugam M, Burgdorf KS, Manichanh C. A human gut microbial gene catalogue established by metagenomic sequencing. Nature. 2010;464(7285):59-65. doi: 10.1038/nature08821, PMID 20203603.
Hooper LV, Gordon JI. Commensal host-bacterial relationships in the gut. Science. 2001;292(5519):1115-8. doi: 10.1126/science.1058709, PMID 11352068.
Laforest Lapointe I, Arrieta MC. Patterns of early-life gut microbial colonization during human immune development: an ecological perspective. Front Immunol. 2017;8:788. doi: 10.3389/fimmu.2017.00788, PMID 28740492.
Adamek K, Skonieczna Zydecka K, Węgrzyn D, Loniewska B. Prenatal and early childhood development of gut microbiota. Eur Rev Med Pharmacol Sci. 2019;23(21):9667-80. doi: 10.26355/eurrev_201911_19461, PMID 31773718.
Wang JZ, Du WT, Xu YL, Cheng SZ, Liu ZJ. Gut microbiome-based medical methodologies for early-stage disease prevention. Microb Pathog. 2017;105:122-30. doi: 10.1016/j.micpath.2017.02.024, PMID 28219830.
Tang ML, Mullins RJ. Food allergy: is prevalence increasing? Intern Med J. 2017;47(3):256-61. doi: 10.1111/imj.13362, PMID 28260260.
Dharmage SC, Perret JL, Custovic A. Epidemiology of asthma in children and adults. Front Pediatr. 2019;7:246. doi: 10.3389/fped.2019.00246, PMID 31275909.
Fang X, Henao Mejia J, Henrickson SE. Obesity and immune status in children. Curr Opin Pediatr. 2020;32(6):805-15. doi: 10.1097/MOP.0000000000000953, PMID 33105275.
Cook M, Douglass J, Mallon D, Smith J, Wong M, Mullins R. Economic impact of allergies. Access Econ. 2007. Available from: https://www.allergy.org.au/images/stories/pospapers/2007_economic_impact_allergies_report_13nov.pdf.
Renz H, Skevaki C. Early life microbial exposures and allergy risks: opportunities for prevention. Nat Rev Immunol. 2021;21(3):177-91. doi: 10.1038/s41577-020-00420-y, PMID 32918062.
Lambrecht BN, Hammad H. The immunology of the allergy epidemic and the hygiene hypothesis. Nat Immunol. 2017;18(10):1076-83. doi: 10.1038/ni.3829, PMID 28926539.
De Filippo C, Cavalieri D, Di Paola M, Ramazzotti M, Poullet JB, Massart S. Impact of diet in shaping gut microbiota revealed by a comparative study in children from Europe and rural Africa. Proc Natl Acad Sci USA. 2010;107(33):14691-6. doi: 10.1073/pnas.1005963107, PMID 20679230.
Spor A, Koren O, Ley R. Unravelling the effects of the environment and host genotype on the gut microbiome. Nat Rev Microbiol. 2011;9(4):279-90. doi: 10.1038/nrmicro2540, PMID 21407244.
Rodriguez JM, Murphy K, Stanton C, Ross RP, Kober OI, Juge N. The composition of the gut microbiota throughout life with an emphasis on early life. Microb Ecol Health Dis. 2015;26:26050. doi: 10.3402/mehd.v26.26050, PMID 25651996.
Yatsunenko T, Rey FE, Manary MJ, Trehan I, Dominguez Bello MG, Contreras M. Human gut microbiome viewed across age and geography. Nature. 2012;486(7402):222-7. doi: 10.1038/nature11053, PMID 22699611.
Perez Munoz ME, Arrieta MC, Ramer Tait AE, Walter J. A critical assessment of the “sterile womb” and in utero colonization” hypotheses: implications for research on the pioneer infant microbiome. Microbiome. 2017;5(1):48. doi: 10.1186/s40168-017-0268-4, PMID 28454555.
Mishra A, Lai GC, Yao LJ, Aung TT, Shental N, Rotter Maskowitz A. Microbial exposure during early human development primes fetal immune cells. Cell. 2021;184(13):3394-3409.e20. doi: 10.1016/j.cell.2021.04.039, PMID 34077752.
Tanaka M, Nakayama J. Development of the gut microbiota in infancy and its impact on health in later life. Allergol Int. 2017;66(4):515-22. doi: 10.1016/j.alit.2017.07.010, PMID 28826938.
Aagaard K, Ma J, Antony KM, Ganu R, Petrosino J, Versalovic J. The placenta harbors a unique microbiome. Sci Transl Med. 2014;6(237):237ra65. doi: 10.1126/scitranslmed.3008599, PMID 24848255.
Rautava S, Collado MC, Salminen S, Isolauri E. Probiotics modulate host-microbe interaction in the placenta and fetal gut: a randomized double-blind placebo-controlled trial. Neonatology. 2012;102(3):178-84. doi: 10.1159/000339182, PMID 22776980.
Jimenez E, Fernandez L, Marin ML, Martin R, Odriozola JM, Nueno Palop C. Isolation of commensal bacteria from umbilical cord blood of healthy neonates born by cesarean section. Curr Microbiol. 2005;51(4):270-4. doi: 10.1007/s00284-005-0020-3, PMID 16187156.
Jimenez E, Marin ML, Martin R, Odriozola JM, Olivares M, Xaus J. Is meconium from healthy newborns actually sterile? Res Microbiol. 2008;159(3):187-93. doi: 10.1016/j.resmic.2007.12.007, PMID 18281199.
Houghteling PD, Walker WA. Why is initial bacterial colonization of the intestine important to infants and children’s health? J Pediatr Gastroenterol Nutr. 2015;60(3):294-307. doi: 10.1097/MPG.0000000000000597, PMID 25313849.
Dogra S, Sakwinska O, Soh SE, Ngom Bru C, Bruck WM, Berger B. Rate of establishing the gut microbiota in infancy has consequences for future health. Gut Microbes. 2015;6(5):321-5. doi: 10.1080/19490976.2015.1078051, PMID 26516657.
Backhed F, Roswall J, Peng Y, Feng Q, Jia H, Kovatcheva-Datchary P. Dynamics and stabilization of the human gut microbiome during the first year of life. Cell Host Microbe. 2015;17(6):852. doi: 10.1016/j.chom.2015.05.012, PMID 26308884.
Almgren M. Benefits of skin-to-skin contact during the neonatal period: governed by epigenetic mechanisms? Genes Dis. 2018;5(1):24-6. doi: 10.1016/j.gendis.2018.01.004, PMID 30258931.
Van Den Elsen LW, Garssen J, Burcelin R, Verhasselt V. Shaping the gut microbiota by breastfeeding: the gateway to allergy prevention? Front Pediatr. 2019;7:47. doi: 10.3389/fped.2019.00047, PMID 30873394.
Sela DA, Chapman J, Adeuya A, Kim JH, Chen F, Whitehead TR. The genome sequence of Bifidobacterium longum subsp. infantis reveals adaptations for milk utilization within the infant microbiome. Proc Natl Acad Sci USA. 2008;105(48):18964-9. doi: 10.1073/pnas.0809584105, PMID 19033196.
Bergstrom A, Skov TH, Bahl MI, Roager HM, Christensen LB, Ejlerskov KT. Establishment of intestinal microbiota during early life: a longitudinal explorative study of a large cohort of Danish infants. Appl Environ Microbiol. 2014;80(9):2889-900. doi: 10.1128/AEM.00342-14, PMID 24584251.
Faith JJ, Guruge JL, Charbonneau M, Subramanian S, Seedorf H, Goodman AL. The long-term stability of the human gut microbiota. Science. 2013;341(6141):1237439. doi: 10.1126/science.1237439, PMID 23828941.
Rajilic Stojanovic M, Heilig HG, Tims S, Zoetendal EG, De Vos WM. Long-term monitoring of the human intestinal microbiota composition. Environ Microbiol. 2013;15(4):1146-59. doi: 10.1111/1462-2920.12023, PMID 23286720.
Yassour M, Vatanen T, Siljander H, Hamalainen AM, Harkonen T, Ryhanen SJ. Natural history of the infant gut microbiome and impact of antibiotic treatment on bacterial strain diversity and stability. Sci Transl Med. 2016;8(343):343ra81. doi: 10.1126/scitranslmed.aad0917, PMID 27306663.
Koenig JE, Spor A, Scalfone N, Fricker AD, Stombaugh J, Knight R. Succession of microbial consortia in the developing infant gut microbiome. Proc Natl Acad Sci USA. 2011;108 (Suppl 1):4578-85. doi: 10.1073/pnas.1000081107, PMID 20668239.
Derrien M, Alvarez AS, De Vos WM. The gut microbiota in the first decade of life. Trends Microbiol. 2019;27(12):997-1010. doi: 10.1016/j.tim.2019.08.001, PMID 31474424.
Zhong H, Penders J, Shi Z, Ren H, Cai K, Fang C. Impact of early events and lifestyle on the gut microbiota and metabolic phenotypes in young school-age children. Microbiome. 2019;7(1):2. doi: 10.1186/s40168-018-0608-z, PMID 30609941.
Deering KE, Devine A, O’ Sullivan TA, Lo J, Boyce MC, Christophersen CT. Characterizing the composition of the pediatric gut microbiome: a systematic review. Nutrients. 2019;12(1):16. doi: 10.3390/nu12010016, PMID 31861722.
Wang J, Thingholm LB, Skieceviciene J, Rausch P, Kummen M, Hov JR. Genome-wide association analysis identifies variation in vitamin D receptor and other host factors influencing the gut microbiota. Nat Genet. 2016;48(11):1396-406. doi: 10.1038/ng.3695, PMID 27723756.
Bonder MJ, Kurilshikov A, Tigchelaar EF, Mujagic Z, Imhann F, Vila AV. The effect of host genetics on the gut microbiome. Nat Genet. 2016;48(11):1407-12. doi: 10.1038/ng.3663, PMID 27694959.
Blekhman R, Goodrich JK, Huang K, Sun Q, Bukowski R, Bell JT. Host genetic variation impacts microbiome composition across human body sites. Genome Biol. 2015;16(1):191. doi: 10.1186/s13059-015-0759-1, PMID 26374288.
Xu F, Fu Y, Sun TY, Jiang Z, Miao Z, Shuai M. The interplay between host genetics and the gut microbiome reveals common and distinct microbiome features for complex human diseases. Microbiome. 2020;8(1):145. doi: 10.1186/s40168-020-00923-9, PMID 33032658.
Turpin W, Espin Garcia O, Xu W, Silverberg MS, Kevans D, Smith MI. Association of host genome with intestinal microbial composition in a large healthy cohort. Nat Genet. 2016;48(11):1413-7. doi: 10.1038/ng.3693, PMID 27694960.
Goodrich JK, Waters JL, Poole AC, Sutter JL, Koren O, Blekhman R. Human genetics shape the gut microbiome. Cell. 2014;159(4):789-99. doi: 10.1016/j.cell.2014.09.053, PMID 25417156.
Human Microbiome Project Consortium. Structure function and diversity of the healthy human microbiome. Nature. 2012;486(7402):207-14. doi: 10.1038/nature11234, PMID 22699609.
Deschasaux M, Bouter KE, Prodan A, Levin E, Groen AK, Herrema H. Depicting the composition of gut microbiota in a population with varied ethnic origins but shared geography. Nat Med. 2018;24(10):1526-31. doi: 10.1038/s41591-018-0160-1, PMID 30150717.
Gaulke CA, Sharpton TJ. The influence of ethnicity and geography on human gut microbiome composition. Nat Med. 2018;24(10):1495-6. doi: 10.1038/s41591-018-0210-8, PMID 30275567.
Wang M, Li M, Wu S, Lebrilla CB, Chapkin RS, Ivanov I. Fecal microbiota composition of breast-fed infants is correlated with human milk oligosaccharides consumed. J Pediatr Gastroenterol Nutr. 2015;60(6):825-33. doi: 10.1097/MPG.0000000000000752, PMID 25651488.
Iyengar SR, Walker WA. Immune factors in breast milk and the development of atopic disease. J Pediatr Gastroenterol Nutr. 2012;55(6):641-7. doi: 10.1097/MPG.0b013e3182617a9d, PMID 22684347.
Dzidic M, Mira A, Artacho A, Abrahamsson TR, Jenmalm MC, Collado MC. Allergy development is associated with consumption of breastmilk with a reduced microbial richness in the first month of life. Pediatr Allergy Immunol. 2019;31(3):250-7. doi: 10.1111/pai.13176, PMID 31736150.
Cacho NT, Lawrence RM. Innate immunity and breast milk. Front Immunol. 2017;8:584. doi: 10.3389/fimmu.2017.00584, PMID 28611768.
Reguigne Arnould I, Couillin P, Mollicone R, Faure S, Fletcher A, Kelly RJ. Relative positions of two clusters of human alpha-L-fucosyltransferases in 19q (FUT1-FUT2) and 19p (FUT6-FUT3-FUT5) within the microsatellite genetic map of chromosome 19. Cytogenet Cell Genet. 1995;71(2):158-62. doi: 10.1159/000134098, PMID 7656588.
Hao H, Zhu L, Faden HS. The milk-based diet of infancy and the gut microbiome. Gastroenterol Rep (Oxf). 2019;7(4):246-9. doi: 10.1093/gastro/goz031, PMID 31413830.
Turpin W, Bedrani L, Espin Garcia O, Xu W, Silverberg MS, Smith MI. FUT2 genotype and secretory status are not associated with fecal microbial composition and inferred function in healthy subjects. Gut Microbes. 2018;9(4):357-68. doi: 10.1080/19490976.2018.1445956, PMID 29533703.
Palmer C, Bik EM, Di Giulio DB, Relman DA, Brown PO. Development of the human infant intestinal microbiota. PLOS Biol. 2007;5(7):e177. doi: 10.1371/journal.pbio.0050177, PMID 17594176.
Differding MK, Benjamin Neelon SE, Hoyo C, Ostbye T, Mueller NT. Timing of complementary feeding is associated with gut microbiota diversity and composition and short chain fatty acid concentrations over the first year of life. BMC Microbiol. 2020;20(1):56. doi: 10.1186/s12866-020-01723-9, PMID 32160858.
Rutayisire E, Huang K, Liu Y, Tao F. The mode of delivery affects the diversity and colonization pattern of the gut microbiota during the first year of infants’ life: a systematic review. BMC Gastroenterol. 2016;16(1):86. doi: 10.1186/s12876-016-0498-0, PMID 27475754.
Sevelsted A, Stokholm J, Bonnelykke K, Bisgaard H. Cesarean section and chronic immune disorders. Pediatrics. 2015;135(1):e92-8. doi: 10.1542/peds.2014-0596, PMID 25452656.
Dominguez Bello MG, De Jesus Laboy KM, Shen N, Cox LM, Amir A, Gonzalez A. Partial restoration of the microbiota of cesarean-born infants via vaginal microbial transfer. Nat Med. 2016;22(3):250-3. doi: 10.1038/nm.4039, PMID 26828196.
Betran AP, Ye J, Moller AB, Zhang J, Gulmezoglu AM, Torloni MR. The increasing trend in caesarean section rates: global regional and national estimates: 1990-2014. PLOS One. 2016;11(2):e0148343. doi: 10.1371/journal.pone.0148343, PMID 26849801.
Dominguez Bello MG, Costello EK, Contreras M, Magris M, Hidalgo G, Fierer N. Delivery mode shapes the acquisition and structure of the initial microbiota across multiple body habitats in newborns. Proc Natl Acad Sci USA. 2010;107(26):11971-5. doi: 10.1073/pnas.1002601107, PMID 20566857.
Levin AM, Sitarik AR, Havstad SL, Fujimura KE, Wegienka G, Cassidy Bushrow AE. Joint effects of pregnancy sociocultural and environmental factors on early life gut microbiome structure and diversity. Sci Rep. 2016;6:31775. doi: 10.1038/srep31775, PMID 27558272.
Dogra S, Sakwinska O, Soh SE, Ngom Bru C, Bruck WM, Berger B. Dynamics of infant gut microbiota are influenced by delivery mode and gestational duration and are associated with subsequent adiposity. mBio. 2015;6(1):e02419-14. doi: 10.1128/mBio.02419-14, PMID 25650398.
Hesla HM, Stenius F, Jaderlund L, Nelson R, Engstrand L, Alm J. Impact of lifestyle on the gut microbiota of healthy infants and their mothers the ALADDIN birth cohort. FEMS Microbiol Ecol. 2014;90(3):791-801. doi: 10.1111/1574-6941.12434, PMID 25290507.
Renz Polster H, David MR, Buist AS, Vollmer WM, O’Connor EA, Frazier EA. Caesarean section delivery and the risk of allergic disorders in childhood. Clin Exp Allergy. 2005;35(11):1466-72. doi: 10.1111/j.1365-2222.2005.02356.x, PMID 16297144.
Imoto N, Morita H, Amanuma F, Maruyama H, Watanabe S, Hashiguchi N. Maternal antimicrobial use at delivery has a stronger impact than mode of delivery on bifidobacterial colonization in infants: a pilot study. J Perinatol. 2018;38(9):1174-81. doi: 10.1038/s41372-018-0172-1, PMID 30042470.
Kingsbury MA, Bilbo SD. The inflammatory event of birth: how oxytocin signaling may guide the development of the brain and gastrointestinal system. Front Neuroendocrinol. 2019;55:100794. doi: 10.1016/j.yfrne.2019.100794, PMID 31560883.
Strzępa A, Lobo FM, Majewska Szczepanik M, Szczepanik M. Antibiotics and autoimmune and allergy diseases: causative factor or treatment? Int Immunopharmacol. 2018;65:328-41. doi: 10.1016/j.intimp.2018.10.021, PMID 30359934.
Metsala J, Lundqvist A, Virta LJ, Kaila M, Gissler M, Virtanen SM. Prenatal and post-natal exposure to antibiotics and risk of asthma in childhood. Clin Exp Allergy. 2015;45(1):137-45. doi: 10.1111/cea.12356, PMID 24943808.
Van Der Waaij D, Nord CE. Development and persistence of multi-resistance to antibiotics in bacteria; an analysis and a new approach to this urgent problem. Int J Antimicrob Agents. 2000;16(3):191-7. doi: 10.1016/S0924-8579(00)00227-2, PMID 11091035.
Belizario JE, Faintuch J. Microbiome and gut dysbiosis. Exp Suppl. 2018;109:459-76. doi: 10.1007/978-3-319-74932-7_13, PMID 30535609.
Zimmermann P, Curtis N. The effect of antibiotics on the composition of the intestinal microbiota a systematic review. J Infect. 2019;79(6):471-89. doi: 10.1016/j.jinf.2019.10.008, PMID 31629863.
Hagan T, Cortese M, Rouphael N, Boudreau C, Linde C, Maddur MS. Antibiotics-driven gut microbiome perturbation alters immunity to vaccines in humans. Cell. 2019;178(6):1313-28.e13. doi: 10.1016/j.cell.2019.08.010, PMID 31491384.
Palleja A, Mikkelsen KH, Forslund SK, Kashani A, Allin KH, Nielsen T. Recovery of gut microbiota of healthy adults following antibiotic exposure. Nat Microbiol. 2018;3(11):1255-65. doi: 10.1038/s41564-018-0257-9, PMID 30349083.
Korpela K, Salonen A, Saxen H, Nikkonen A, Peltola V, Jaakkola T. Antibiotics in early life associate with specific gut microbiota signatures in a prospective longitudinal infant cohort. Pediatr Res. 2020;88(3):438-43. doi: 10.1038/s41390-020-0761-5, PMID 31954376.
Sun L, Zhang X, Zhang Y, Zheng K, Xiang Q, Chen N. Antibiotic-induced disruption of gut microbiota alters local metabolomes and immune responses. Front Cell Infect Microbiol. 2019;9:99. doi: 10.3389/fcimb.2019.00099, PMID 31069173.
Singh RK, Zogg H, Ghoshal UC, Ro S. Current treatment options and therapeutic insights for gastrointestinal dysmotility and functional gastrointestinal disorders. Front Pharmacol. 2022;13:808195. doi: 10.3389/fphar.2022.808195, PMID 35145413.
Camilleri M, Atieh J. New developments in prokinetic therapy for gastric motility disorders. Front Pharmacol. 2021;12:711500. doi: 10.3389/fphar.2021.711500, PMID 34504426.
Camilleri M, Kerstens R, Rykx A, Vandeplassche L. A placebo-controlled trial of prucalopride for severe chronic constipation. N Engl J Med. 2008;358(22):2344-54. doi: 10.1056/NEJMoa0800670, PMID 18509121.
Camilleri M. Leaky gut: mechanisms measurement and clinical implications in humans. Gut. 2019;68(8):1516-26. doi: 10.1136/gutjnl-2019-318427, PMID 31076401.
Camilleri M, Madsen K, Spiller R, Greenwood Van Meerveld B, Verne GN. Intestinal barrier function in health and gastrointestinal disease. Neurogastroenterol Motil. 2012;24(6):503-12. doi: 10.1111/j.1365-2982.2012.01921.x, PMID 22583600.
Camilleri M. New drugs on the horizon for functional and motility gastrointestinal disorders. Gastroenterology. 2021;161(3):761-4. doi: 10.1053/j.gastro.2021.04.079, PMID 33989661.
Camilleri M, Nord SL, Burton D, Oduyebo I, Zhang Y, Chen J. Randomised clinical trial: significant biochemical and colonic transit effects of the farnesoid X receptor agonist tropifexor in patients with primary bile acid diarrhoea. Aliment Pharmacol Ther. 2020;52(5):808-20. doi: 10.1111/apt.15967, PMID 32702169.
Camilleri M, Stanghellini V. Current management strategies and emerging treatments for functional dyspepsia. Nat Rev Gastroenterol Hepatol. 2013;10(3):187-94. doi: 10.1038/nrgastro.2013.11, PMID 23381190.
Charbonneau D, Gibb RD, Quigley EM. Fecal excretion of Bifidobacterium infantis 35624 and changes in fecal microbiota after eight weeks of oral supplementation with encapsulated probiotic. Gut Microbes. 2013;4(3):201-11. doi: 10.4161/gmic.24196, PMID 23549409.
Chedid V, Camilleri M. Relamorelin for the treatment of gastrointestinal motility disorders. Expert Opin Investig Drugs. 2017;26(10):1189-97. doi: 10.1080/13543784.2017.1373088, PMID 28847163.
Lee JY, Kim N, Choi YJ, Park JH, Ashktorab H, Smoot DT. Expression of tight junction proteins according to functional dyspepsia subtype and sex. J Neurogastroenterol Motil. 2020;26(2):248-58. doi: 10.5056/jnm19208, PMID 32235032.
Shetty GB, Rameshwar TK, Sumana K. Preliminary oral probiotics bacterial profile in neonatal and pediatrics and its clinical evaluation. Int J Curr Pharm Sci. 2022;14(4):5-9. doi: 10.22159/ijcpr.2022v14i4.2005.
Mandal B. Bacteriocin produced by lactic acid bacteria: a probiotic. Int J Pharm Pharm Sci. 2024;16(3):1-7. doi: 10.22159/ijpps.2024v16i3.50326.