
Int J App Pharm, Vol 18, Issue 1, 2026, 314-324Original Article
COMPUTATIONAL INVESTIGATION OF THE ANTICOAGULANT POTENTIAL OF SARGASSUM SP. BIOACTIVE COMPOUNDS
FRANCISCA DIANA ALEXANDRA1,2*, MARIANTI A. MANGGAU3, MUH. NASSRUM MASSI4, YANTI LEMAN5, YSRAFIL YSRAFIL2
1Faculty of Medicine, Hasanuddin University, Tamalanrea, Makassar-90245, Indonesia. 2Department of Pharmacotherapy, Faculty of Medicine, Universitas Palangka Raya, Palangka Raya-73111, Indonesia. 3Department of Pharmacy, Faculty of Pharmacy, Hasanuddin University, Tamalanrea, Makassar-90245, Indonesia. 4Department of Microbiology, Faculty of Medicine, University of Hasanuddin, Makassar, Indonesia. 5Department of Pharmacology, Faculty of Medicine, Universitas Hasanuddin, Makassar, Indonesia
*Corresponding author: Francisca Diana Alexandra; *Email: francisca@med.upr.ac.id
Received: 31 Aug 2025, Revised and Accepted: 05 Dec 2025
ABSTRACT
Objective: This study investigates the anticoagulant potential of bioactive compounds from Sargassum sp. using computational techniques. The goal is to explore their ability to modulate thrombosis, coagulation, and inflammation-related pathways through molecular docking, network pharmacology, and molecular dynamics (MD) simulations.
Methods: Network pharmacology was applied to identify target proteins related to thrombosis and coagulation. Gene Ontology (GO) and KEGG enrichment analyses highlighted relevant biological functions. Molecular docking simulations assessed binding interactions between bioactive compounds and coagulation-related proteins, while molecular dynamics simulations evaluated the stability of these complexes.
Results: A total of 109 coagulation-related proteins were identified, with 35 core proteins forming a highly connected PPI network. Enriched pathways included platelet aggregation and endothelial function. Docking results showed stable binding of alginate and fucoxanthin to SRC and HSP90AA1, with binding scores of-4.7 and-4.3 kcal/mol, respectively. Fucoidan demonstrated stronger binding to MAPK1 (-2.6 kcal/mol). Molecular dynamics simulations confirmed stable complexes, but further simulations (100 ns) are recommended to refine the findings.
Conclusion: Sargassum compounds, particularly alginate and fucoxanthin, may modulate thrombosis and coagulation pathways, likely through indirect mechanisms like inflammation and platelet aggregation. Experimental validation is required to confirm these computational predictions. This study underscores the value of computational models in hypothesis generation and the need for experimental confirmation.
Keywords: Anticoagulant, Bioactive compounds, Molecular dynamics, Network pharmacology, Sargassum sp
© 2026 The Authors. Published by Innovare Academic Sciences Pvt Ltd. This is an open access article under the CC BY license (https://creativecommons.org/licenses/by/4.0/)
DOI: https://dx.doi.org/10.22159/ijap.2026v18i1.56715 Journal homepage: https://innovareacademics.in/journals/index.php/ijap
INTRODUCTION
Thromboembolic disorders, including deep vein thrombosis (DVT), pulmonary embolism (PE), and stroke associated with atrial fibrillation, remain significant global health challenges, contributing to high rates of morbidity and mortality. These conditions arise from abnormal blood clot formation, which can migrate to vital organs, causing severe complications such as embolism or ischemic stroke. DVT and PE are ubiquitous in hospitalized patients, accounting for millions of deaths annually and imposing substantial economic and healthcare burdens. Similarly, atrial fibrillation-related strokes are a leading cause of long-term disability and death.
Current anticoagulant therapies, such as vitamin K antagonists (VKAs), heparin-based treatments, and direct oral anticoagulants (DOACs), are effective but have notable limitations, including bleeding risks, patient variability, and the need for continuous monitoring [1, 2]. Warfarin, a common VKA, requires frequent International Normalized Ratio (INR) checks due to its narrow therapeutic window [3]. Heparin and low-molecular-weight heparins (LMWHs) offer rapid anticoagulation but require parenteral administration and are associated with complications such as heparin-induced thrombocytopenia [4, 5]. DOACs provide predictable pharmacokinetics but are unsuitable for patients with renal or cardiovascular comorbidities. These challenges underscore the need for safer and more accessible anticoagulants [6, 7].
Marine-derived bioactive compounds, particularly polysaccharides from brown algae, have emerged as promising candidates for developing natural anticoagulants. Fucoidan, a sulfated polysaccharide found in Sargassum sp., enhances antithrombin III activity and exhibits anticoagulant, antioxidant, and anti-inflammatory properties beneficial for cardiovascular health [8-10]. Several studies have shown that sulfated polysaccharides from algae modulate clotting factors and inhibit platelet aggregation, with biological activity strongly influenced by sulfation patterns [11, 12]. However, despite increasing evidence of anticoagulant potential, the molecular mechanisms by which Sargassum sp. compounds interact with coagulation pathways remain poorly defined [13].
Recent advances in computational biology have enabled the exploration of such mechanisms through network pharmacology and molecular docking. These methods predict bioactive compound-protein interactions and identify key targets across multiple biological pathways [14-17]. Molecular dynamics (MD) simulations further provide insight into protein-ligand complexes' stability and conformational dynamics, bridging theoretical predictions with biological relevance.
The present study integrates these in silico methods, network pharmacology, molecular docking, and MD simulations, to elucidate the potential anticoagulant mechanisms of Sargassum sp. compounds. This multi-step computational approach aims to identify key coagulation-related protein targets, predict binding interactions, and assess molecular stability, thereby providing a mechanistic foundation for the therapeutic application of marine-derived bioactives. By addressing existing gaps in molecular characterization, this study contributes to the rational development of safer and more effective anticoagulant agents derived from marine sources.
MATERIALSAND METHODS
Network pharmacology
The network pharmacology workflow involved multiple stages, including protein screening, compound sampling, protein-target mapping, protein interaction analysis, identification of core protein, and pathway enrichment analysis. To strengthen the interpretation of pathway enrichment, statistical significance thresholds (p<0.05) were applied to ensure robust selection criteria for relevant pathways.
Finding of coagulation related proteins
This analysis was conducted to identify proteins involved in the molecular coagulation mechanism. This analysis was conducted using human GeneCards (https://www.genecards.org), a human gene database containing information on genes and/or proteins in the human body, including those involved in a disease. In addition, we also use Online Mendelian Inheritance in Man (OMIM) (https://www.omim.org) as comprehensive and authoritative database on human genes and genetic phenotypes that is available free of charge and updated daily. The finding of coagulation-related proteins in both databases was performed using the keyword ‘coagulation’. Protein-related data sets were combined and duplicate findings were removed [18, 19].
Analysis of Sargassumsp compounds target
Validated bioactive compounds of Sargassum sp. reported in literature, including fucoidan, fucoxanthine, laminarin, phloroglucinol, and alginate were use in this research. These compounds were retrieved from PubChem for further analysed their potential target proteins. The analysis is performed using tools Pharm Mapper (http://lilab-ecust.cn/pharmmapper/index.html) as updated integrated pharmacophore matching platform with statistial method for compounds potential target identification. Te tools were set in ‘Human Only’ mode and selection the target by z-score greater than 1.5 following prior literature recommendations for optimal target selection. Additionally, we also use swiss target prediction as tools that could provide most probable protein targets of small molecules. This is intended to investigate more targets of the compound. The target proteins of each compound are combined and the repeated ones are removed [19, 20].
Relation of Sargassumsp. target on coagulation
The target compounds of Sargassum sp. obtained were further intersected with coagulation-related proteins identified previously. This was conducted using a Venn diagram, Venny 2.1.0 (https://bioinfogp.cnb.csic.es/tools/venny/) to visualize the overlapped and particular targets. The overlapped protein indicated as coagulation related protein that targeted by Sargassumsp compounds.
Protein-protein interaction network construction and identification of core proteins
Overlapping protein targets obtained in previous step were further build in form of a protein-protein interaction (PPI) network to reflect the interaction between proteins targeted by Sargassum sp compounds in coagulation disease. The construction of PPI network were conduct using STRING (Search Tool for the Retrieval of Interacting Genes/Proteins) database. The PPI were construct following setting: species “homo sapiens”, required score “highest confidence (0.900)” and false discovery rate (FDR) stringency “medium (5 percent)”. The results obtained in the visualisation were further processed using Cytoscape 3.9.1. Proteins not included in the main network were removed, and the remainder were further analysed to determine protein topology. Furthermore, analysis core protein were conduct using CytoNCA, a plug in of cytoscape that provide precision in identification of core protein. The determination of this potential is based on several centrality parameters, namely degree (DC), betweenness (BC), closeness (CC) and eigenvector (EC) [19].
Functional enrichment analysis
Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment analyses were performed using ShinyGo (https://bioinformatics.sdstate.edu/go80/) on the genes/proteins included in the main network obtained from STRING screening and topological analysis described in the previous section. In this study, a total of 35 core proteins identified from the topological analysis were subjected to GO and KEGG enrichment analyses. GO analysis classified the proteins according to their biological processes, molecular functions, and cellular components, while KEGG analysis identified significantly enriched signaling pathways related to anticoagulation. The tool was set to the “human” species mode, with classification based on enrichment level and a false discovery rate (FDR) cutoff value of 0.05. The top 10 enriched terms from each analysis were selected and visualized as lollipop plots.
Preparation for molecular docking
Molecular docking was performed to predict the binding interactions between Sargassum sp. bioactive compounds and coagulation-related proteins. Molecular docking were conduct using MGL tools package and autodock vina. Prior to docking, the preparation of protein and ligand were conducted properelly. The 3D structure of protein were retrieve from RCSB PDB that further prepare by Biovia Discovery Studio Visualizer and Autodock. The prepared protein subsequently determine their Gridbox and validated by Autodock (valid protein were assumed by<2 Å of RMSD). Menwhile, The 3D structures of selected compounds were optimized using Marvin Sketch and autodock. Both of protein and compounds were saved in PDBQT format prior to docking [19, 21, 22].
Molecular dynamics (MD) simulations
To evaluate the stability of the ligand–protein complexes, molecular dynamics (MD) simulations were performed using YASARA Dynamics (https://www.yasara.org/). Two representative complexes, HSP90AA1–Fucoxanthine and SRC–Alginate, were selected for detailed simulations. The MD simulations were run under physiological conditions (310 K, pH 7.4) for 20 ns using the AMBER14 force field. The systems were solvated using the TIP3P water model with an ionic strength of 0.9% NaCl to mimic physiological conditions.
MD simulations allow for the study of the dynamic behavior of protein-ligand complexes, providing insights into the stability and flexibility of the interactions. Several analyses were conducted to assess the stability of the complexes during the simulations:
Potential Energy Analysis: Fluctuations in the total potential energy were examined to assess system stability.
Solvent Accessible Surface Area (SASA): Changes in SASA were used to infer ligand exposure to solvent molecules, which provides insights into the protein's conformational changes during ligand binding.
Root mean Square Deviation (RMSD): The RMSD of the ligand-protein complexes was tracked over simulation time to monitor the structural stability of the complex.
Root mean Square Fluctuation (RMSF): Residue-level fluctuations were analyzed to identify flexible regions of the protein that may play a role in ligand binding.
Molecular Mechanics Poisson-Boltzmann Surface Area (MMPBSA): Binding free energies were calculated to predict ligand affinity under solvated conditions.
While a 20 ns simulation provides initial insights, extending the simulation time to 100 ns or longer would provide more reliable results regarding the equilibrium and stability of the protein-ligand complexes. These analyses will be crucial for understanding the long-term stability and biological relevance of the interactions observed in the docking simulations [23].
RESULTS
Identification of potential therapeutic targets
This study identified five bioactive compounds from Sargassum sp., namely fucoxanthine, laminarin, phloroglucinol, alginate, and fucoidan. These compounds were selected based on their previously reported bioactivity and compatibility with coagulation-related proteins. The identification process was facilitated using Swiss Target Prediction and PharmMapper, two well-established databases for predicting compound-target interactions. A total of 5050 coagulation-related proteins were initially identified through these platforms. After further filtering, 188 proteins targeted explicitly by the Sargassum sp. compounds were identified. A Venn diagram was constructed to visualize the intersection of the target proteins to narrow down the potential coagulation-related targets. The results revealed an overlap of 109 coagulation-associated proteins targeted by multiple bioactive compounds from Sargassum sp. (fig. 1). These overlapping targets suggest a multi-target action of Sargassum sp. compounds, reinforcing their potential as candidates for anticoagulant therapy.
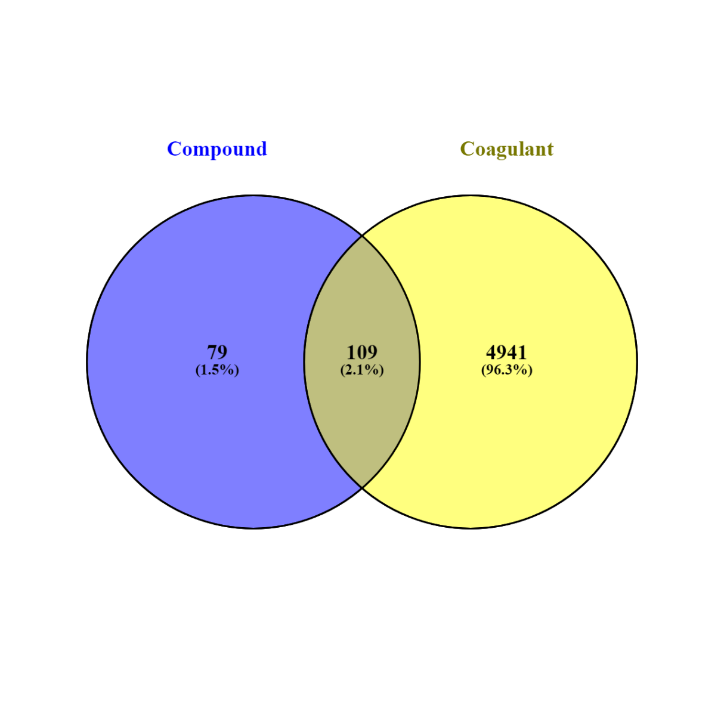
Fig. 1: Venn diagram of coagulation-related proteins and Sargassum sp. Targets
Analysis of protein-protein interaction networks and identification of core proteins
A protein-protein interaction (PPI) network was constructed using the STRING database to understand further the role of Sargassum sp. compounds in coagulation pathways. The analysis was performed in "Homo sapiens" mode with a confidence threshold set at>0.9, ensuring the inclusion of high-confidence interactions. After filtering out duplicate and irrelevant data, 35 potential proteins with 35 nodes and 65 edges were identified, demonstrating strong connectivity within the coagulation cascades. The network analysis revealed an average node degree and clustering coefficient that suggested significant biological interaction between the identified proteins (fig. 2).
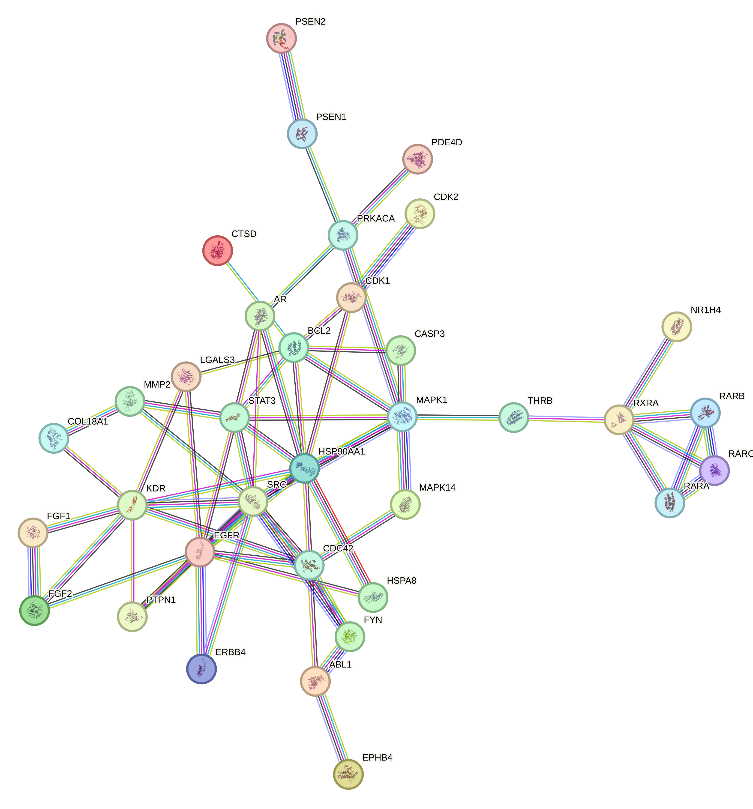
Fig. 2: Protein-protein interaction network involving coagulation targets of Sargassum sp. compounds after filtering
To identify the most influential proteins within this network, the CytoNCA plugin in Cytoscape was used to calculate centrality parameters, including degree (DC), betweenness (BC), closeness (CC), and eigenvector centrality (EC). These parameters are commonly employed in network pharmacology to pinpoint core proteins central to the biological network. These analyses identified three core proteins—SRC, MAPK1, and HSP90AA1—as the network's most influential nodes. These proteins are well-documented in coagulation and inflammatory signaling pathways, indicating that Sargassum sp. compounds may exert their anticoagulant effects by modulating these key regulatory nodes (fig. 3A).
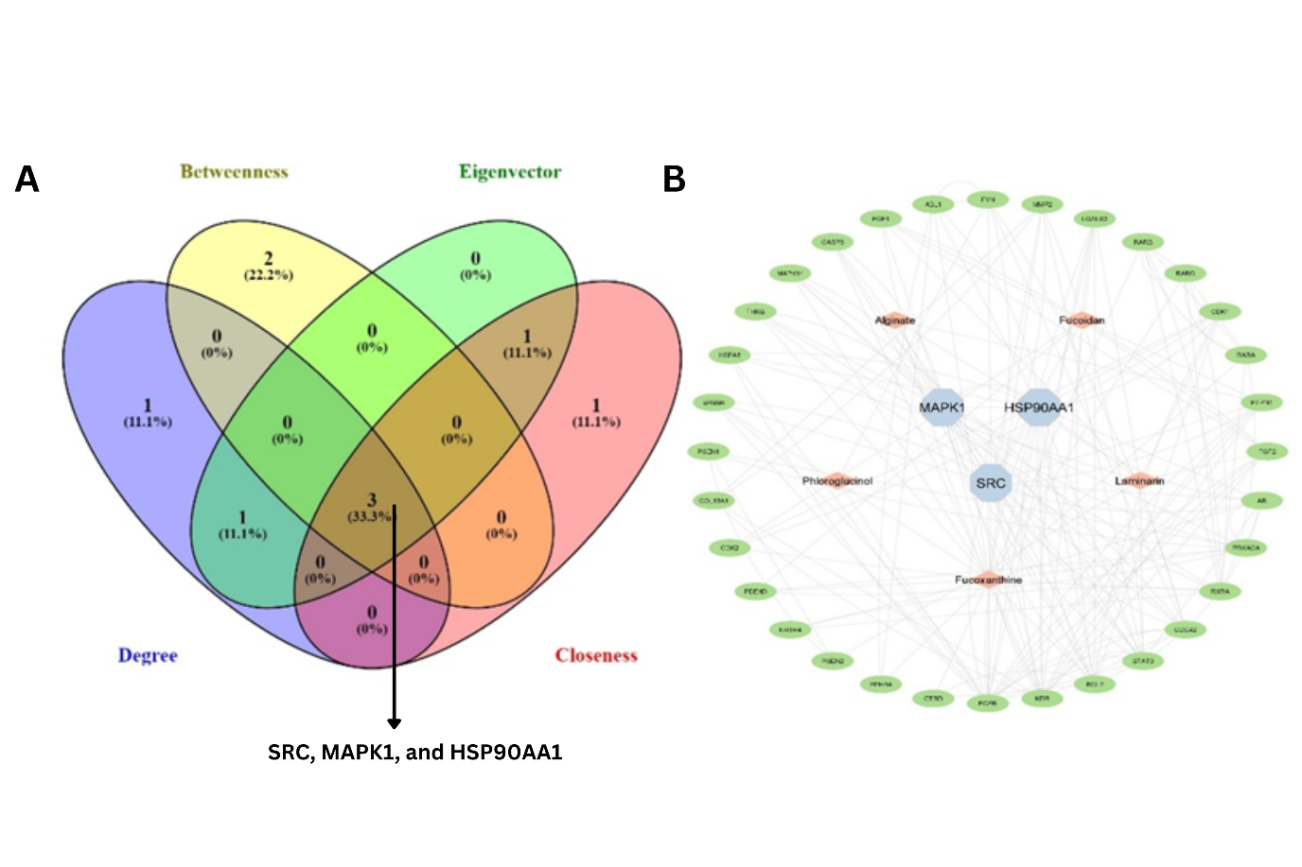
Fig. 3: (A) Top five proteins in four centrality parameters, including Degree, Betweenness, Eigenvector, and Closeness, specifying three core proteins involved and targeted by Sargassum sp. compounds in thrombosis. (B) Compound-target disease network showing the three potential core proteins
The protein-target interaction network constructed using Cytoscape (fig. 3B) further illustrates the multi-target nature of Sargassum sp. compounds, highlighting their ability to target multiple proteins involved in coagulation pathways simultaneously. This multi-target action could contribute to the overall anticoagulant effects observed in both computational and experimental studies.
Mechanism analysis of coagulation pathway modulation
Gene Ontology (GO) and KEGG pathway enrichment analyses were performed to gain deeper insights into the identified target proteins' biological functions. The analysis was conducted using the ShinyGO v0.77 database, which is widely used for functional enrichment analysis. Thirty-five coagulation-related proteins were selected for enrichment analysis based on their centrality scores and direct involvement in coagulation cascades. The statistical significance of the enriched pathways was determined using FDR (p<0.05) to ensure robustness in the biological interpretation.
The KEGG pathway analysis identified several significant enriched pathways, and the top 10 pathway were depicted in fig. 4D. As show in figure, some of pathway enriched associated with coagulation including key pathways such as EGFR tyrosine kinase inhibitor resistance, estrogen signalling, and adherens junction pathways. These pathways are crucial in regulating vascular homeostasis, endothelial function, and thrombosis. These findings support the hypothesis that Sargassum sp. compounds may exert their effects by modulating these mechanisms, which are critical in coagulation and thrombus formation.
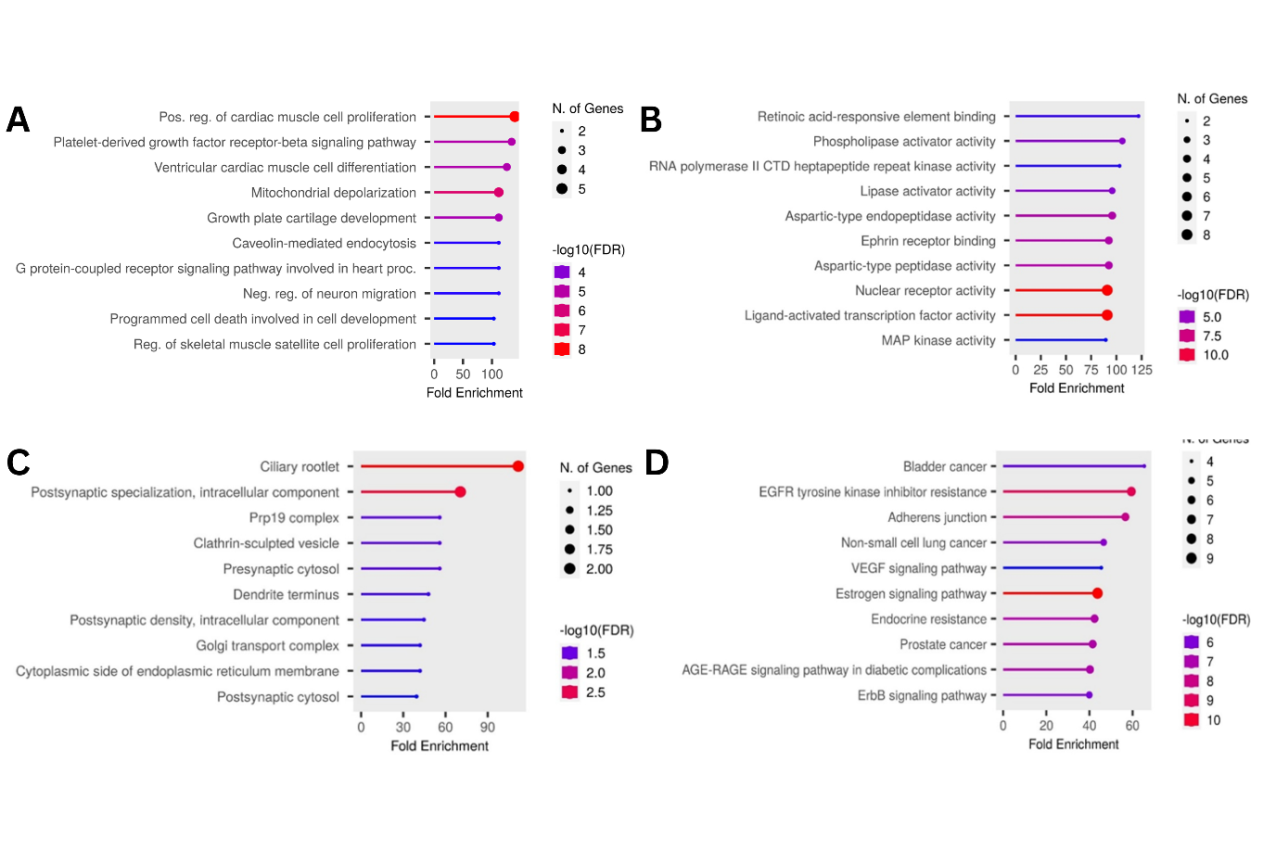
Fig. 4: Gene ontology and KEGG pathway analysis of coagulation-related proteins targeted by bioactive compounds from Sargassum sp. (A) Top 10 biological processes, (B) Top 10 molecular functions, (C) Top 10 cellular components, (D) Signaling pathways
GO analysis provided more profound insights into the molecular functions, cellular components, and biological processes associated with the identified proteins. The analysis of biological processes revealed that the 35 proteins involved in various biological processes in humans. The top 10 of the biological process affected were sow in fig. 4A that associated with blood coagulation key processes such as cardiac muscle cell proliferation, platelet-derived growth factor-beta signalling, ventricular cardiac muscle cell differentiation, mitochondrial depolarization, and cell adhesion regulation, among others. Meanwhile, the top 10 of molecular function that enriched including highlight that the proteins involved on nuclear receptor activity, ligand-activated transcription factor activity, and enzyme-binding interactions (fig. 4B). These molecular functions are critical in signal transduction and homeostatic regulation, further reinforcing the therapeutic potential of Sargassum sp. compounds.
The top 10 of cellular component analysis that significantly enriched highlight several key components such as the ciliary rootlet, extracellular matrix, and postsynaptic specialization (fig. 4C). These components are involved in intracellular signalling, vascular integrity, and endothelial function, suggesting that Sargassum sp. compounds may regulate coagulation at multiple structural and functional levels.
Molecular docking analysis of Sargassum sp. compounds with coagulation-related proteins
Molecular docking simulations were conducted to evaluate the binding interactions between Sargassum sp. bioactive compounds and three key coagulation-related proteins—SRC, MAPK1, and HSP90AA1—identified in the pharmacological network analysis. The docking studies were performed using AutoDock 4 and AutoDock Vina, employing a flexible ligand docking protocol to improve the accuracy of predicting binding affinities and interaction dynamics. Post-docking analysis was carried out to examine hydrogen bonding, hydrophobic interactions, and molecular surface complementarity.
As shown in table 1, the docking results revealed that alginate exhibited the lowest docking score (-4.7 kcal/mol) against SRC, indicating a relatively higher binding affinity compared to the other compounds (fig. 6A). Meanwhile, fucoidan showed a binding affinity of-2.6 kcal/mol toward MAPK1, while fucoxanthin displayed the strongest interaction with HSP90AA1, with a docking score of-4.3 kcal/mol (fig. 5B). However, it is important to note that these binding energies correspond to weak-to-moderate affinities, suggesting that the interactions may not be sufficiently strong to indicate promising anticoagulant potential. This result is still weaker compared to several compounds that act as inhibitors of the three proteins (table 1). Nonetheless, these results should be regarded as preliminary predictive indicators rather than experimental confirmation of biological activity. Further in vitro and in vivo studies are required to validate these computational predictions and confirm the actual anticoagulant effects of these compounds.
Table 1: Molecular docking results of Sargassum sp. bioactive compounds against core proteins
| Molecular | SRC | MAPK1 | HSP90AA1 |
| Fucoidan | -2,6 | -2,6 | -3,8 |
| Fucoxanthine | -1,9 | -1,9 | -5,0 |
| Laminarin | -2,5 | -2,5 | -4 |
| Phloroglucinol | -2,4 | -2 | -2,6 |
| Alginate | -4,7 | -2 | -3,1 |
| Dasatinib (SRC Inhibitor) | -7.7 | - | - |
| U0126 | - | -6.6 | - |
| Luminespib | - | - | -8.5 |
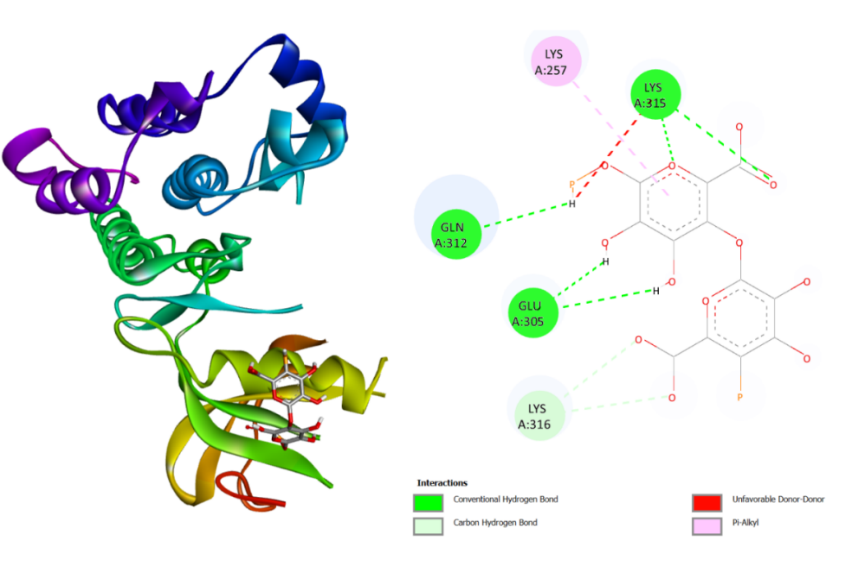
Fig. 5A: 3D and 2D visualization of SRC-Alginate binding
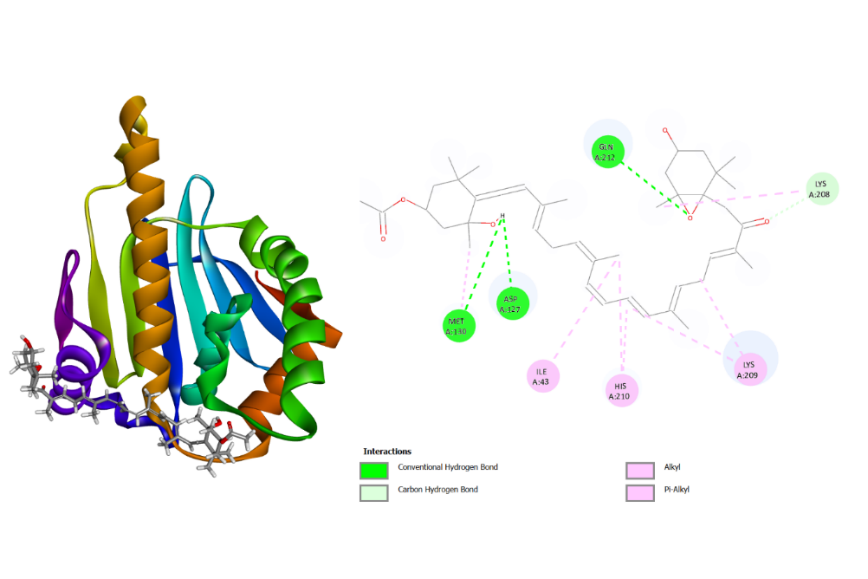
Fig. 5B: 3D and 2D visualization of HSP90AA1-fucoxanthine binding
Molecular dynamics simulation analysis
Molecular dynamics (MD) simulations were conducted to components involved in stabilityand binding interactions with key coagulation-related proteins. The analysis incorporated several structural and energetic parameters, including potential energy fluctuations, molecular mechanics Poisson-Boltzmann surface area (MMPBSA), solvent accessible surface area (SASA), root mean square deviation (RMSD), root mean square fluctuation (RMSF), and radius of gyration (RG). These parameters collectively provide a comprehensive understanding of complex stability, ligand binding affinity, and conformational adaptability over the simulation period.
Potential energy and binding free energy analysis
The total potential energy of the complexes exhibited significant initial fluctuations within the first 0.025 ns, indicative of system equilibration and energy minimization (fig. 6). The fluctuations suggest that ligand binding modulates protein stability by forming stabilizing molecular interactions, as reflected in subsequent convergence toward a stable energy state. The MMPBSA analysis further supported these findings, with binding free energy values ranging between-27,397 and-21,137 kJ/mol over the 20 ns simulation period (fig. 7). These values suggest thermodynamically favorable interactions between Sargassum sp. compounds and their target proteins, as lower binding free energy correlates with stronger and more stable ligand-protein interactions.
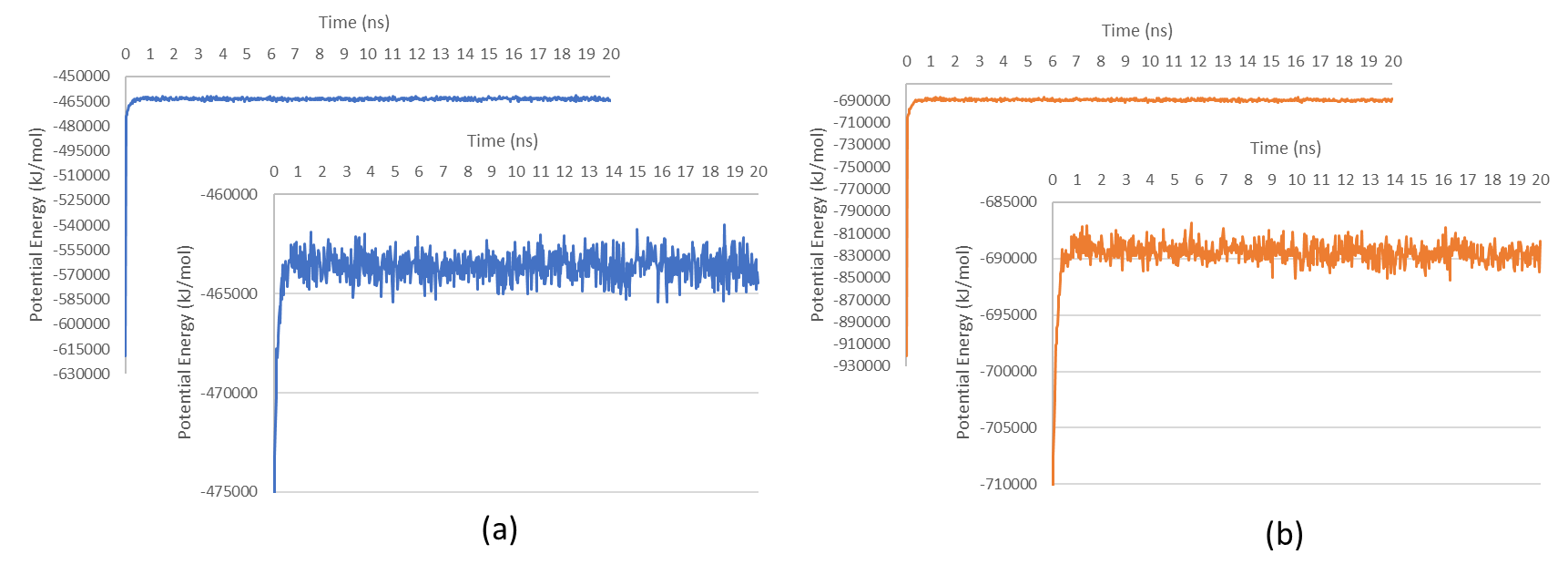
Fig. 6: Potential energy fluctuations of (a) HSP90AA1-Fucoxanthine and (b) SRC-Alginate complexes over the simulation period
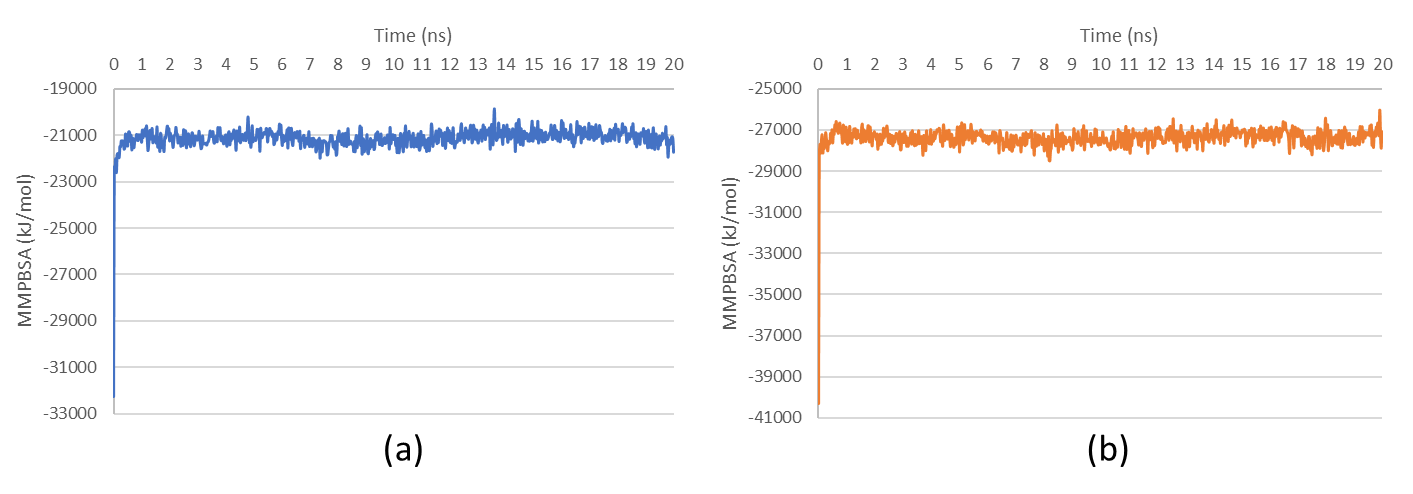
Fig. 7: Binding free energy analysis of (a) HSP90AA1-Fucoxanthine and (b) SRC-Alginate complexes using MMPBSA calculations
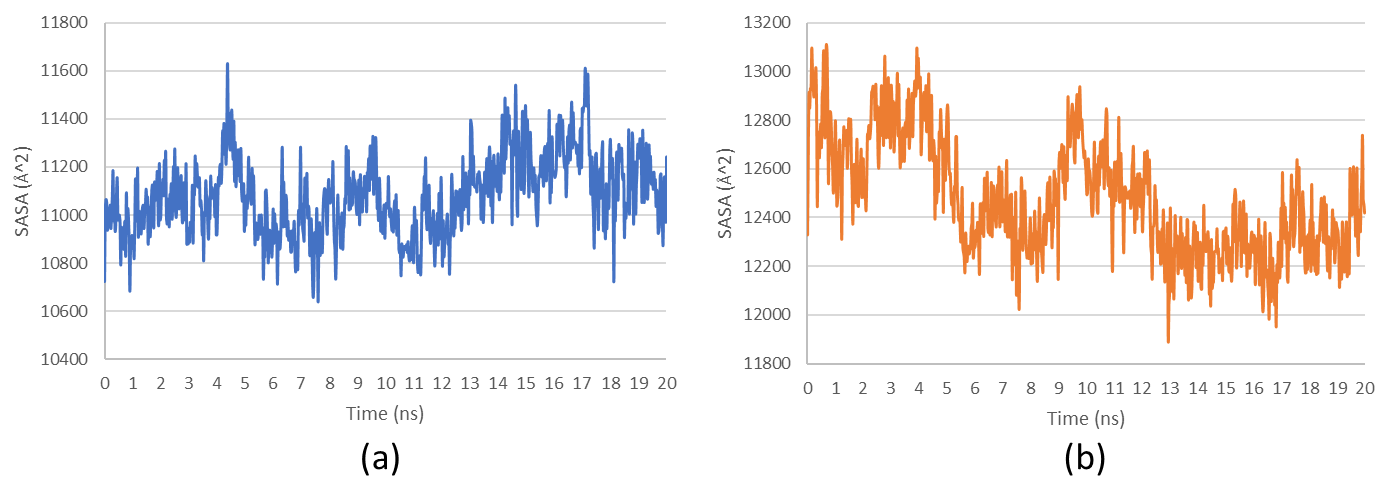
Fig. 8: Solvent accessible surface area (SASA) analysis of (a) HSP90AA1-Fucoxanthine and (b) SRC-Alginate complexes over the simulation period
Solvent accessible surface area (SASA) analysis
SASA values were evaluated to analyze the protein structures' conformational dynamics and solvent exposure during ligand binding. Higher SASA values indicate increased surface exposure, often associated with structural expansion or unfolding, while lower SASA values suggest a more compact and stable conformation. Changes in SASA can reflect ligand-induced conformational rearrangements, which are crucial for understanding the stability of ligand–protein complexes. The results showed a stable SASA pattern for HSP90AA1–Fucoxanthine, suggesting minimal structural perturbations upon ligand binding. In contrast, SRC–Alginate gradually declined in SASA values, indicating a progressive reduction in solvent exposure (fig. 8). However, it is important to note that a decrease in SASA may represent either conformational tightening that enhances structural stability or potential partial denaturation, depending on the protein’s folding behavior and interaction environment. This interpretation highlights the distinct dynamic responses of the two complexes to ligand binding.
Root mean square deviation (RMSD) and conformational stability
RMSD analysis was conducted to assess structural deviations and conformational stability of the protein-ligand complexes over the simulation period. The overall RMSD trends demonstrated an increasing tendency with periodic fluctuations, suggesting inherent conformational flexibility (fig. 9). However, the final RMSD stabilization toward the end of the simulation period indicates that the complexes reached a dynamic equilibrium, reinforcing the reliability of binding interactions. Notably, SRC-Alginate exhibited maximum RMSD values exceeding 3 Å, indicating significant structural rearrangements, which may suggest ligand-induced conformational adaptation or protein flexibility.

Fig. 9: Root mean square deviation (RMSD) analysis of (a) HSP90AA1-Fucoxanthine and (b) SRC-Alginate complexes over the simulation period
Root mean square fluctuation (RMSF) and residue-level conformational changes
RMSF analysis was performed to evaluate the flexibility of individual amino acid residues and their susceptibility to conformational changes upon ligand binding. Higher RMSF values indicate regions with increased mobility, which may correspond to the protein's loop regions, active sites, or flexible terminal regions. Identifying these flexible regions is essential for understanding ligand-induced structural adaptations. The RMSF values for both complexes showed maximum fluctuations exceeding 3 Å at specific residue positions (fig. 10), indicating localized flexibility rather than global structural instability.
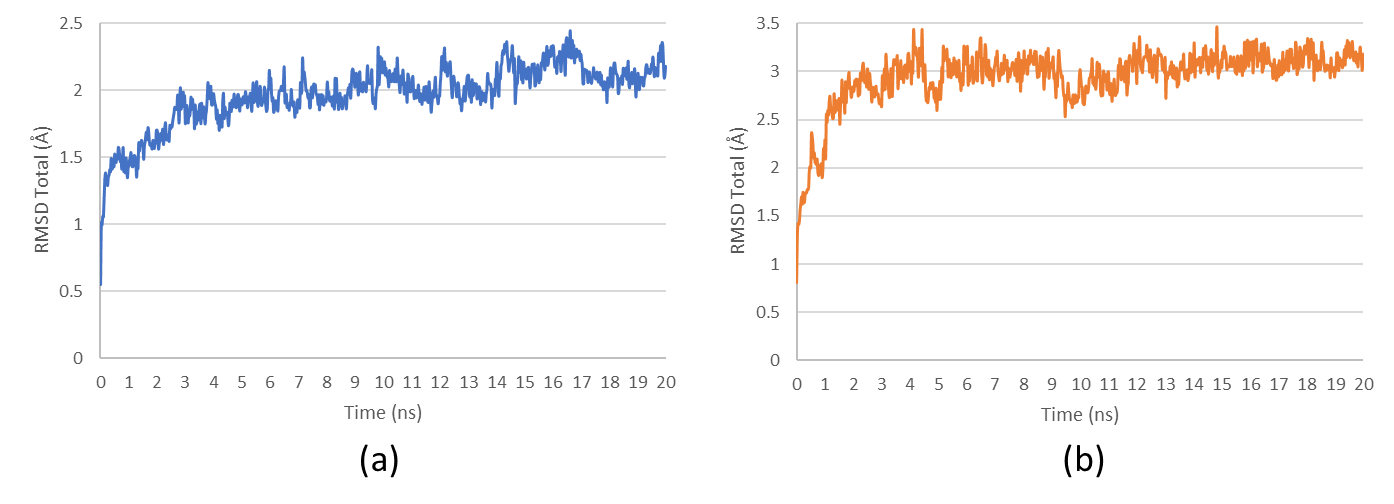
Fig. 10: Root mean square fluctuation (RMSF) analysis of (a) HSP90AA1-Fucoxanthine and (b) SRC-Alginate complexes, highlighting regions of structural flexibility
Radius of gyration (RG) and protein compactness
RG analysis assessed protein compactness and overall structural stability over the simulation period. The RG trends for both complexes correlated with SASA fluctuations, with HSP90AA1–Fucoxanthine maintaining a stable RG profile, indicative of minimal structural perturbations and a well-folded state. In contrast, SRC–Alginate gradually declined in RG values, suggesting progressive protein compaction following ligand binding (fig. 11).
These findings suggest that SRC–Alginate undergoes ligand-induced structural reorganization, potentially leading to a more compact functional state, while HSP90AA1–Fucoxanthine maintains a stable conformation. However, it should be noted that decreases in Rg may reflect either conformational tightening that stabilizes the structure or potential partial denaturation, depending on the protein’s local environment and folding dynamics. This highlights the differences in the binding mechanisms of these bioactive compounds.
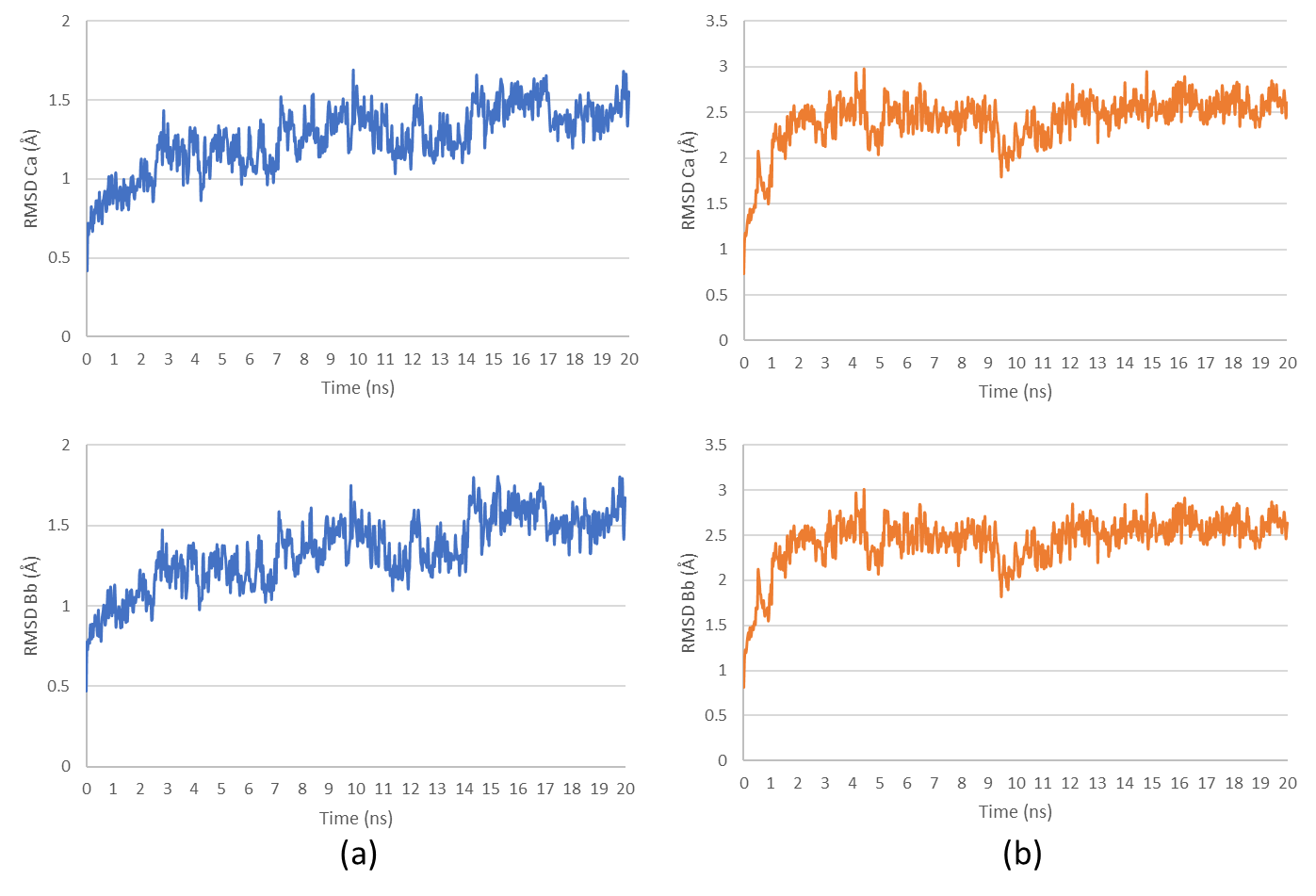
Fig. 11: Radius of gyration (RG) analysis of (a) HSP90AA1-Fucoxanthine and (b) SRC-Alginate complexes, demonstrating structural compactness trends
RMSF analysis of amino acid residues
RMSF (Root mean Square Fluctuation) analysis provides detailed insights into residue-level flexibility and conformational dynamics of the protein-ligand complex. Higher RMSF values indicate regions of greater mobility, such as flexible loops or active sites, which may play a role in ligand binding and allosteric regulation. This analysis is critical for understanding how ligand interactions influence local structural adaptability. The RMSF analysis of amino acid residues in the HSP90AA1-Fucoxanthine and SRC-Alginate complexes revealed maximum fluctuations exceeding 3 Å at specific residue positions (fig. 12). These fluctuations suggest localized flexibility, particularly in loop regions or near binding pockets, where structural adaptations may facilitate ligand accommodation. These findings suggest localized bond relaxation, which may contribute to protein flexibility and ligand-induced conformational adjustments. However, no residues exhibited RMSF values exceeding 5 Å, indicating that while certain regions experience dynamic fluctuations, the overall protein structure remains stable without significant unfolding or bond dissociation. This stability reinforces the reliability of the ligand-protein interactions observed in docking and MD simulations.
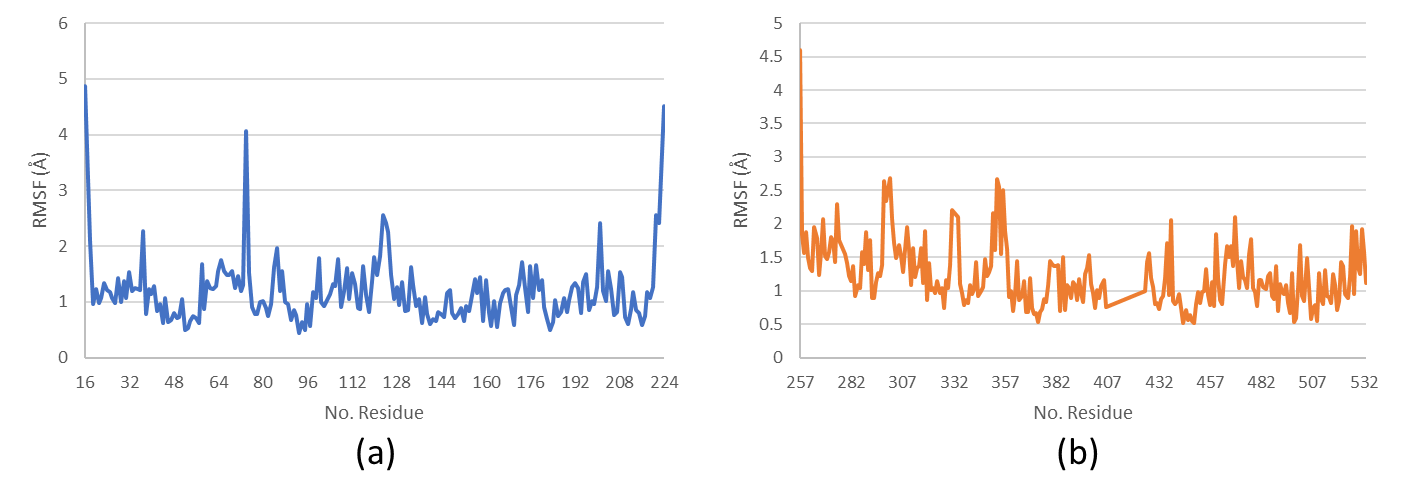
Fig. 12: Root mean square fluctuation (RMSF) analysis of amino acid residues in (a) HSP90AA1-Fucoxanthine and (b) SRC-Alginate complexes, highlighting regions of structural flexibility
DISCUSSION
The study investigates the anticoagulant potential of bioactive compounds derived from Sargassum sp., a genus of brown algae, using network pharmacology, molecular docking, and molecular dynamics simulations. Network pharmacology analysis revealed that several Sargassum-derived compounds, including fucoidan, fucoxanthin, laminarin, phloroglucinol, and alginate, may exert anticoagulant effects by targeting multiple proteins involved in the coagulation pathway. Previous studies have demonstrated that compounds such as fucoidan possess diverse biological activities, including antioxidant, anticoagulant, antithrombotic, and anti-inflammatory effects [24]. In addition, Sargassum has been shown to exert notable anticoagulant effects by binding to positively charged regions of coagulation proteins, thereby inhibiting blood clot formation [25].
The analysis of PPI found three key coagulation-related proteins targeted by Sargassumsp compounds, including SRC, MAPK1, and HSP90AA1. These proteins have been reported as critical regulators of platelet aggregation, endothelial function, and inflammation, all contributing to thrombus formation [26-28]. SRC plays a pivotal role in platelet-mediated coagulation by regulating integrin αIIbβ3 signaling. Through interaction between the SH3 domain of SRC and the C-terminal region of the β3 cytoplasmic tail, SRC becomes activated, facilitating “outside-in” signaling essential for platelet aggregation and spreading—key steps in thrombus formation [29]. Furthermore, MAPK1 lead platelet activation and coagulation by regulating multiple signalling pathways involved in thrombus formation. Its activity promotes platelet aggregation, granule release, αIIbβ3 activation, calcium mobilization, and clot retraction through phosphorylation of downstream targets such as p38, ERK5, JNK, ASK1, talin1, and β3. Deficiency or inhibition of MAPK1 disrupts these processes, resulting in impaired hemostasis and reduced thrombotic potential[30]. On the other hand, HSP90AA1, which functions as a molecular chaperone, may play a role in platelet activation. However, this protein can also trigger the uPA effect, which increases the risk of bleeding. The use of HSP90 inhibitors has been demonstrated to suppress hyperfibrinolysis and help maintain the balance between clot formation and dissolution [31, 32].
The molecular docking and network pharmacology approaches used in this study provide strong initial evidence for the anticoagulant potential of Sargassum sp. compounds. The docking results demonstrate that fucoidan, alginate, and other bioactive compounds from Sargassum sp. show strong binding affinities with key coagulation-related proteins such as SRC, MAPK1, and HSP90AA1. While the docking scores for SRC and MAPK1 were moderate, they were consistent with the idea that the interaction of these compounds with the proteins could lead to the modulation of coagulation pathways. The docking scores for fucoidan and alginate are relatively low, and these findings remain preliminary and hypothesis-generating.
The moderate binding affinity of these compounds toward SRC and MAPK suggests that Sargassum-derived compounds may inhibit coagulation by suppressing αIIbβ3 activation, as well as platelet aggregation, granule release, calcium mobilization, and clot retraction, consistent with the roles of these two proteins in the coagulation process [29, 30]. Meanwhile, their moderate binding potential toward HSP90AA1 also supports this hypothesis. The relatively weak binding observed in this study may represent an interesting finding, as stronger inhibition of HSP90AA1 could have negative effects, given that this protein exhibits dual and nearly opposing roles in cellular processes [31, 32].
Although involving different pathways, the findings of this study support previous reports that Sargassum spp. compounds, including fucoidan, exhibit anticoagulant effects by directly modulating thrombin and factor Xa [33, 34]. In particular, fucoidan, a sulfated polysaccharide, has been shown to inhibit thrombin and factor Xa, both central components of the coagulation cascade [35]. This study extends these findings by proposing that Sargassum sp. compounds may act as selective inhibitors of coagulation-related proteins, thus offering a novel mechanism for anticoagulation therapy.
A significant strength of this study lies in its multi-target approach, where bioactive compounds from Sargassum sp. were found to interact with multiple proteins involved in coagulation, signalling, and inflammation. Such interactions could offer a more comprehensive therapeutic strategy for managing thrombotic diseases. Unlike conventional anticoagulants, such as warfarin and heparin, which primarily target coagulation factors directly involved in thrombus formation, As shown in this study, Sargassum compounds may exert anticoagulant effects by suppressing SRC, MAPK1, and HSP90AA1, thereby inhibiting αIIbβ3 activation, platelet aggregation, granule release, calcium mobilization, and clot retraction. Furthermore, several other studies have reported that these compounds provide additional benefits by modulating inflammatory and endothelial pathways that contribute to thrombosis. This multi-target potential is consistent with the growing interest in polypharmacology, where compounds are designed to interact with multiple targets to improve therapeutic benefits [36, 37]. By targeting both coagulation factors and inflammatory pathways, Sargassum sp. compounds offer an advantage in efficacy and safety, potentially reducing the bleeding risks often associated with conventional therapies.
In molecular dynamic symulation, the RMSD, Rg, and SASA are valuable, it is important to interpret these results cautiously. The gradual decrease in Rg and SASA may suggest conformational changes in the protein-ligand complex. However, such reductions could indicate protein collapse or denaturation rather than proper "stabilization." The decrease in SASA and Rg values over time might represent the structural compaction of the complex; however, this could also signal instability if the structural changes lead to protein misfolding or unfolding. Therefore, it is important to analyze the MD trajectories further to determine whether the complex has reached a stable conformation or if these changes reflect the loss of stability. However, although the molecular dynamics (MD) simulations offer valuable insights into the short-term stability of the protein–ligand complexes,the 20 ns simulation duration provides only limited information on transient stability and does not confirm equilibrium binding or long-term conformational stability of the complexes. The fluctuations in the RMSD values observed in this study, particularly for the SRC-Alginate complex, suggest that the system has not fully equilibrated, and longer simulations are necessary to confirm the stability of these interactions. It is widely accepted in the field that 100 ns or longer simulations are required to properly evaluate the stability of protein-ligand interactions [38, 39].
This study provides new insights into the development of Sargassum sp.-based anticoagulants. The computational models employed offer valuable understanding of the potential mechanisms of action of Sargassum sp. compounds. However, as with any in silico approach, the accuracy of predictions depends heavily on the quality of the databases and the assumptions applied during the modeling process. In this study, molecular docking was performed using standard protocols primarily designed for small molecules, which may not be optimal for large and flexible polysaccharides such as fucoidan and alginate. The conformational flexibility of these macromolecules is not fully captured by conventional docking methods. To address this limitation, future studies are recommended to employ specialized docking approaches, such as the GLYCAM force field, which is specifically developed for modeling polysaccharide–protein interactions. This approach is expected to provide more accurate predictions of the binding mechanisms and anticoagulant activity of Sargassum sp.-derived compounds [40].
While the computational predictions in this study are promising, they must be validated through experimental studies. Computational models, such as molecular docking and molecular dynamics simulations, provide valuable insights into the potential interactions of bioactive compounds with their targets. However, experimental validation is necessary to confirm these interactions and assess their biological relevance. In vitro assays, such as coagulation tests and platelet aggregation studies, would help validate Sargassum sp. compounds' anticoagulant effects in biological systems. Furthermore, in vivo preclinical trials are needed to evaluate these compounds' pharmacokinetics, bioavailability, and long-term safety before clinical application.
However, if these findings are validated through wet-laboratory experiments and confirmed in clinical trials, they could represent a significantly beneficial discovery. When comparing the results of this study with existing anticoagulants, Sargassum sp. compounds show promising advantages. Warfarin, a widely used vitamin K antagonist, has a narrow therapeutic index, requiring frequent monitoring to prevent under-or over-anticoagulation [41]. Similarly, heparin, while effective, carries a risk of bleeding and requires close monitoring. Direct oral anticoagulants (DOACs), such as rivaroxaban and apixaban, offer convenience and predictable pharmacokinetics but still carry bleeding risks, particularly in patients with renal impairment [42, 43]. In contrast, Sargassum sp. compounds may provide a safer alternative by selectively modulating coagulation pathways with fewer adverse effects and a more favorable therapeutic window. This is especially important in clinical settings, where managing bleeding risks is critical.
In addition to the primary focus on coagulation, Sargassum sp. compounds may have other therapeutic properties that could complement their anticoagulant effects. Studies have shown that marine-derived polysaccharides, particularly fucoidan, exhibit anti-inflammatory, antioxidant, and immunomodulatory effects [44]. These properties could further enhance the therapeutic potential of Sargassum sp. compounds, making them suitable for treating thrombotic diseases associated with inflammation and endothelial dysfunction.
Finally, the computational models used in this study may not fully account for the complexity of biological systems. Anticoagulant activity is influenced by numerous physiological factors, including liver metabolism, renal clearance, and feedback mechanisms within the coagulation cascade [45]. These factors may not be captured by the computational models alone. Thus, experimental studies are essential to validate the potential of Sargassum sp. compounds as anticoagulant agents in real-world settings. Further research should focus on integrating computational predictions with experimental validation to bridge the gap between theoretical models and clinical application.
CONCLUSION
In this study, we employed in silico methods to predict the anticoagulant potential of Sargassum compounds, such as alginate and fucoidan, by evaluating their interactions with coagulation-related proteins like SRC and MAPK1. The results suggest that these compounds may modulate coagulation through their interactions with these targets. However, it is important to recognize that these findings are preliminary and hypothesis-generating, and computational predictions, including docking scores and MMPBSA binding free energies, provide valuable insights into potential protein–ligand interactions. Despite these promising results, experimental validation is essential to confirm the anticoagulant effects of Sargassum compounds. Specifically, in vitro clotting assays such as activated partial thromboplastin time (aPTT), prothrombin time (PT), and thrombin time (TT) should be performed to validate the predicted anticoagulant activity. Additionally, in vivo studies are necessary to confirm the clinical potential of these compounds as anticoagulant agents. In conclusion, this computational study provides promising preliminary insights, but further validation through clotting assays and in vivo testing is strongly recommended to confirm the anticoagulant potential of Sargassum compounds.
ACKNOWLEDGEMENT
Gratefully thank to Faculty of Medicine Universitas Palangka Raya as well as our research team in Faculty of Medicine and Faculty of Pharmacy, Universitas Hasanuddin.
FUNDING
Declared none
AUTHORS CONTRIBUTIONS
Conceptualization, Francisca Diana Alexandra and Ysrafil Ysrafil; Data curation, Francisca Diana Alexandra, Marianti A. Manggau and Ysrafil Ysrafil.; Formal analysis, Muh. Nassrum Massi, Yanti Leman.; Funding acquisition, Francisca Diana Alexandra; Investigation, Francisca Diana Alexandra, Marianti A. Manggau, Muh. Nassrum Massi, Yanti Leman, Ysrafil Ysrafil.; Methodology, Francisca Diana Alexandra and Ysrafil Ysrafil; Project administration, Francisca Diana Alexandra, Marianti A. Manggau and Ysrafil Ysrafil.; Resources, Ysrafil Ysrafil and Marianti A. Manggau; Software, Ysrafil Ysrafil and Francisca Diana Alexandra; Supervision, Marianti A. Manggau, Muh. Nassrum Massi, Yanti Leman; Validation,Marianti A. Manggau, Muh. Nassrum Massi, Yanti Leman, and Ysrafil Ysrafil; Visualization, Francisca Diana Alexandra and Ysrafil Ysrafil; Writing—original draft preparation, All authors; writing—review and editing, All authors; All authors revised the manuscript into its final form and given the approval for submission.
CONFLICT OF INTERESTS
There are no confilic interest with this study
REFERENCES
Cairns JA. Efficacy and limitations of warfarin and novel oral anticoagulants with atrial fibrillation. Left atrial appendage closure: mechanical approaches to stroke prevention in atrial fibrillation: Springer; 2016. p. 17-36.
Wang M, Zeraatkar D, Obeda M, Lee M, Garcia C, Nguyen L. Drug-drug interactions with warfarin: a systematic review and meta-analysis. Br J Clin Pharmacol. 2021;87(11):4051-100. doi: 10.1111/bcp.14833, PMID 33769581.
Tan J, Bae S, Segal JB, Zhu J, Alexander GC, Segev DL. Warfarin use and the risk of stroke, bleeding and mortality in older adults on dialysis with incident atrial fibrillation. Nephrology (Carlton). 2019;24(2):234-44. doi: 10.1111/nep.13207, PMID 29219209, PMCID PMC5993567.
Patriota YB, Chaves LL, Gocke EH, Severino P, Soares MF, Soares Sobrinho JL. Applied nanotechnologies in anticoagulant therapy: from anticoagulants to coagulation test performance of drug delivery systems. Appl Nano. 2021;2(2):98-117. doi: 10.3390/applnano2020009.
Shute JK. Heparin low molecular weight heparin and non-anticoagulant derivatives for the treatment of inflammatory lung disease. Pharmaceuticals (Basel, Switzerland). 2023;16(4):584. doi: 10.3390/ph16040584, PMID 37111341, PMCID PMC10141002.
Jackevicius CA, Lu L, Ghaznavi Z, Warner AL. Bleeding risk of direct oral anticoagulants in patients with heart failure and atrial fibrillation. Circ Cardiovasc Qual Outcomes. 2021;14(2):e007230. doi: 10.1161/circoutcomes.120.007230, PMID 33541109.
Welander F, Renlund H, Dimeny E, Holmberg H, Sjalander A. Direct oral anticoagulants versus warfarin in patients with non-valvular atrial fibrillation and CKD G3-G5D. Clin Kidney J. 2023;16(5):835-44. doi: 10.1093/ckj/sfad004, PMID 37151425, PMCID PMC10157779.
Dwivedi R, Pomin VH. Marine antithrombotics. Mar Drugs. 2020;18(10):514. doi: 10.3390/md18100514, PMID 33066214, PMCID PMC7602030.
Jayawardena TU, Sanjeewa KK, Nagahawatta DP, Lee HG, Lu YA, Vaas AP. Anti-inflammatory effects of sulfated polysaccharide from Sargassum swartzii in macrophages via blocking TLR/NF-Κb signal transduction. Mar Drugs. 2020;18(12):601. doi: 10.3390/md18120601, PMID 33260666, PMCID PMC7760840.
Li Y, Zheng Y, Zhang Y, Yang Y, Wang P, Imre B. Brown algae carbohydrates: structures pharmaceutical properties and research challenges. Mar Drugs. 2021;19(11):620. doi: 10.3390/md19110620, PMID 34822491, PMCID PMC8623139.
Sanger G, Wonggo D, Montolalu L, Dotulong V. Pigments constituents phenolic content and antioxidant activity of brown seaweed Sargassum sp. IOP Conf S Earth Environ Sci. 2022;1033(1):012057. doi: 10.1088/1755-1315/1033/1/012057.
Pandian P, Srinivasan NJ. Anticoagulant properties of marine algae: a systematic review. JoRiPS. 2021;12(2):1529‑38.
Saraswati GPE, Giriwono PE, Iskandriati D. Screening of in vitro anti-inflammatory and antioxidant activity of sargassum ilicifolium crude lipid extracts from different coastal areas in Indonesia. Mar Drugs. 2021;19(5):252. doi: 10.3390/md19050252, PMID 33925071, PMCID PMC8146618.
El Kalyoubi SA, Ragab A, Abu Ali OA, Ammar YA, Seadawy MG, Ahmed A. One-pot synthesis and molecular modeling studies of new bioactive spiro-oxindoles based on uracil derivatives as SARS-CoV-2 inhibitors targeting RNA polymerase and spike glycoprotein. Pharmaceuticals (Basel, Switzerland). 2022;15(3):376. doi: 10.3390/ph15030376, PMID 35337173, PMCID PMC8954694.
Gentile F, Agrawal V, Hsing M, Ban F, Norinder U, Gleave ME. Deep docking a deep learning approach for Virtual Screening of Big Chemical Datasets. bioRxiv. 2019:15.877316. doi: 10.1101/2019.12.15.877316.
Jia LY, Cao GY, Li J, Gan L, Li JX, Lan XY. Investigating the pharmacological mechanisms of SheXiang XinTongNing against coronary heart disease based on network pharmacology and experimental evaluation. Front Pharmacol. 2021;12:698981. doi: 10.3389/fphar.2021.698981, PMID 34335263, PMCID PMC8316858.
Wang Y, Yang SH, Zhong K, Jiang T, Zhang M, Kwan HY. Network pharmacology-based strategy for the investigation of the anti-obesity effects of an ethanolic extract of zanthoxylum bungeanum maxim. Front Pharmacol. 2020;11:572387. doi: 10.3389/fphar.2020.572387, PMID 33364948, PMCID PMC7751641.
Alotaibi NM, Alotaibi MO, Alshammari N, Adnan M, Patel M. Network pharmacology combined with molecular docking molecular dynamics and in vitro experimental validation reveals the therapeutic potential of thymus vulgaris L. essential oil (thyme oil) against human breast cancer. ACS Omega. 2023;8(50):48344-59. doi: 10.1021/acsomega.3c07782, PMID 38144096, PMCID PMC10734022.
Syahputra D, Ysrafil Y, Alexandra FD, Praja RK, Fatmaria F, Pratika RA. Network pharmacology combined with molecular docking and molecular dynamics to verify the therapeutic potential of mung beans (Vigna radiata) against prostate cancer. Beni-Suef Univ J Basic Appl Sci. 2024;13(1):100. doi: 10.1186/s43088-024-00552-3.
Nurhasanah N, Fadilah F, Bahtiar A. Prediction of active compounds of muntingia calabura as potential treatment for chronic obstructive pulmonary diseases by network pharmacology integrated with molecular docking. Int J App Pharm. 2023;15(1):274-9. doi: 10.22159/ijap.2023v15i1.46281.
Arwansyah A, Lewa AF, Muliani M, Warnasih S, Mustopa AZ, Arif AR. Molecular recognition of Moringa oleifera active compounds for stunted growth prevention using network pharmacology and molecular modeling approach. ACS Omega. 2023;8(46):44121-38. doi: 10.1021/acsomega.3c06379, PMID 38027368, PMCID PMC10666129.
Fitriana M, Mun’im A, Firdayani WA, Adikusuma W. Exploration of the active compounds of Moringa Oleifera lam as Hiv-1 reverse transcriptase inhibitor: a network pharmacology and molecular docking approach. Int J App Pharm. 2024;16(2):237-46. doi: 10.22159/ijap.2024v16i2.49855.
Zhang L, Wang P, Yang Z, Du F, Li Z, Wu C. Molecular dynamics simulation exploration of the interaction between curcumin and myosin combined with the results of spectroscopy techniques. Food Hydrocoll. 2020;101:105455. doi: 10.1016/j.foodhyd.2019.105455.
Wang Y, Xing M, Cao Q, Ji A, Liang H, Song S. Biological activities of fucoidan and the factors mediating its therapeutic effects: a review of recent studies. Mar Drugs. 2019;17(3):183. doi: 10.3390/md17030183, PMID 30897733, PMCID PMC6471298.
George AM, Chakraborty K, Paulose SK, Bose C, Dhara S. Anti-coagulant effects of α-(1 → 3)/(1 → 4) linked sulfated galactofucan from brown macroalga Sargassum plagiophyllum (C. Agardh) in human umbilical vein endothelial cells. Food Biosci. 2024;62:105467. doi: 10.1016/j.fbio.2024.105467.
Guo F, Yang R, Xiong X, Yuan Y, Zhang L, Hua C. Taxifolin inhibits platelet activation and thrombosis by regulating the PI3K/Akt and MAPK signaling pathways. Mol Nutr Food Res. 2025;69(16):e70129. doi: 10.1002/mnfr.70129, PMID 40635239.
Li J, Tao Q, Xie Y, Wang P, Jin R, Huang X. Exploring the targets and molecular mechanisms of thalidomide in the treatment of ulcerative colitis: network pharmacology and experimental validation. Curr Pharm Des. 2023;29(34):2721-37. doi: 10.2174/0113816128272502231101114727, PMID 37961863.
Zan X, Zhang S, Liu Q, Wang F, Tian M, Cao Y. Network pharmacology molecular docking and in vivo study on oleanolic acid against psoriasis. Front Med. 2025;12:1616886. doi: 10.3389/fmed.2025.1616886, PMID 41122516, PMCID PMC12535998.
De Kock L, Freson K. The (patho) biology of SRC kinase in platelets and megakaryocytes. Med (Kaunas Lith). 2020;56(12):633. doi: 10.3390/medicina56120633, PMID 33255186, PMCID PMC7759910.
Xu X, Dai Y, Zhu H, Zhang J, Li Y, Jiang H. Mitogen-activated protein kinase 1 regulates platelet function and thrombus formation. J Thromb Haemost. 2025;23(9):2944-57. doi: 10.1016/j.jtha.2025.05.022, PMID 40447106.
Maguire PB, Parsons ME, Szklanna PB, Zdanyte M, Münzer P, Chatterjee M. Comparative platelet releasate proteomic profiling of acute coronary syndrome versus stable coronary artery disease. Front Cardiovasc Med. 2020;7:101. doi: 10.3389/fcvm.2020.00101, PMID 32671099, PMCID PMC7328343.
Li Y, Luo L, Wu J, Wu Z, Jiang H, Chen Z. IL-1β induce CAR-T-associated coagulopathy via the Hsp90/kallikrein/uPA pathway. Blood. 2024;144(Suppl 1):1197. doi: 10.1182/blood-2024-202845.
Hardiningtyas SD, Jauharah N, Tarman K. Characteristics of bioactive components in fermented Sargassum and Ulva Seaweed using SCOBY as potential anti-diabetic candidates. BIO Web Conf. 2024;106:02013. doi: 10.1051/bioconf/202410602013.
Hidayati JR, Yudiati E, Pringgenies D, Arifin Z, Oktaviyanti DT. Antioxidant activities total phenolic compound and pigment contents of tropical Sargassum sp. extract macerated in different solvents polarity. J Kelautan Tropis. 2019;22(1):73-80. doi: 10.14710/jkt.v22i1.4404.
Tanna B, Mishra A. Nutraceutical potential of seaweed polysaccharides: structure, bioactivity safety and toxicity. Compr Rev Food Sci Food Saf. 2019;18(3):817-31. doi: 10.1111/1541-4337.12441, PMID 33336929.
Xiong Z, Yang H. Unraveling the pharmacological intricacies of Kurarinone in osteoporosis: insights from a network pharmacology approach. Pharmacogn Mag. 2024;20(4):1350-8. doi: 10.1177/09731296241259563.
Kampen S, Duy Vo D, Zhang X, Panel N, Yang Y, Jaiteh M. Structure-guided design of G-protein-coupled receptor polypharmacology. Angew Chem Int Ed Engl. 2021;60(33):18022-30. doi: 10.1002/anie.202101478, PMID 33904641, PMCID PMC8456950.
Mohan A, Rendine N, Mohammed MK, Jeeva A, Ji HF, Talluri VR. Structure-based virtual screening in silico docking ADME properties prediction and molecular dynamics studies for the identification of potential inhibitors against SARS-CoV-2 Mpro. Mol Divers. 2022;26(3):1645-61. doi: 10.1007/s11030-021-10298-0, PMID 34480682, PMCID PMC8417657.
Pavan M, Menin S, Bassani D, Sturlese M, Moro S. Qualitative estimation of protein-ligand complex stability through thermal titration molecular dynamics simulations. J Chem Inf Model. 2022;62(22):5715-28. doi: 10.1021/acs.jcim.2c00995, PMID 36315402, PMCID PMC9709921.
Nieto Fabregat F, Lenza MP, Marseglia A, Di Carluccio C, Molinaro A, Silipo A. Computational toolbox for the analysis of protein-glycan interactions. Beilstein J Org Chem. 2024;20:2084-107. doi: 10.3762/bjoc.20.180, PMID 39189002, PMCID PMC11346309.
Bosch FT, Nisio MD, Buller HR, Van Es N. Diagnostic and therapeutic management of upper extremity deep vein thrombosis. J Clin Med. 2020;9(7):2069. doi: 10.3390/jcm9072069, PMID 32630244, PMCID PMC7408847.
Kao TW, Chen ZW, Lin YH. Anticoagulation for patients with concomitant atrial fibrillation and end-stage renal disease: a systematic review and network meta-analysis. J Am Heart Assoc. 2024;13(8):e034176. doi: 10.1161/jaha.123.034176, PMID 38606775, PMCID PMC11262503.
Stanifer JW, Pokorney SD, Chertow GM, Hohnloser SH, Wojdyla DM, Garonzik S. Apixaban versus warfarin in patients with atrial fibrillation and advanced chronic kidney disease. Circulation. 2020;141(17):1384-92. doi: 10.1161/circulationaha.119.044059, PMID 32160801.
Manggau M, Kasim S, Fitri N, Aulia NS, Agustiani AN, Raihan M. Antioxidant anti-inflammatory and anticoagulant activities of sulfate polysaccharide isolate from brown alga Sargassum policystum. IOP Conf S Earth Environ Sci. 2022;967(1):012029. doi: 10.1088/1755-1315/967/1/012029.
Wu YW, Ta GH, Lung YC, Weng CF, Leong MK. In silico prediction of skin permeability using a two-QSAR approach. Pharmaceutics. 2022;14(5):961. doi: 10.3390/pharmaceutics14050961, PMID 35631545, PMCID PMC9143389.