
Int J App Pharm, Vol 18, Issue 2, 2026, 398-409Original Article
DESIGN, OPTIMIZATION AND IN SILICO STUDIES OF FOLIC ACID-CONJUGATED DOCETAXEL-LOADED LIPOSOMES FOR TARGETED DELIVERY TO A549 LUNG ADENOCARCINOMA CELLS
AYUSHI PRADHAN1, GURUDUTTA PATTNAIK1*, CH NIRANJAN PATRA2, DIBYALOCHAN MOHANTY3, GNYANA RANJAN PARIDA1, MANI SHARMA3
1School of Pharmacy and Life Sciences, Centurion University of Technology and Management, Bhubaneswar, Odisha, India. 2Department of Pharmaceutics, Roland Institute of Pharmaceutical Sciences, Berhampur, Ganjam, Odisha, India. 3Department of Pharmaceutics, School of Pharmacy, Anurag University, Hyderabad, Telangana-500088, India
*Corresponding author: Gurudutta Pattnaik; *Email: gurudutta.pattnaik@cutm.ac.in
Received: 05 Nov 2025, Revised and Accepted: 09 Jan 2026
ABSTRACT
Objective: This study aimed to develop and optimize a folic acid-conjugated liposomal formulation of docetaxel (FA-Opt-DTXL) for enhanced delivery and cytotoxic efficacy against folate receptor–overexpressing A549 lung adenocarcinoma cells.
Methods: A Box–Behnken design (BBD) coupled with response surface methodology was employed to optimize critical formulation parameter soya lecithin quantity, cholesterol content, and sonication time, affecting liposomal characteristics. The optimized docetaxel-loaded liposomes (Opt-DTXL) were characterized for particle size, entrapment efficiency, and zeta potential. FA–PEG–Cholesterol conjugate was synthesized via NHS-activated ester coupling and incorporated into preformed liposomes using the post-insertion method to yield FA-Opt-DTXL. Morphological analysis was performed using TEM, and cytotoxicity was assessed via IC₅₀ determination and fluorescence-based apoptosis assays.
Results: The optimized Opt-DTXL formulation exhibited a particle size of 239.60 nm, entrapment efficiency of 84.03%, and zeta potential of –22.85 mV, closely matching predicted values. TEM imaging confirmed spherical, well-defined vesicles. Compared to Pure-DXT and Opt-DTXL, FA-Opt-DTXL demonstrated superior cytotoxicity, with an IC₅₀ of 16.67 µg/ml. Apoptotic assays revealed dominant red fluorescence and chromatin fragmentation in FA-Opt-DTXL–treated cells, indicating enhanced receptor-mediated uptake and intracellular drug accumulation. The integration of statistical optimization and Network pharmacology and folate-targeted surface modification yielded a potent nanocarrier system with improved cytotoxicity against A549 lung adenocarcinoma cells.
Conclusion: FA-Opt-DTXL represents a promising platform for site-specific docetaxel delivery in folate receptor–positive malignancies, offering translational potential for targeted cancer therapy, also supported by network pharmacology and molecular docking studies.
Keywords: Docetaxel, Liposome, A549 lung adenocarcinoma cells, Folic acid, NHS
© 2026 The Authors.Published by Innovare Academic Sciences Pvt Ltd. This is an open access article under the CC BY license (https://creativecommons.org/licenses/by/4.0/)
DOI: https://dx.doi.org/10.22159/ijap.2026v18i2.57010 Journal homepage: https://innovareacademics.in/journals/index.php/ijap
INTRODUCTION
Lung adenocarcinoma, a major subtype of non-small cell lung carcinoma (NSCLC), originates from glandular epithelial cells lining the alveoli and bronchioles [1]. It is characterized by abnormal proliferation of these cells, often driven by genetic mutations such as EGFR (Epidermal Growth Factor Receptor) and KRAS (Kirsten Rat Sarcoma Viral Oncogene Homolog) rearrangements [2]. These mutations disrupt normal cell signalling pathways-particularly those regulating growth, apoptosis, and angiogenesis leading to uncontrolled tumour expansion and metastasis. Unlike squamous cell carcinoma, adenocarcinoma frequently arises in the peripheral regions of the lung and is more common in non-smokers and females. Patho physiologically, the tumour microenvironment plays a critical role in disease progression [3]. Hypoxia, chronic inflammation, and immune evasion mechanisms such as PD-L1 overexpression contribute to tumor survival and resistance to therapy [4]. As the tumor grows, it compromises pulmonary function, leading to symptoms like dyspnea, persistent cough, hemoptysis, and chest pain. Advanced stage disease often involves metastasis to the brain, bones, liver, and adrenal glands, further complicating treatment and prognosis. Despite therapeutic advances, including targeted therapies and immunotherapy, the overall survival rate remains low, underscoring the need for improved drug delivery systems and early diagnostic strategies
Docetaxel, a second-generation taxane, has shown substantial cytotoxic activity against NSCLC by stabilizing microtubules and disrupting mitotic spindle formation, thereby inducing apoptosis in rapidly dividing tumorcells [5]. However, its clinical application is significantly hindered by several pharmacokinetic and pharmacodynamic limitations. These include poor aqueous solubility, which necessitates the use of toxic solubilizing agents; non-specific biodistribution, leading to off-target effects; and systemic toxicity, manifesting as neutropenia, hypersensitivity reactions, and gastrointestinal disturbances [6]. Collectively, these drawbacks reduce the therapeutic index of docetaxel and compromise patient outcomes.
Docetaxel loaded nano delivery systems including nanocapsules [7], nanomicelles [8], dendrimers [9], solid lipid nanoparticles [10], SNEDDS [11] and electrospunnanofibers [12] have been extensively investigated to address the drug’s poor solubility, systemic toxicity, and limited tumor penetration. Among these, liposomal formulations have emerged as the most clinically favourable platform due to their exceptional biocompatibility, high drug encapsulation efficiency, and ability to prolong systemic circulation. Liposomes not only enhance the bioavailability of hydrophobic agents like docetaxel but also facilitate preferential tumor accumulation via the enhanced permeability and retention (EPR) effect [13]. Compared to other nanocarriers, liposomes offer superior control over drug release kinetics, reduced immunogenicity, and scalability for clinical translation [14]. Their versatility in surface modification, stability under lyophilization, and proven efficacy in preclinical and early clinical models position liposomal systems as the leading strategy for optimized docetaxel delivery.
Folate Receptor α (FR-α) is a high-affinity, glycosylphosphatidylinositol-anchored membrane protein that mediates cellular folate uptake. Upon binding folic acid, the receptor–ligand complex undergoes receptor-mediated endocytosis, facilitating selective internalization of FA-decorated nanocarriers [15]. This mechanism enables enhanced intracellular drug delivery, bypasses efflux-mediated resistance, and provides tumor selectivity due to the minimal FR-α expression in most normal tissues [16, 17].
The present study aims to develop and optimize a novel docetaxel-loaded liposomal lyophilized powder using response surface methodology to enhance targeted cytotoxicity against A549 cells by in vitro and also showcasing the efficacy and binding capability of docetaxel and folic acid against few receptors and genes shortlisted and analyzed through network pharmacology and molecular docking studies. Addressing the limitations of conventional docetaxel formulations, it integrates nanotechnology and statistical modeling to deliver a stable, reconstitutable system with improved tumor selectivity and reduced systemic toxicity.
MATERIALS AND METHODS
Materials
Docetaxelwasa gift sample from Alembic Research Centre, Gujurat, India. Soya lecithin, Cholesterol and Chloroform were purchased from S. D Fine Mumbai, India Ultrapure water was prepared by Milli-Q purification system (Millipore Corp., Bedford, MA, USA).
Preparation method docetaxel loaded liposome
Liposomes were prepared by the thin film hydration method (TFH) [18]. The 10 mg Docetaxel constant for all batches and the required quantities of Soya lecithin and cholesterol were taken in a 100 ml round bottom flask and dissolved in 10 ml chloroform. All the batches were prepared according to the experimental design. Chloroform was evaporated using rotary vacuum evaporator and kept overnight under vacuum. Then it was hydrated by 15 ml of phosphate buffer pH 7.4 for 1 h with 10 min of extensive vertexing. The suspension of liposomes was subjected to probe sonication for 10 min in the water bath at 60 °C to reduce the size of liposomes. Final liposome dispersion was subjected to freeze drying or lyophilization at-80 °C for 24 h. Final liposome dried powder was subjected to further characterization study.
Application of experimental design
A Box–Behnken design (BBD) was employed to optimize the formulation parameters of Docetaxel-loaded liposomes using response surface methodology [19]. Three independent variables soya lecithin concentration (mg), cholesterol content (mg), and sonication time (min) were selected as critical formulation independent factors based on their influence on liposomal characteristics. The dependent variables evaluated included particle size (nm), entrapment efficiency (%), and zeta potential (mV), which collectively determine the physicochemical performance of the liposomal system. Design Expert software (version 12, Stat-Ease Inc., Minneapolis, MN, USA) was utilized to construct the experimental matrix comprising 17 trial formulations, including five replicates at the center point to ensure model robustness. Mathematical modelling, statistical analysis, and model validation were conducted to establish predictive equations and assess the significance of factor–response relationships. Model adequacy was confirmed based on p-values, correlation coefficients (R²), and predicted residual error sum of squares (PRESS). Optimization of the formulation was achieved through a numerical desirability function, identifying the most suitable combination of input variables to meet predefined response criteria [20]. Model validation was performed using checkpoint formulations, wherein predicted values were compared against experimental outcomes. A prediction error within ±5% was considered acceptable, confirming the reliability and predictive capability of the established model.
Formulation of FA cojugated Opt-DTXL (FA-Opt-DTXL)
Folic acid (10 mg) was first dissolved in 2 ml of dimethyl sulfoxide (DMSO) and activated using N-hydroxysuccinimide (NHS, 3 mg). The activated folic acid was then reacted with PEG6000 (60 mg) and cholesterol (15 mg), which had been co-dissolved in 2 ml of chloroform. The reaction was carried out under a nitrogen atmosphere with continuous stirring at room temperature for 12–16 h to facilitate the formation of the FA–PEG–Cholesterol conjugate. The resulting product was subsequently lyophilized by adding sucrose as cryoprotectant for use in liposomal functionalization. For targeted incorporation, the FA–PEG–Cholesterol conjugate (typically prepared at 2 mg/ml in phosphate-buffered saline, PBS) was added dropwise to 10 ml of preformed liposomal dispersion maintained at 60 °C, a temperature above the lipid phase transition threshold. The mixture was gently stirred for 30–60 min to allow spontaneous insertion of the conjugate into the liposomal bilayer, thereby imparting folate receptor-targeting capability to the formulation.
Table 1: Application of box-behnken design to develop drug-loaded liposome
| Independent variables (X) | Coded value | ||
| Low (-1) | Mid (0) | High (+1) | |
| Soya lecithin(X1) (mg) | 150 | 200 | 250 |
| Cholesterol (X2) (mg) | 20 | 25 | 30 |
| Sonication time (X3)(min) | 3 | 5 | 7 |
| Responses (Y) | Constraint | ||
Particle size (Y1) Entrapment efficiency (Y2) Zeta potential (Y3) |
Minimize Maximize Between-20 to-30 |
||
Table 2: Application of box-behnken design to develop drug-loaded liposome
| Formulation | X1:Soyalecithin | X2:Cholesterol | X3:Sonication time (min) | Y1:Particle size (nm) | EE% | Zeta potential(mV) |
| DTXL1 | 250 | 30 | 5 | 252±11.1 | 92±1.6 | -18.01±0.3 |
| DTXL2 | 200 | 25 | 5 | 238±13.7 | 85±1.2 | -22.12±0.2 |
| DTXL3 | 200 | 25 | 5 | 239±12.6 | 85±1.3 | -22.17±0.1 |
| DTXL4 | 200 | 30 | 7 | 241±11.4 | 87±1.7 | -19.12±0.5 |
| DTXL5 | 200 | 20 | 7 | 235±13.1 | 83±1.1 | -26.01±0.3 |
| DTXL6 | 200 | 20 | 3 | 237±11.6 | 83±1.8 | -28.96±0.4 |
| DTXL7 | 150 | 25 | 7 | 227±11.4 | 78±1.6 | -22.14±0.6 |
| DTXL8 | 150 | 25 | 3 | 229±13.7 | 78±1.5 | -22.37±0.8 |
| DTXL9 | 200 | 30 | 3 | 232±11.7 | 87±1.4 | -18.51±0.7 |
| DTXL10 | 200 | 25 | 5 | 241±13.1 | 85±1.3 | -22.14±0.1 |
| DTXL11 | 250 | 25 | 3 | 251±10.5 | 90±1.2 | -22.18±0.2 |
| DTXL12 | 200 | 25 | 5 | 240±11.6 | 85±1.4 | -22.09±0.6 |
| DTXL13 | 200 | 25 | 5 | 237±12.6 | 85±1.6 | -22.14±0.3 |
| DTXL14 | 250 | 25 | 7 | 251±13.1 | 90±1.3 | -22.04±0.2 |
| DTXL15 | 150 | 20 | 5 | 225±13.4 | 75±1.2 | -26.51±0.4 |
| DTXL16 | 250 | 20 | 5 | 246±11.4 | 88±1.1 | -26.04±0.3 |
| DTXL17 | 150 | 30 | 5 | 222±12.3 | 80±1.5 | -18.56±0.2 |
Results are given in mean± SD, n=3
Characterization of docetaxel-loaded liposome
Particle size and zeta potential
The average particle size and particle size distribution were measured by the light dynamic scattering (DLS) method using Delsa™ Nano Submicron Particle Size Analyzer (A53878, Beckman Coulter Inc., Indianapolis, IN, USA). The Zeta potential was measured by the laser doppler anemometry (LDA) on zeta Plus Zeta Potential Analyzer (Brookheaven Instruments Corporation) [21].
Entrapment efficiency (%EE)
The encapsulation efficiency of various docetaxel-loaded liposomal formulations was evaluated by quantifying the unencapsulated drug present in the dispersion medium. 3 mL aliquot of each liposomal dispersion was transferred into a centrifuge tube and subjected to centrifugation at 10,000 rpm for 30 min. Post-centrifugation, the supernatant was carefully collected and filtered using Whatman filter paper to remove residual particulates. The filtered supernatant was subsequently washed, appropriately diluted, and analyzed using a UV–visible spectrophotometer (Model 1700, Shimadzu) to determine the concentration of free docetaxel. The encapsulation efficiency (EE%) of the nanoparticles was calculated using the standard equation referenced in [22]. The resulting values for each formulation are presented below.
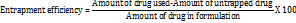
Morphological study by transmission electron microscopy (TEM)
Using TEM (JEOL 100CX; JEOL Inc., Peabody, MA, USA) at an accelerating voltage of 80 kV, the morphology of the Optimized-Docetaxel-loaded liposome (Opt-DTXL) was examined. The Opt-DTXL formulation was diluted from 1 to 0.01% w/w and treated in an ultrasonic bath (Model 3510, Branson MS) in order to reduce particle aggregation on the copper grid. For TEM analysis, a single drop of the modified mixture was dried and put on a copper grid coated with carbon [23].
In vitro release study
The Docetaxel release from the pure liposome suspension and Opt-DTXL were evaluated by a dialysis bag method using buffers with pH 6.8. The dialysis bags, with pure Docetaxel or liposomes containing DTX, were immersed in buffers (pH 6.8) containing 0.5% Tween 80 (dissolution medium) and were stirred (50 r/min) at 37 °C. At different time intervals up to 24 h, the sample was taken out and the fresh PBS was replaced correspondingly. The drug content in the sample was analysed by UV method.
MTT assay procedure
The NCCS in Pune, India, providedA549 lung adenocarcinoma cells. The cells were kept in DMEM media with high glucose supplemented with 10% FBS and 1% antibiotic-antimycotic solution in a CO2 incubator with 5% CO2, 18% O2, and 37 °C. They were subcultured every two days. Cell viability was evaluated using the MTT test, and the cell inhibitory rate was calculated using the formula below [24]. A549 cells were cultured for 24 h after being treated with Pure-Docetaxel, Opt-DTXL and FA-Opt-DTXL at Concentration of 0.1,1,10,25,50,75 and 100 µg/ml. All MTT assays were performed in triplicate (n=3) across three independent experiments to ensure reproducibility and statistical reliability. IC₅₀ values were determined using non-linear regression analysis the sigmoidal dose-response curve produced by GraphPad Prism was used to calculate the IC50. The following formula was used to determine the percentage of cell viability:
% cell viability=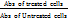 × 100
× 100
Apoptosis study against A549 lung adenocarcinoma cells
Acridine orange and ethidium bromide double staining
Acridine orange (AO) and ethidium bromide (EB) double staining is a fluorescence-based technique employed to distinguish viable, apoptotic, and necrotic cells. AO, a cell-permeable dye, stains both live and dead cells, emitting green fluorescence upon binding to double-stranded DNA in viable cells. In contrast, EB penetrates only cells with compromised membranes, binding to DNA and emitting red fluorescence. This dual-staining approach enables clear differentiation: viable cells appear green, early apoptotic cells exhibit yellow-green fluorescence with condensed or fragmented chromatin, and necrotic or late apoptotic cells fluoresce red [25]. For microscopic analysis, A549 lung adenocarcinoma cells were seeded at a density of 1 × 10⁵ cells/ml in 6-well or 24-well plates. Following 24 h treatment with Pure-Docetaxel, Opt-DTXL, and FA-Opt-DTXL at their respective IC₅₀ concentrations, cells were fixed using 4% paraformaldehyde and stained with 10 μl of a 1 mg/ml AO/EB mixture per well. Fluorescence images were captured using a Nikon Eclipse Ti2 microscope (Japan) at 20× magnification and subsequently processed using ImageJ Software v1.48.
Network pharmacology and molecular docking
Software and web servers used to perform molecular docking, various tools were employed for evaluating binding affinities and ligand–receptor interactions. For network pharmacology analysis, databases and platforms like STRING, STITCH, Swiss Target Prediction, and Cytoscape were used to predict, visualize, and analyze protein–protein interactions and target networks associated with selected bioactive compounds. All software and ebsites are enlisted in table 3.
Table 3: Detail list of software’s and web servers used in molecular docking with their links
| S. No. | Software | Link/Webserver |
| 1. | CHEBI | https://pubchem.ncbi.nlm.nih.gov/ |
| 2. | PubChem data-base | https://www.molsoft.com/mprop/ |
| 3. | Molsoft L. L. | https://www.molsoft.com/mprop/ |
| 4. | Swiss target prediction | http://www.swisstargetprediction.ch/ |
| 5. | DisGeNET | https://www.disgenet.org/search |
| 6. | Therapeutic Target Database (TTD) | https://idrblab.net/ttd/ |
| 7. | venny (2.1.0) | https://bioinfogp.cnb.csic.es/tools/venny/ |
| 8. | string | https://string-db.org/ |
| 9. | Cytoscape (Version: 3.10.2) | https://cytoscape.org/ |
| 10. | DAVID database | https://david.ncifcrf.gov/tools.jsp |
| 11. | Protein Data Bank | https://www.rcsb.org/ |
Identification of active components and their targets collection
The key moieties of Docetaxel and Folic acid were sourced from IMPPAT (Indian Medici-nal Plant, Phytochemistry and Therapeutics, https://cb.imsc.res.in/imppat/) [26] as well as CHEBI (Chemical Entities of Biological Intrest) (European Molecular Biology Laboratory 2018) database, which cata-logs chemical entities of biological interest. Subsequently, the canonical smiles reprentations of these compounds were retrived from the PubChem database (https://pubchem.ncbi.nlm.nih.gov/). Further, Molsoft L. L. (https://www.molsoft.com/mprop/) [27] was used for drug likeness and molecular property prediction. The compounds were then shortlisted based on their drug likeness score with positive readings. The drug likeness score is taken on the bases of positive readings (+), which are included, andthe negative readings are excluded. Because the closer the score is to 1, the more drug-like the molecule. Swiss target prediction (http://www.swisstargetprediction.ch/) and SEA (https://sea.bkslab.org/) are employed for target prediction [28].
Collection of disease targets
Targets for the lung cancer were gathered from various data-bases. Like DisGeNET (https://www.disgenet.org/search) in that specifically, diabetes-related targets, including those for insulin dependence, were identified [29] Additionally, targets were sourced from the Therapeutic Target Database (TTD, https://idrblab.net/ttd/)[30].
Common targets
By using venny 2.1.0 (https://bioinfogp.cnb.csic.es/tools/venny/) to evaluate common targets from disease targets and compound targets, the common targets were identified. In this the list of disease targets is filled in one box, naming dis-ease tar and the list of compound targets is filled in another box naming com tar, from this analysis, resulting in the creation of a venn diagram fig. 10b [31].
Protein–protein interaction network
The string (https://string-db.org/) platform was utilized for protein-protein interaction analysis. The list of common targets was pasted into the platform and filtered for Homo sapiens, and in advance settings, the conditions applied are, for Network type:-full STRING network, for Required score:-medium confidence (0.400) and for FDR stringency:-medium (5%) and click for search, resulting in the generation of the image structure [32].
Network construction
Cytoscape is employed to construct a network. In the net-work section, navigate to the string subsection and paste the common targets obtained from venny. Then, in the cyto-Hubba bar, choose calculate, followed by top, then enter 10, then degree, and select to display the shortest path before submitting. This will provide the top 10 genes for the path-ways [33].
Compound-gene network construction
Cytoscape (Version: 3.10.2 https://cytoscape.org/) is utilized to construct the network diagram by inserting the excel tables of columns, namely source, targets, and attribution. The active components and top 10 genes is constructed.
Molecular docking
Molecular docking can be utilized to validate the key targets in network pharmacology. Firstly search the shortlisted compounds in the PubChem database, and 3D structures of those compounds were downloaded. Next, the 3D structures of the top 10 proteins were obtained from the Protein Data Bank (https://www.rcsb.org/). Specifically the human models are selected, then the BIOVIA discovery studio is utilized to convert the downloaded files into PDB format. Next, the docking process was carried out in PyRx [34]. Software and the docking results were obtained. Then the docking structures were obtained by utilizing the BIOVIA Discovery studio software and both 3D and 2D structures are saved [35].
RESULTS AND DISCUSSION
Particle size
The particle size of liposomes is a critical parameter that directly influences their stability, drug delivery efficiency, biodistribution, and therapeutic performance. Particle size found in a range of 222±12.3 nm to 252±11.1 nm (table 2). The polynomial equation presented in table 3 reveals that particle size was significantly influenced by the independent variables and their interactions
Y1=+239.46+11.93X1+2.23X2-0.8250X3+2.25X1X2+0.6000X1X3+6.20X2X3+2.02 X12 – 4.93 X22-1.48X32
Table 3 presents the quadratic model analysis, revealing a statistically significant response with an F-value of 36.43 and a P-value less than 0.0001.
An increase in the quantity of soya lecithin (X1) leads to a significant rise in liposome particle size due to enhanced bilayer formation and lipid packing. Higher lecithin content contributes to the development of thicker or multilamellar vesicles, which expand the overall vesicle diameter. Additionally, increased lipid concentration elevates the viscosity of the dispersion, reducing the efficiency of size-reduction techniques such as sonication, thereby yielding larger particles. This trend is supported by Dwiastuti et al. in 2016 [36] who observed that higher concentrations of soy lecithin phospholipids led to larger liposomal vesicles. Cholesterol quantity (X2) exerts a synergistic influence on liposome particle size by modulating bilayer rigidity and structural dynamics. At elevated levels, cholesterol integrates into the phospholipid bilayer, enhancing membrane stability and reducing fluidity through tighter lipid packing. This synergistic stabilization effect limits the extent of vesicle size reduction during sonication or extrusion, resulting in larger liposomal particles. The interaction between cholesterol and phospholipids not only reinforces bilayer integrity but also imparts resistance to mechanical stress, thereby preserving vesicle architecture. Similar findings were reported by Briuglia et al. in 2015 [37]. Probe sonication time (X3) plays a critical role in reducing liposome particle size by delivering high-energy acoustic waves directly into the lipid dispersion. This mechanical energy disrupts larger vesicles and promotes the formation of smaller, more uniform liposomes through cavitation and shear forces, similar findings reported by Yun et al. in 2023 [38] (fig. 1).
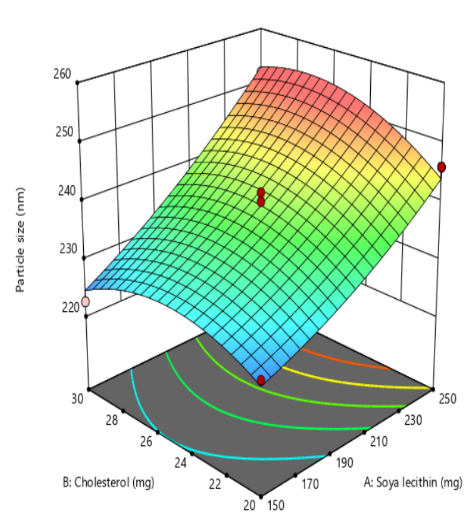
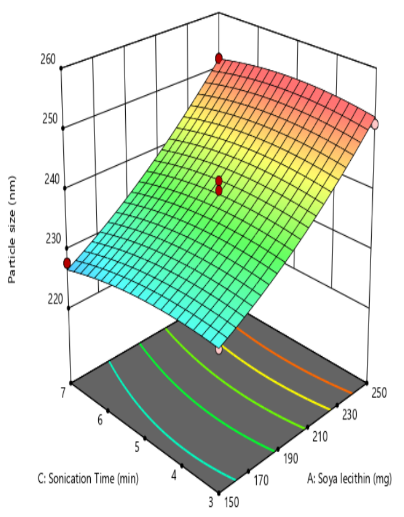
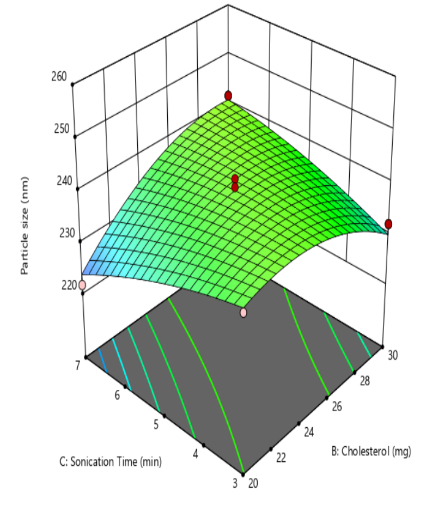
Fig. 1: Effect of independent variable on particle size
Entrapment efficiency analysis
The encapsulation efficiency (%EE) in liposome loading Docetaxel significantly influences drug loading capacity and release kinetics. Entrapment efficiency (EE%) for the 17 experimental formulations ranged from 75±1.2% to 92±1.6%, as detailed in table 2. The quadratic model derived from the Box–Behnken design provided a predictive equation for EE%, incorporating the influence of formulation variables. This equation enables the identification of optimal conditions for achieving high entrapment efficiency and supports the rational design of nanoparticle-based drug delivery systems.
Y2=+85.06+6.13X1+2.13X2+0.0001X3-0.2500X1X2+0.0000X1X3+0.0000X2X3-1.15X12-0.1550X22+0.0950X32
The quadratic model analysis for entrapment efficiency (EE%) presented in table 3 yield an F-value of 826.85 and a p-value<0.0001, indicating a statistically significant model fit. The quadratic model revealed a statistically significant and synergistic relationship between soya lecithin quantity (X1) and entrapment efficiency increasing lecithin levels enhances drug encapsulation, This effect is attributed to lecithin’s amphiphilic nature and its ability to form stable bilayer vesicles that serve as effective carriers for both hydrophilic and hydrophobic drugs. Higher lecithin levels provide a greater lipid matrix, facilitating improved drug partitioning into the bilayer and reducing leakage from the vesicular system. Additionally, the structural integrity and packing density of the lipid membrane are reinforced at elevated lecithin concentrations, resulting in more efficient entrapment of the drug. Comparable findings were reported by Ahmed et al. (2023) [39], who demonstrated that increasing soya lecithin concentration significantly improved drug encapsulation efficiency (fig. 2). The amount of cholesterol (X₁) plays a critical role in modulating the physicochemical properties of liposomal bilayers, exerting a synergistic influence on encapsulation efficiency (EE%). By enhancing bilayer rigidity and reducing membrane permeability, cholesterol effectively minimizes drug leakage and promotes retention of the encapsulated payload. Recent findings by Fukuoka et al. (2025) demonstrated that cholesterol-containing liposomes exhibited reduced permeability and enhanced structural integrity, reinforcing its role in improving encapsulation performance [40].
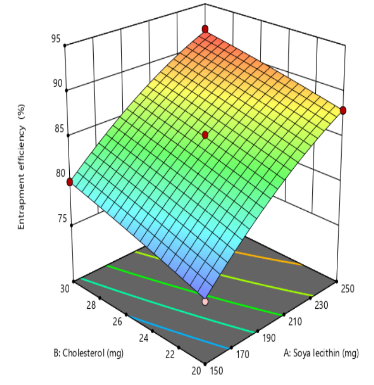
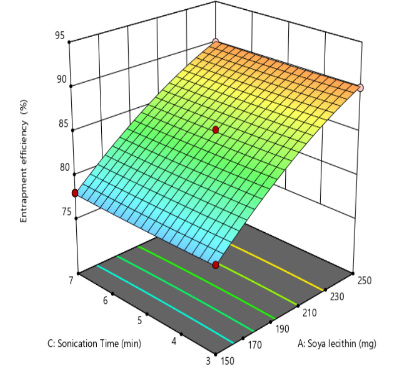
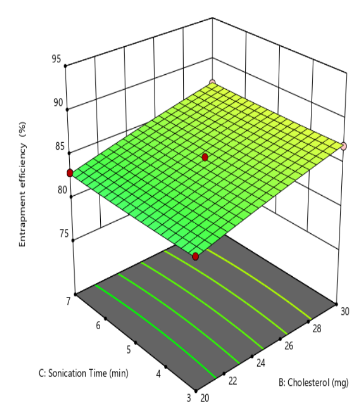
Fig. 2: Effect of independent variable on entrapment efficiency
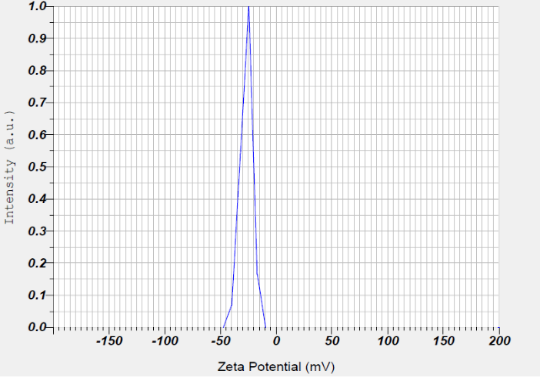
Zeta potential
Zeta potential reflects the electrostatic potential at the slipping plane of liposomal vesicles in suspension and serves as a key indicator of colloidal stability. A higher magnitude of zeta potential, whether strongly positive or negative, generates greater electrostatic repulsion between liposomes, thereby minimizing vesicle aggregation and enhancing formulation stability [41]. The zeta potential values obtained ranged from-18.01±0.3 mV to-28.96 ±0.4 mV, as presented in table 2. The corresponding quadratic equation describing the zeta potential response is as follows
Y3=-22.28+0.1550X1+4.17X2+0.3375X3+0.0075X1X2-0.0225X1X3-0.8925X2X3+0.4903X12-0.4748X22-0.3948X32
The Quadratic model (table 3) for zeta potential demonstrated a significant response, with an F-value of 79.51 and a P-value less than 0.0001. From above equation it is clear that Quantity of Cholesterol(X2) exerts an inverse effect on the zeta potential of liposomes. As cholesterol content increases, it disrupts the orientation and density of charged phospholipid head groups, leading to a reduction in the magnitude of zeta potential. This effect is attributed to cholesterol’s ability to condense the bilayer structure and reduce the exposure of ionic groups to the surrounding medium, thereby diminishing electrostatic repulsion between vesicles. Similar work was reported by Pavlović et al. (2025), who demonstrated that increasing cholesterol content in liposomal formulations led to a reduction in zeta potential [42]. The quantity of soya lecithin(X2) and the duration of sonication (X3) exhibited an insignificant effect on the zeta potential of liposomal formulations, indicating that variations in these parameters did not substantially alter the surface charge characteristics or electrostatic stability of the vesicles (fig. 3).
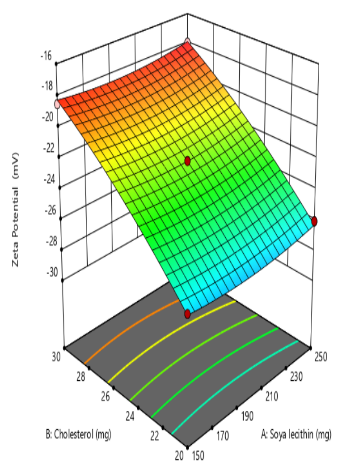
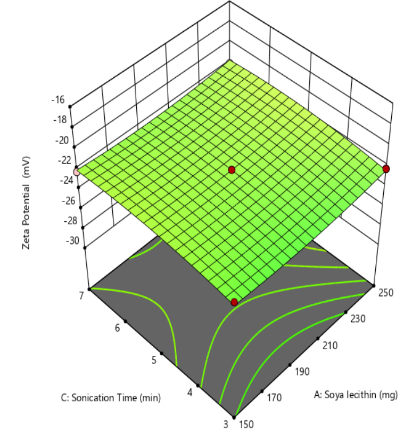
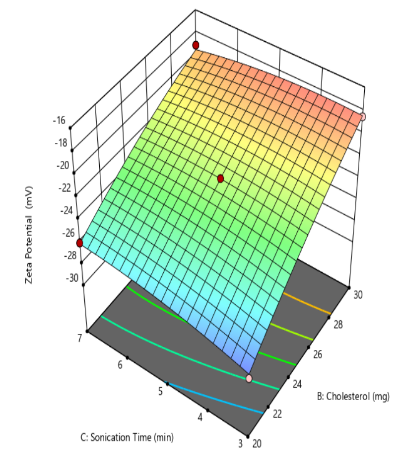
Fig. 3: Effect of independent variable on zeta potential
Table 4: Regression values for responses
| Parameter | Responses |
|||||
Particle size (Y1) |
Entrapment efficiency (Y2) |
Zeta potential (Y3) |
||||
F-value |
p-value |
F-value |
p-value |
F-value |
p-value |
|
| Model | 36.43 |
<0.0001 |
826.85 |
<0.0001 |
79.51 |
<0.0001 |
| Lack of fit | 3.38 |
0.1351 |
4.63 |
0.0864 |
1.48 |
0.3466 |
| R2 | 0.97910.9991 | |||||
| Adjusted R2 | 0.95220.99790.9779 | |||||
| Predicted R2 | 0.75090.98800.9112 | |||||
| Suggested model | Quadratic | Quadratic | Quadratic | |||
Y1=+239.46+11.93X1+2.23X2-0.8250X3+2.25X1X2+0.6000X1X3+6.20X2X3+2.02 X12–4.93 X22-1.48X32 Y2=+85.06+6.13X1+2.13X2+0.0001X3-0.2500X1X2+0.0000X1X3+0.0000X2X3-1.15X12-0.1550X22+0.0950X32 Y3=-22.28+0.1550X1+4.17X2+0.3375X3+0.0075X1X2-0.0225X1X3-0.8925X2X3+0.4903X12-0.4748X22-0.3948X32 |
||||||
Optimization and validation of the Opt-DTXL formulation
The optimized Opt-DTXL formulation was identified by applying constraints to the dependent variables, as outlined in table 4. Using point prediction analysis in Design Expert Software version 12, the formulation with the highest desirability approaching a value of 1was selected. The predicted optimal process parameters were: Soya lecithin (𝑋1) at 200 mg, Cholesterol (𝑋2) at 25 mg, and sonication time (𝑋3) at 5 min. corresponding predicted response values included particle size (𝑌1) of 239.46 nm, entrapment efficiency (𝑌2) of 85.06%, and zeta potential (𝑌3) of –22.27mV. The optimized formulation (Opt-DTXL) was subsequently prepared and characterized for particle size, entrapment efficiency, and zeta potential. The experimentally obtained values particle size (𝑌1) of 239.60 nm, entrapment efficiency (𝑌2) of 84.03%, and zeta potential (𝑌₃) of –22.85 mV were found to be in close agreement with the predicted values generated by the Response Surface Methodology (RSM), thereby confirming the reliability and validity of the RSM model [43].
Table 5: Predicted and observed responses for optimized formulation
| Responses | Predicted | Observed | % Error |
| Particle size(nm) | 239.46 | 239.60 | 0.05 |
| Entrapment efficiency (%) | 85.06 | 84.03 | 1.22 |
| Zeta potential(mV) | -22.27 | -22.85 | 2.53 |
Table 6: Particle size, Zeta potential, EE% of FA cojugated Opt-DTXL (FA-Opt-DTXL)
| Formulation | Particle size (nm) | EE% | Zeta potential (mV) | PDI |
| Opt-DTXL | 239.6 | 84.03 | -22.85 | 0.40 |
| FA-Opt-DTXL | 241.0 | 86.01 | -23.74 | 0.46 |
TEM study
TEM imaging revealed that the Docetaxel-loaded liposome (Opt-DTXL) formulations (fig. 4a) exhibited a predominantly spherical morphology with smooth, well-defined boundaries, indicative of uniform vesicle formation. The observed particle size was consistent with dynamic light scattering results, averaging 239.6 nm (fig. 4b), which falls within the optimal range for passive targeting and systemic circulation. Notably, no signs of vesicle aggregation or fusion were detected, suggesting excellent colloidal stability and effective steric repulsion between particles. The absence of aggregation further supports the integrity of the bilayer structure and validates the formulation conditions used, including lipid composition and sonication parameters [44].
Particle size, Zeta potential and PDI of Opt-DTXL and FA-Opt-DTXL
The particle size of Opt-DTXL was recorded at 239.6 nm, while FA-Opt-DTXL exhibited a slightly larger size of 241 nm. This marginal increase can be attributed to the surface conjugation of folic acid, which adds molecular bulk to the liposomal surface without significantly altering the core structure. Both sizes fall within the optimal range (<300 nm). Zeta Potential Opt-DTXL showed a zeta potential of –22.85 mV, whereas FA-Opt-DTXL exhibited a slightly more negative value of –23.74 mV. The increase in negative surface charge upon folate conjugation may result from the ionizable groups present in the folic acid moiety or altered lipid packing at the bilayer interface [45]. A zeta potential magnitude above ±20 mV generally indicates sufficient electrostatic repulsion to prevent vesicle aggregation, suggesting both formulations possess good colloidal stability. Polydispersity Index (PDI) the PDI values for Opt-DTXL and FA-Opt-DTXL were 0.40 and 0.46, respectively. While both values indicate moderate size distribution, the slight increase in PDI for FA-Opt-DTXL may reflect heterogeneity introduced during the conjugation process. Nonetheless, values below 0.5 are considered acceptable for liposomal systems intended for drug delivery, indicating a reasonably uniform population of vesicles.
FTIR studies
The FTIR spectra of pure Docetaxel (DXT), physical mixture, optimized Docetaxel formulation (Opt-DTXL), and folate-conjugated optimized Docetaxel (FA-Opt-DTXL) reveal distinct molecular interactions and confirm the presence of characteristic functional groups (fig. 5). Docetaxel exhibits prominent absorption bands around 3400 cm⁻¹ (–OH stretching), 1730 cm⁻¹ (ester C=O stretching), and 1600 cm⁻¹ (aromatic C=C stretching), consistent with its native chemical structure. The physical mixture retains these peaks with minimal shifts, indicating the absence of significant interactions between Docetaxel and excipients. In contrast, Opt-DTXL shows slight shifts and broadening in the –OH and C=O regions, suggesting hydrogen bonding and encapsulation within the carrier matrix. Notably, FA-Opt-DTXL displays additional peaks and intensified bands near 1650 cm⁻¹ and 1500 cm⁻¹, corresponding to amide and aromatic vibrations, which confirm successful folate conjugation. These spectral modifications collectively validate the structural integrity of Docetaxel and the functionalization achieved through formulation and targeting strategies.
In vitro drug release
The in vitro release of pure docetaxel (Pure-DXT) and Optimized Docetaxel Liposomes (Opt-DTXL) were evaluated over a 24 h period to assess formulation-dependent modulation of drug release. Pure-DXT exhibited a rapid release profile, with 98.32% of the drug released within 8 h, indicative of an immediate-release behavior typical of unencapsulated drug. In contrast, Opt-DTXL demonstrated a sustained release pattern, reaching 79.32% at 8 h and achieving near-complete release (99.32%) by 24 h. This controlled release is attributed to liposomal encapsulation, which likely imparts membrane stability and diffusion resistance. Overall, both liposomal formulations significantly prolonged the release of docetaxel compared to the free drug, with Opt-DTXL (fig. 6).
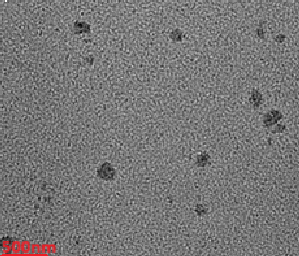

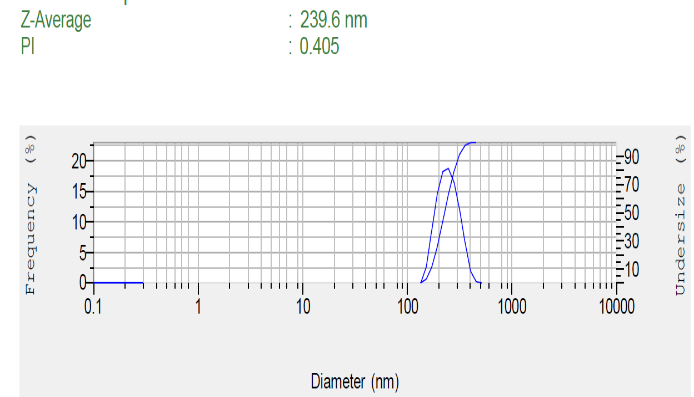
a b
c
Fig. 4: 4a-TEM, 4b-Particle size distribution 4c-zeta potential
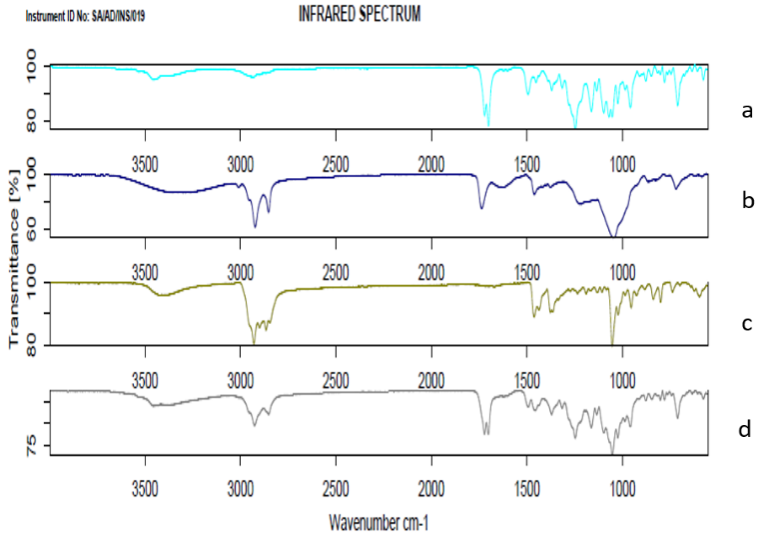
Fig. 5: FTIR study of a: Docetaxel b: Physical mixturec: Opt-DTXL d: FA-Opt-DTXL
MTT assay on A549 lung adenocarcinoma cells
The cytotoxic potential of Pure Docetaxel (Pure-DXT), Optimized Docetaxel Liposomes (Opt-DTXL), and FA-conjugated Optimized Docetaxel Liposomes (FA-Opt-DTXL) was evaluated against A549 lung adenocarcinoma cells across a concentration range of 0.1–100 µg/ml (fig. 7). All formulations exhibited a dose-dependent reduction in cell viability, with FA-Opt-DTXL demonstrating the most pronounced cytotoxic effect at each concentration. At 25 µg/ml, FA-Opt-DTXL reduced cell viability to 36.68%, compared to 49.84% for Opt-DTXL and 51.99% for Pure-DXT. At the highest tested concentration (100 µg/ml), FA-Opt-DTXL achieved near-complete cytotoxicity (5.47%), outperforming Opt-DTXL (10.32%) and Pure-DXT (12.32%). These trends were quantitatively confirmed by the half-maximal inhibitory concentration (IC₅₀) values: FA-Opt-DTXL exhibited the lowest IC₅₀ of 16.67 µg/ml, followed by Opt-DTXL at 27.96 µg/ml and Pure-DXT at 38.06 µg/ml. The IC₅₀ value, defined as the concentration of a drug required to inhibit 50% of cell viability in vitro, serves as a critical indicator of cytotoxic potency. Lower IC₅₀ values reflect higher efficacy, as less drug is needed to achieve the same biological effect. In this study, the superior potency of FA-Opt-DTXL is attributed to its folic acid–mediated targeting mechanism. Folic acid binds selectively to folate receptors, which are overexpressed on A549 cells, facilitating receptor-mediated endocytosis and enhancing intracellular accumulation of docetaxel. This targeted delivery not only improves drug uptake but also amplifies apoptotic induction, resulting in a more robust cytotoxic response. In contrast, Opt-DTXL benefits from liposomal encapsulation, which improves drug stability and cellular uptake compared to free docetaxel, but lacks the active targeting component. Pure-DXT, while effective at higher concentrations, demonstrates limited potency due to rapid clearance and non-specific distribution.
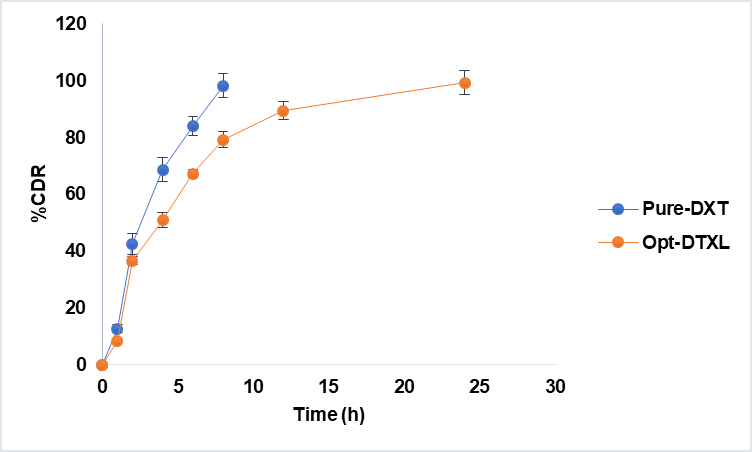
Fig. 6: In vitro drug release study, results are given in mean± SD, n=3
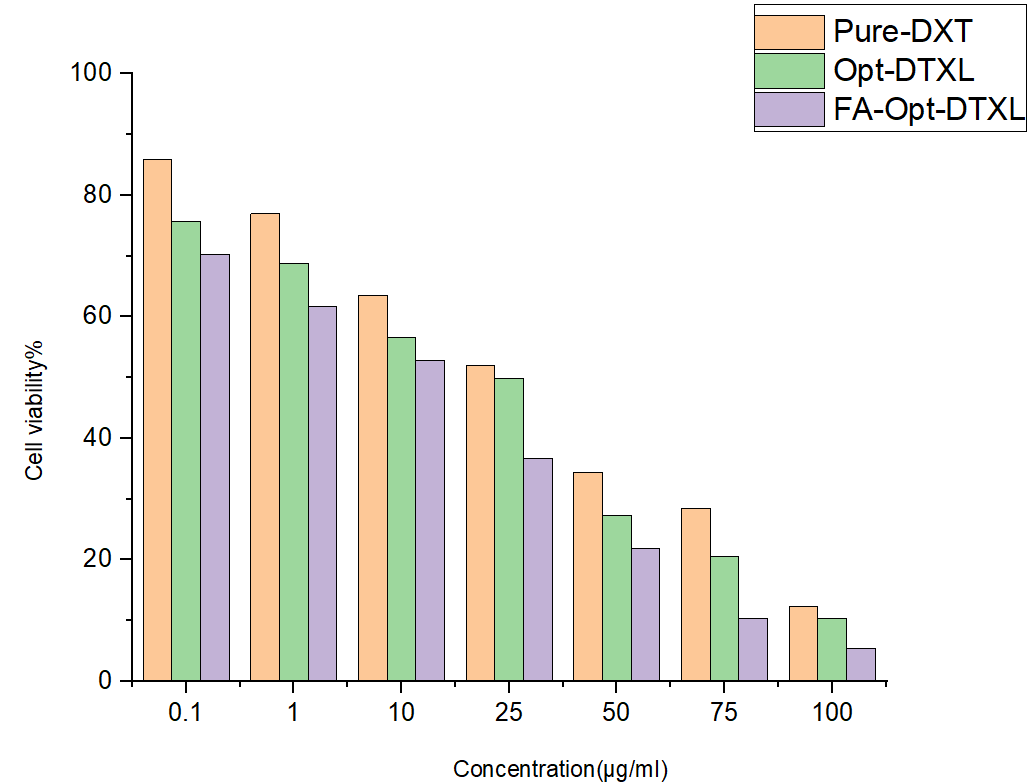
Fig. 7: In vitro drug release study, results are given in mean, n=3
Apoptosis activity against A549 lung adenocarcinoma cells
Dual staining with acridine orange (AO) and ethidium bromide (EB) enabled qualitative assessment of apoptosis in A549 lung adenocarcinoma cells following treatment with Pure Docetaxel, Opt-DTXL, and FA-Opt-DTXL at their respective IC₅₀ concentrations. AO, a cell-permeable dye, stained viable cells green by intercalating into double-stranded DNA, while EB penetrated only cells with compromised membranes, emitting red fluorescence upon DNA binding. This differential uptake allowed clear visualization of apoptotic progression. Cells treated with Pure Docetaxel predominantly exhibited green fluorescence, indicating a higher proportion of viable cells with intact membranes. Occasional yellow-green fluorescence and mild chromatin condensation suggested early apoptotic changes, but the overall apoptotic induction appeared limited. In contrast, Opt-DTXL-treated cells showed a marked increase in yellow-green fluorescence, with visible nuclear condensation and fragmentation. The presence of red-stained cells indicated progression to late apoptosis or secondary necrosis [46]. These findings suggest that liposomal encapsulation of Docetaxel enhances intracellular delivery and apoptotic efficacy compared to the free drug. The most pronounced apoptotic response was observed in the FA-Opt-DTXL group, where cells displayed dominant red fluorescence, extensive chromatin fragmentation, and minimal green signal. This indicates a high proportion of late apoptotic or necrotic cells. The folate-targeted liposomes likely facilitated receptor-mediated endocytosis, resulting in superior cellular uptake and enhanced cytotoxicity. Overall, the fluorescence profiles demonstrate a treatment-dependent escalation in apoptotic activity, with FA-Opt-DTXL showing the highest efficacy. These qualitative observations align with the expected pharmacodynamic behaviour of targeted liposomal formulations and support their potential for improved therapeutic outcomes in lung cancer models [fig. 8].
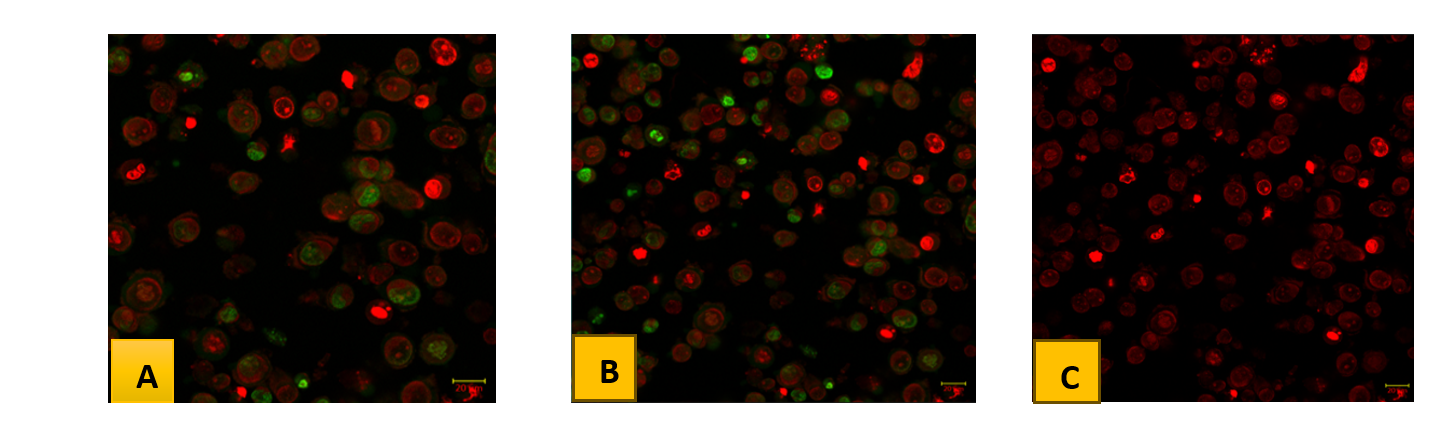
Fig. 8: Acridine orange (AO) and Ethidium bromide (EB) dual staining study of HepG2 cells in Untreated and treated with IC50 concentration of given compounds represented the changes in nuclear morphology of cells. (8A. A549 lung adenocarcinoma cells with pure docetaxel, 8B. A549 lung adenocarcinoma with Opt-DTXLand 8C. A549 lung adenocarcinoma with FA-Opt-DTXL) AO represents viable cells (green) and EtBr represents dead cells (red color). All the images were captured at 25x magnification. Scale bar-20um
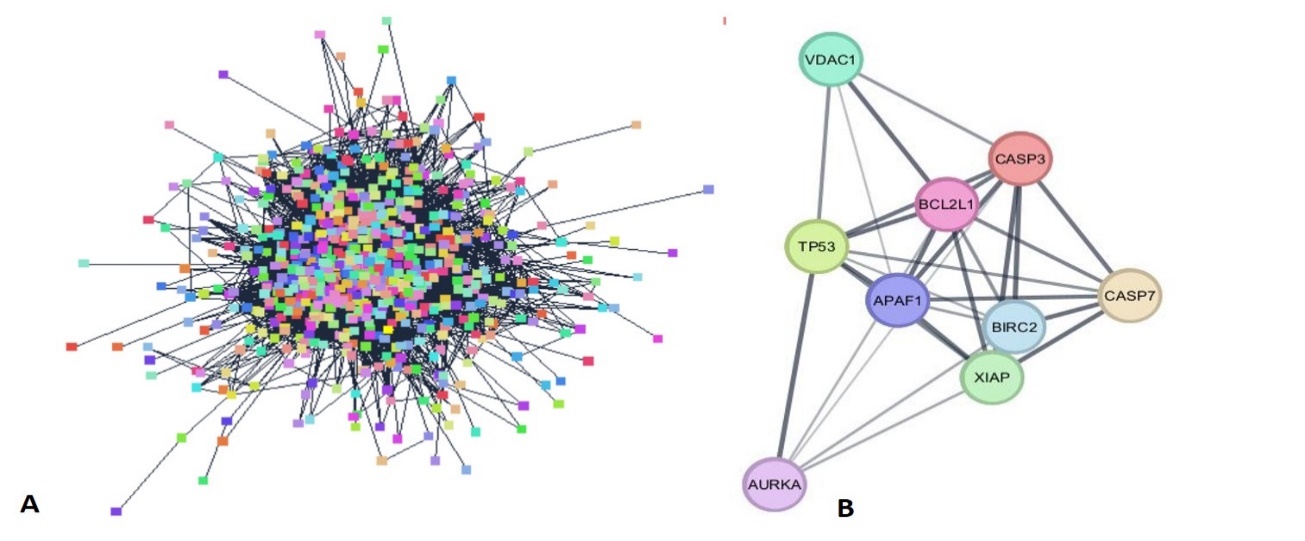
Fig. 9: PPI Network of: A-All 1118 common targets, B-9 best targets as per the closeness in the string network
Table 7: List of target genes showed most closeness as per the data generated by cytohubba
| String canonical name (protein) | Shared string name | Compartment nucleus | Compartment cytosol |
| VDAC1_HUMAN | ENSP00000378487 | 1.696423 | 2.754193 |
| AURKA_HUMAN | ENSP00000216911 | 2.293012 | 2.075812 |
| XIAP_HUMAN | ENSP00000360242 | 2.212035 | 2.177107 |
| P53_HUMAN | ENSP00000269305 | 1.228747 | 1.804924 |
| B2CL1_HUMAN | ENSP00000365230 | 1.679983 | 1.83152 |
| BIRC2_HUMAN | ENSP00000477613 | 2.597501 | 2.54345 |
| APAF_HUMAN | NSP00000448165 | 2.77011 | 2.698812 |
| CASP7_HUMAN | ENSP00000358327 | 1.40803 | 1.682148 |
| CASP3_HUMAN | ENSP00000311032 | 1.677667 | 1.413827 |
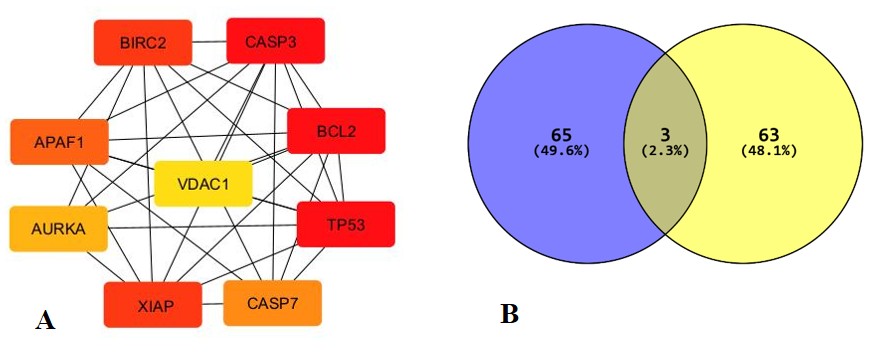
Fig. 10: Top shortest targets (compound network as per degree of closeness) estimated by cytohubba: BCL2L1, BIRC2, CASP3, APAF1, XIAP, CASP7, VDAC1, TP53 and BCL2L1
Network pharmacology and molecular docking
Intersection of docetaxel and active lung cancer targets
After exploring the swiss target prediction and SEA for compound targets. Related targets without duplicate values were obtained. Meanwhile, Targets were identified in DisGeNET and TTD data-base. A total of 200 overlapping targets were obtained by determining the intersection by using Venny diagram as shown in fig. 10B.
Protein-protein Interaction (PPI) of docetaxel and network construction
A total of 131 common targets were input to the String database out of 244 genes, and the PPI network was exported, where the PPI network was visualized and analysed. The network was composed of 120 nodes and 1118 edges, average node degree was found to be 22.5 and average local clustering coefficient is 0.580. PPI enrichment p-value is. As shown in fig. 9A. From common targets top 9 degree shorted targets, fig. 10A were obtained from cytohubba through cytoscape string network as shown in fig. 9B and table 7.
Molecular docking results
Molecular docking studies employing docetaxel and folic acid against the identified key molecular targets-A549, BCL2L1, BIRC2, CASP3, APAF1, XIAP, CASP7, VDAC1, and TP53-were conducted to validate their therapeutic relevance in lung cancer. The docking results revealed strong binding affinities and favorable interaction profiles, suggesting stable ligand–target complexes that are crucial for modulating apoptotic and survival signaling cascades. These findings are consistent with earlier reports demonstrating docetaxel’s high binding propensity toward apoptosis-related proteins, including BCL-2 family members and caspases, thereby contributing to its well-established pro-apoptotic and antimitotic effects in lung cancer therapy [47]. The integration of docking outcomes with protein–protein interaction (PPI) network analysis using Cytoscape further strengthened the biological relevance of the identified targets. Key nodes such as TP53, CASP3, XIAP, and BCL2L1 exhibited high degree centrality and closeness values, indicating their pivotal regulatory roles within apoptotic networks. Similar network-based investigations have highlighted TP53 and caspase family proteins as central hubs governing cancer cell fate decisions, reinforcing the reliability of the current network topology. [48] The prominent positioning of VDAC1 and APAF1 within the network aligns with previously published studies emphasizing their involvement in mitochondrial-mediated apoptosis and cytochrome-c release mechanisms in lung carcinoma cells. Folic acid demonstrated favorable docking interactions with several targets, supporting its role in receptor-mediated targeting strategies. Prior studies have reported that folate-based ligands enhance cellular uptake through folate receptor overexpression in lung cancer cells, thereby improving therapeutic selectivity and intracellular drug accumulation. The observed ligand–target stability in the present docking analysis corroborates these findings and underscores the potential advantage of folate-guided delivery systems in lung cancer management. Furthermore, the combined utilization of molecular docking platforms and GeneBank-derived protein structures provided robust validation of ligand–target compatibility, minimizing structural uncertainty and enhancing predictive accuracy [49]. Comparable studies integrating docking and network pharmacology approaches have demonstrated superior reliability in elucidating multi-target mechanisms of anticancer formulations, particularly for complex diseases such as lung cancer.
Overall, the convergence of docking scores, network centrality parameters, and literature-supported molecular functions substantiates the mechanistic plausibility of the developed formulation in modulating critical apoptotic and survival pathways. These results not only align with existing experimental and computational evidence but also provide a strong foundation for further in-depth investigations, including molecular dynamics simulations, in vitro validation, and in vivo studies. Collectively, this integrated strategy highlights substantial potential for advancing targeted therapeutic development for lung cancer through multi-node molecular modulation.
CONCLUSION
This study successfully optimized docetaxel-loaded liposomes using Box–Behnken design, identifying formulation parameters that enhanced vesicle stability, drug retention, and cellular uptake. The optimized formulation (Opt-DTXL) exhibited a particle size of 239.46 nm, high entrapment efficiency (84.03%), and a zeta potential of –22.85 mV, all closely aligned with predicted model values The folic acid–conjugated formulation (FA-Opt-DTXL) demonstrated the most potent therapeutic activity, with significantly lower IC₅₀ values and pronounced apoptosis induction in A549 cells, confirming the advantage of folate receptor–mediated targeting. Network pharmacology and molecular docking further supported the mechanistic basis of this enhanced efficacy. Collectively, these findings establish FA-Opt-DTXL as a promising nanocarrier for site-specific docetaxel delivery in folate receptor–overexpressing cancers. Future in vivo pharmacokinetic and efficacy studies will be essential to validate its translational relevance and potential clinical application.
ACKNOWLEDGEMENT
I extend my heartfelt gratitude to my supervisor, Dr. Gurudutta Pattnaik, Dean of Centurion University of Technology and Management, for his unwavering support and insightful guidance throughout the course of my research. I am also profoundly thankful to Research Terminal, Hyderabad, for their instrumental role in facilitating formulation characterization studies, which substantially advanced the progress of this project.
FUNDING
Nil
AUTHORS CONTRIBUTIONS
Ayushi Pradhan: Conceptualization; Methodology; Writing – Original Draft Preparation, Gurudutta Pattnaik: Supervision; Project Administration, Ch Niranjan Patra: Data Curation; Formal Analysis, Dibyalochan Mohanty: Review ing; Editing Manuscript, Gnyana Ranjan Parida: Resources; Visualization, Mani Sharma: Data Curation; Formal Analysis Ayushi Pradhan: Gurudutta Pattnaik: Ch Niranjan Patra:
CONFLICT OF INTERESTS
Declared none
REFERENCES
Zhang S. Adenocarcinoma. In: Zhang S, editor. Diagnostic imaging of lung cancers. Singapore: Springer Nature; 2024. p. 3-49. doi: 10.1007/978-981-99-6815-2_1.
Smolarz B, Lukasiewicz H, Samulak D, Piekarska E, Kolacinski R, Romanowicz H. Lung cancer-epidemiology pathogenesis, treatment and molecular aspect (review of literature). Int J Mol Sci. 2025 Feb 26;26(5):2049. doi: 10.3390/ijms26052049, PMID 40076671.
Shi Y, Taherifard E, Saeed A, Saeed A. MASLD-related HCC: a comprehensive review of the trends, pathophysiology tumor microenvironment surveillance and treatment options. Curr Issues Mol Biol. 2024 Jun 13;46(6):5965-83. doi: 10.3390/cimb46060356, PMID 38921027.
Chamani FK, Etebari A, Hajivalili M, Mosaffa N, Jalali SA. Hypoxia and programmed cell death-ligand 1 expression in the tumor microenvironment: a review of the effects of hypoxia-induced factor-1 on immunotherapy. Mol Biol Rep. 2024 Dec;51(1):88. doi: 10.1007/s11033-023-08947-8, PMID 38183512.
Pournami PK, Rasheed N, Raveendran A, Gopinath V. Antineoplastic drug leads from plants and microbes. In: Haridas M, Abdulhameed S, Francis D, Kumar SS, editors. Drugs from nature: targets, assay systems and leads. Singapore: Springer Nature; 2024. p. 287-331. doi: 10.1007/978-981-99-9183-9_12.
Loushambam B, Yanglem S, Krishnaswami V, Kumar M, Vijayaraghavalu S. Nanomedicine: pioneering advances in neural disease, stroke and spinal cord injury treatment. Neuroglia. 2025 Feb 21;6(1):9. doi: 10.3390/neuroglia6010009.
Ibrahim I, Aiman M, Chee CF. Active targeting docetaxel-loaded nanocapsules, primarily composed of polycaprolactone, chitosan-folate and TPGS for lung cancer treatment. Malays J Med Health Sci. 2024 Jan 3;20(Suppl 2):52.
Mehta D, Dua C, Chakraborty R, Yadav P, Dasgupta U, Bajaj A. Docetaxel-conjugated bile acid-derived nanomicelles can inhibit tumour progression with reduced toxicity. Nanoscale Adv. 2025;7(7):2003-10. doi: 10.1039/D4NA00715H, PMID 39967859.
Korake S, Salve R, Gajbhiye V, Pawar A. αvβ3 integrin-targeted pH-responsive dendritic nanocarriers for enhanced anti-tumor efficacy of docetaxel against breast cancer. J Drug Deliv Sci Technol. 2024 Sep 1;99:105946. doi: 10.1016/j.jddst.2024.105946.
Chaudhuri A, Naveen Kumar D, Kumar D, Kumar Agrawal A. Functionalized solid lipid nanoparticles combining docetaxel and erlotinib synergize the anticancer efficacy against triple-negative breast cancer. Eur J Pharm Biopharm. 2024 Aug 1;201:114386. doi: 10.1016/j.ejpb.2024.114386, PMID 38950717.
Pandey D, Bhardwaj S, Roy KK. Self-nanoemulsifying drug delivery systems for oral anticancer drugs: development, application, and challenges. In: Singh SK, Mehndiratta S, Paudel KR, De Rubis G, Dureja H, Dua K, editors. Application of Self-Nanoemulsifying Drug Delivery Systems in Inflammatory Diseases. Singapore: CRC Press; 2025 Jun 27. p. 167-84. doi: 10.1201/9781032697475-13.
Li X, Yuan K, Yin Y, Tian Y, Guo Z, Qin Z. Docetaxel-loaded electrospun nanofibrous mats for local chemotherapy targeting positive surgical margins in prostate cancer. Mol Pharm. 2025 Mar 12;22(4):2213-23. doi: 10.1021/acs.molpharmaceut.4c01440, PMID 40073383.
Rodriques P, Thacker K, Bhupendra GP. Exploring lipid-based drug delivery in cancer therapy via liposomal formulations. Asian J Pharm Clin Res. 2022 May 7;15(5):15-22. doi: 10.22159/ajpcr.2022.v15i5.43668.
Agrawal SS, Baliga V, Londhe VY. Liposomal formulations: a recent update. Pharmaceutics. 2024 Dec 30;17(1):36. doi: 10.3390/pharmaceutics17010036, PMID 39861685.
Narmani A, Rezvani M, Farhood B, Darkhor P, Mohammadnejad J, Amini B. Folic acid functionalized nanoparticles as pharmaceutical carriers in drug delivery systems. Drug Dev Res. 2019 Jun;80(4):404-24. doi: 10.1002/ddr.21545, PMID 31140629.
Rajoriya V, Gupta R. Folate functionalized nanostructured lipid carriers of paclitaxel for site-specific A549 lung adenocarcinoma targeting. BioNanoScience. 2025 Sep;15(3):1-3. doi: 10.1007/s12668-025-02133-9.
Patel P, Raval M, Airao V, Ali N, Shazly GA, Khan R. Formulation of folate receptor-targeted silibinin-loaded inhalable chitosan nanoparticles by the QbD approach for lung cancer targeted delivery. ACS Omega. 2024 Feb 24;9(9):10353-70. doi: 10.1021/acsomega.3c07954, PMID 38463259.
Wang J, He W, Cheng L, Zhang H, Wang Y, Liu C. A modified thin film method for large scale production of dimeric artesunate phospholipid liposomes and comparison with conventional approaches. Int J Pharm. 2022 May 10;619:121714. doi: 10.1016/j.ijpharm.2022.121714, PMID 35367585.
Vardhan H, Mittal P, Adena SK, Upadhyay M, Yadav SK, Mishra B. Process optimization and in vivo performance of docetaxel loaded PHBV-TPGS therapeutic vesicles: a synergistic approach. Int J Biol Macromol. 2018 Mar 1;108:729-43. doi: 10.1016/j.ijbiomac.2017.10.172, PMID 29111267.
Podder S, Mukherjee S. Response surface methodology (RSM) as a tool in pharmaceutical formulation development. Asian J Pharm Clin Res. 2024 Nov 7;17(11):18–25. doi: 10.22159/ajpcr.2024v17i11.52149.
Jastrzebska AM, Kurtycz P, Olszyna A, Karwowska E, Miaskiewicz Peska E, Zaleska Radziwill M. The impact of zeta potential and physicochemical properties of TiO2–based nanocomposites on their biological activity. Int J Applied Ceramic Tech. 2015 Nov;12(6):1157-73. doi: 10.1111/ijac.12340.
Odeh F, Naffa R, Azzam H, Mahmoud IS, Alshaer W, Al Bawab A. Co-encapsulation of thymoquinone with docetaxel enhances the encapsulation efficiency into pegylated liposomes and the chemosensitivity of MCF7 breast cancer cells to docetaxel. Heliyon. 2019 Nov 1;5(11):e02919. doi: 10.1016/j.heliyon.2019.e02919, PMID 31844767.
Han J, Zhou C, French AD, Han G, Wu Q. Characterization of cellulose II nanoparticles regenerated from 1-butyl-3-methylimidazolium chloride. Carbohydr Polym. 2013 May 15;94(2):773-81. doi: 10.1016/j.carbpol.2013.02.003, PMID 23544632.
Kurkute P, Jadhav A, Pandit SV. Anticancer potential and cytotoxic activity of NN-32, a snake venom peptide, against A549, lung cancer cell line. Int J Pept Res Ther. 2023 Jun 8;29(4):67. doi: 10.1007/s10989-023-10527-0.
Liu K, Liu PC, Liu R, Wu X. Dual AO/EB staining to detect apoptosis in osteosarcoma cells compared with flow cytometry. Med Sci Monit Basic Res. 2015 Feb 9;21:15-20. doi: 10.12659/MSMBR.893327, PMID 25664686.
Vivek-Ananth RP, Mohanraj K, Sahoo AK, Samal A. IMPPAT 2.0: an enhanced and expanded phytochemical atlas of Indian medicinal plants. ACS Omega. 2023 Feb 23;8(9):8827-45. doi: 10.1021/acsomega.3c00156, PMID 36910986.
Borul SB, More SP. Study of drug likeness properties substituted indazoles. Int J Creat Res Thoughts. 2022;10(2):54-61.
Desai V, Shaikhsurab MZ, Varghese N, Ashtekar H. Molecular docking and network pharmacology study on active compounds of Cyprus rotundus for the treatment of diabetes mellitus. In Silico Pharmacol. 2024 Nov 2;12(2):98. doi: 10.1007/s40203-024-00273-6, PMID 39498162.
Singh A, Bocher O, Zeggini E. Insights into the molecular underpinning of type 2 diabetes complications. Hum Mol Genet. 2025 Mar 15;34(6):469-80. doi: 10.1093/hmg/ddae203, PMID 39807636.
Meng J, Qiu Y, Zhang Y, Zhao H, Shi S. CMI: CRISPR/Cas9-based efficient multiplexed integration in Saccharomyces cerevisiae. ACS Synth Biol. 2023 Feb 28;12(5):1408-14. doi: 10.1021/acssynbio.2c00591, PMID 36853024.
Jia A, Xu L, Wang Y. Venn diagrams in bioinformatics. Brief Bioinform. 2021 Sep;22(5):bbab108. doi: 10.1093/bib/bbab108, PMID 33839742.
Lee M. Recent advances in deep learning for protein-protein interaction analysis: a comprehensive review. Molecules. 2023 Jul 2;28(13):5169. doi: 10.3390/molecules28135169, PMID 37446831.
Jethalia V, Hasyagar SV, Bhamidipati K, Chatterjee J. Analysing the role of saraswatarishta in the treatment of neurological disorders based on network pharmacology. Neurosci Res Notes. 2021 Sep 18;3(5):23-35. doi: 10.31117/neuroscirn.v3i5.106.
Ayodele PF, Bamigbade A, Bamigbade OO, Adeniyi IA, Tachin ES, Seweje AJ. Illustrated procedure to perform molecular docking using PyRx and Biovia Discovery Studio Visualizer: a case study of 10kt with atropine. Prog Drug Discov Biomed Sci. 2023 Dec 20;6(1):e00045. doi: 10.36877/pddbs.a0000424.
Ashtekar SS, Bhatia NM, Bhatia MS. Exploration of leads from natural domain targeting HER2 in breast cancer: an in-silico approach. Int J Pept Res Ther. 2019 Jun 1;25(2):659-67. doi: 10.1007/s10989-018-9712-y.
Dwiastuti R, Noegrohati S, Istyastono EP, Marchaban. Formulation and physical properties observations of soy lecithin liposome containing 4-n-butylresorcinol. AIP Conf Proc. 2016 Jul 21;1755(1):160005.
Briuglia ML, Rotella C, McFarlane A, Lamprou DA. Influence of cholesterol on liposome stability and on in vitro drug release. Drug Deliv Transl Res. 2015 Jun;5(3):231-42. doi: 10.1007/s13346-015-0220-8, PMID 25787731.
Yun JS, Hwangbo SA, Jeong YG. Preparation of uniform nano liposomes using focused ultrasonic technology. Nanomaterials (Basel). 2023 Sep 22;13(19):2618. doi: 10.3390/nano13192618, PMID 37836259.
Ahmed SA, Salama AA, Gaber MH, Ali SA. Development and characterization of soy lecithin liposome as potential drug carrier systems for doxorubicin. J Pharm Innov. 2023 Sep;18(3):1415-26. doi: 10.1007/s12247-023-09732-7.
Fukuoka S, Shimomura A, Katsumura Y, Oki M, Tsuji G. Transcription and translation efficiency is reduced in cholesterol-containing liposomes. J Biochem. 2025 Jun 13;178(3):193-200. doi: 10.1093/jb/mvaf032, PMID 40511712.
Holsæter AM, Wizgird K, Karlsen I, Hemmingsen JF, Brandl M, Skalko Basnet N. How docetaxel entrapment vesicle size zeta potential and stability change with liposome composition-a formulation screening study. Eur J Pharm Sci. 2022 Oct 1;177:106267. doi: 10.1016/j.ejps.2022.106267, PMID 35872073.
Pavlovic N, Mijalkovic J, Balanc B, Lukovic N, Knezevic Jugovic Z. Role of cholesterol in modifying the physical and stability properties of liposomes and in vitro release of vitamin B12. Eng Proc. 2025 Jun 12;99(1):10. doi: 10.3390/engproc2025099010.
Bakhaidar RB, Naveen NR, Basim P, Murshid SS, Kurakula M, Alamoudi AJ. Response surface methodology (RSM) powered formulation development, optimization and evaluation of thiolated-based mucoadhesive nanocrystals for local delivery of simvastatin. Polymers (Basel). 2022 Nov 28;14(23):5184. doi: 10.3390/polym14235184, PMID 36501579.
Keck CM, Specht D, Brußler J. Influence of lipid matrix composition on biopharmaceutical properties of lipid nanoparticles. J Control Release. 2021 Oct 10;338:149-63. doi: 10.1016/j.jconrel.2021.08.016, PMID 34389366.
Kumar P, Huo P, Liu B. Formulation strategies for folate-targeted liposomes and their biomedical applications. Pharmaceutics. 2019 Aug 2;11(8):381. doi: 10.3390/pharmaceutics11080381, PMID 31382369.
Panzarini E, Tenuzzo B, Palazzo F, Chionna A, Dini L. Apoptosis induction and mitochondria alteration in human HeLa tumour cells by photoproducts of rose bengal acetate. J Photochem Photobiol B. 2006 Apr 3;83(1):39-47. doi: 10.1016/j.jphotobiol.2005.11.014, PMID 16427301.
Gu J. Anti-cancer effects of statins on castration- and chemotherapy-resistant prostate cancer cells. Hamburg, Germany: Universität Hamburg; 2024.
Ilyas S, Lee D. Exploring the role of BCL2 interactome in cancer: a protein/residue interaction network analysis. Biology (Basel). 2025 Mar 5;14(3):261. doi: 10.3390/biology14030261, PMID 40136517.
Singh V, Sinha S, Verma J. Bioinformatics models in drug delivery: predicting biomaterial-biological interactions for targeted therapies. Next Nanotechnol. 2026 Jun 1;9:100335. doi: 10.1016/j.nxnano.2025.100335.