
Int J App Pharm, Vol 18, Issue 2, 2026, 150-159Original Article
IN SILICO INVESTIGATION OF CONVOLVULUS PROSTRATUS AS A POTENTIAL SOURCE OF Α-GLUCOSIDASE INHIBITORS FOR DIABETIC MANAGEMENT
MAHENDRA GOWDRU SRINIVASA1, AMITHA SHETTY2*, SHREYA H. KANCHAN1, PRANAMYA1, KRISHNA YALLAPPA KOLACHI3, PRABITHA PRABHAKARAN3
1Nitte (Deemed to be University), NGSM Institute of Pharmaceutical Sciences (NGSMIPS), Department of Pharmaceutical Chemistry, Mangalore-575018, India. 2Nitte (Deemed to be University), NGSM Institute of Pharmaceutical Sciences (NGSMIPS), Department of Pharmaceutics, Mangalore-575018, India. 3Department of Pharmaceutical Chemistry, JSS College of Pharmacy, JSS Academy of Higher Education and Research, Mysuru-570015, Karnataka, India
*Corresponding author: Amitha Shetty; *Email: amithashetty@nitte.edu.in
Received: 28 Oct 2025, Revised and Accepted: 26 Dec 2025
ABSTRACT
Objective: Diabetes mellitus (DM) remains a major global health concern, and inhibition of intestinal α-glucosidase is a well-established therapeutic approach for managing postprandial hyperglycemia. Convolvulus prostrates (C. prostratus) contains several phytochemicals with reported antidiabetic relevance. This study aimed to computationally assess selected constituents from C. prostratus for their potential α-glucosidase interactions using an integrated in silico strategy.
Methods: Molecular docking, molecular mechanics–generalized born surface area binding free energy calculations, 100 ns molecular dynamics (MD) simulations, and absorption, distribution, metabolism, excretion, and toxicity predictions were performed to evaluate the interaction profiles and drug-likeness of prioritized phytoconstituents against human maltase–glucoamylase (PDB ID: 3TOP).
Results: Among the screened compounds, A18 (identified as quercetin, a well-known flavonoid previously reported in C. prostratus), showed comparatively favorable docking affinity (ΔG = −9.549 kcal/mol) and MM-GBSA binding energy (ΔG_bind = −31.44 kcal/mol) relative to the other constituents evaluated. MD simulations indicated that the quercetin–enzyme complex maintained stable binding, with consistent interactions involving key catalytic residues such as Asp1157, Asp1279, and Asp1526. ADMET profiling suggested good oral absorption and acceptable physicochemical characteristics, while noting that its predicted blood–brain barrier permeability may represent a potential liability for an intestinal enzyme inhibitor.
Conclusion: This study provides computational support for quercetin’s contributory role as one of the phytochemicals in C. prostratus capable of interacting with α-glucosidase. While quercetin demonstrated favorable in silico interaction and pharmacokinetic features compared with the other evaluated constituents, these findings primarily reinforce its known bioactivity and highlight the need for further in vitro and in vivo validation to substantiate its therapeutic relevance.
Keywords: α-glucosidase inhibitors, Convolvulus prostratus, Molecular docking, Molecular dynamics simulation, Diabetes mellitus
© 2026 The Authors.Published by Innovare Academic Sciences Pvt Ltd. This is an open access article under the CC BY license (https://creativecommons.org/licenses/by/4.0/)
DOI: https://dx.doi.org/10.22159/ijap.2026v18i2.57305 Journal homepage: https://innovareacademics.in/journals/index.php/ijap
INTRODUCTION
DM is marked by insufficient insulin production or impaired insulin action, causing chronic hyperglycemia and posing a major global health burden [1, 2]. About 10.5% of adults (536.6 million) were living with diabetes in 2021 [3, 4], projected to rise to 12.2% (783.2 million) by 2045 [5]. By 2030, diabetes is expected to become the sixth leading cause of death worldwide, highlighting the urgent need for effective therapies [6].
Type 2 diabetes mellitus (T2DM) is the most common form, characterized by insulin resistance and progressive β-cell dysfunction, leading to chronic hyperglycemia and cardiovascular complications [7]. Current treatments aim to control postprandial glucose, with α-glucosidase inhibition being a key therapeutic strategy [8]. α-Glucosidase, a brush border enzyme in the small intestine [9], hydrolyzes 1–4 glycosidic bonds to release absorbable glucose [10]. Its enhanced activity contributes to postprandial hyperglycemia in diabetics [11]. Thus, α-glucosidase inhibitors reduce carbohydrate digestion and glucose absorption, lowering postprandial glucose levels [12, 13].
Approved inhibitors such as acarbose, voglibose, and miglitol show clinical benefits [14], but their gastrointestinal side effects (diarrhea, flatulence, abdominal pain) limit adherence [15]. Hence, safer and more effective alternatives are needed.
Natural products offer diverse structures, established safety, and potentially fewer side effects, making them promising for antidiabetic drug discovery [16]. Medicinal plants have shown notable α-glucosidase inhibitory potential [17]. Convolvulus prostratus Forssk. (Shankhpushpi), widely used in Ayurveda [18, 19], is traditionally employed for diabetes, neurological disorders, anxiety, and cardiovascular issues. Its phytochemical richness alkaloids (convolamine, convolvine, convolvidine), flavonoids (kaempferol), phenolics (scopoletin, β-sitosterol), terpenoids, and glycosides contributes to its therapeutic effects [20].
Recent phytochemical studies reveal that extracts of C. prostratus possess moderate to strong α-glucosidase inhibitory activity, particularly the ethyl acetate fraction, with IC₅₀ values comparable to standard antidiabetic drugs. The activity is largely attributed to its flavonoid and phenolic content.
Computational approaches molecular docking, virtual screening, pharmacokinetic and ADMET prediction, accelerate natural product-based drug discovery by reducing cost, time, and animal use, while enabling efficient screening and optimization of bioactive compounds [21].
This study investigates the α-glucosidase inhibitory potential of C. prostratus phytoconstituents using in silico techniques, integrating traditional medicinal knowledge with modern computational tools to identify safer natural alternatives to existing synthetic inhibitors for T2DM management.
MATERIALS AND METHODS
Various components of the plant, such as its seeds, flesh, and even its flowers, have been harnessed for their prospective therapeutic attributes within the realm of traditional medicine/healing customs across the globe [22].
Convolvulus prostratus compounds
The candidate compounds from C. prostratus were obtained from the imppat database, pubchem, and literature [23].
Bioinformatics tools
Imppat database (https://cb.imsc.res.in/imppat) [24], chemdraw version-20.1.1 (https://informatics-support.perkinelmer.com), PubChem (www.pubchem.com), rcsb pdb (http://www.rscb.org/pdb), schrodinger software (https://www.schrodinger.com), swissadme (http://www.swissadme.ch), protox ii (https://tox-new.charite.de/protox_II [25].
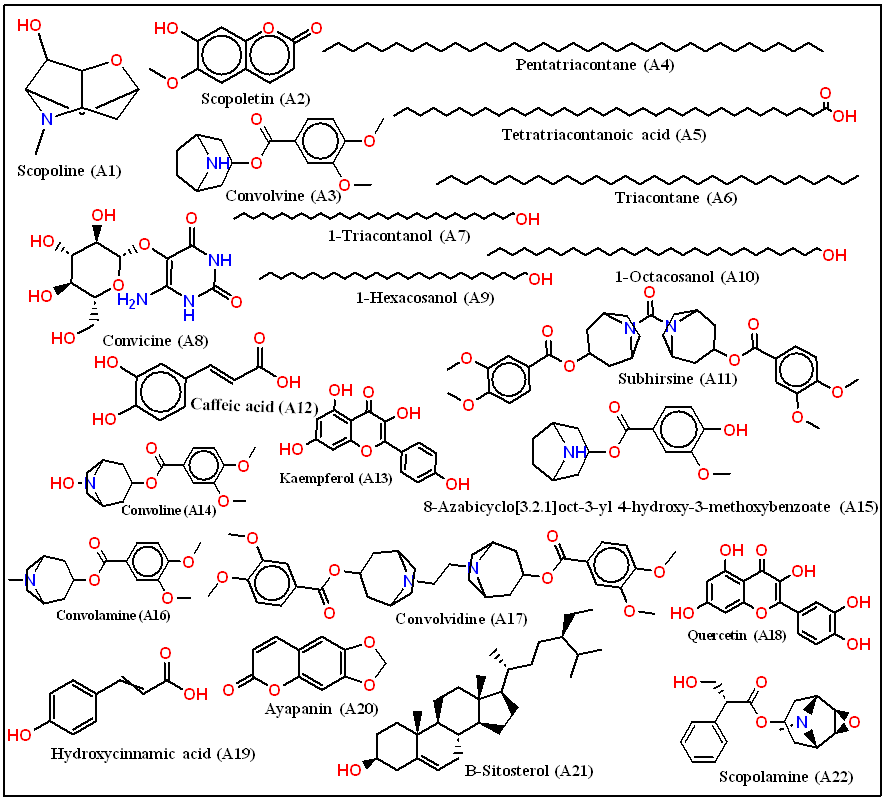
Fig. 1: Chemical structures of lead phytoconstituents identified from Convolvulus prostrates
Table 1: Lead three bioactive phytoconstituents
| Code | A15 | A17 | A18 |
| Common/chemical name | Vanilloyltropine | Convolvidine | Quercetin |
| IUPAC name | 8-Azabicyclo[3.2.1]oct-3-yl 4-hydroxy-3-methoxybenzoate | (1S,3R,5S)-3-(3,4-dimethoxybenzyl)-8-methyl-8-azabicyclo[3.2.1]octan-3-ol | 2-(3,4 dihydroxyphenyl)-3,5,7-trihydroxy-4H-chromen-4-one |
| Canonical SMILES | CN1[C@@H]2CC[C@H]1CC(C2)OC(=O)C3=CC(=C(C=C3)O)OC | CN1[C@H]2CC[C@H]1CC(C2)OC(=O)[C@H](CO)C3=CC=CC=C3 | C1=CC(=C(C=C1C2=C(C(=O)C3=C(C=C(C=C3O2)O)O)O)O)O |
| PubChem CID | 933171 | 6931560 | 5280343 |
Docking studies
Docking investigations were conducted using the Glide module within the Schrödinger 2020-3 suite, specifically the Maestro platform (https://www.schrodinger.com). These analyses were carried out on a Linux workstation [26].
Ligand preparation
The canonical SMILES representations of the 22 constituents from C. prostratus were sourced from pubchem and imppat (https://cb.imsc.res.in/imppat). All phytoconstituents evaluated in this study are presented in fig. 1, while comprehensive details of the lead molecules are provided in table 1. The top 10 compounds with the most favourable docking scores in their 2D structures, depicted in fig. 2, were singled out for subsequent investigation. These selected compounds underwent further processing using the ligprep module of the schrödinger suite (https://www.schrodinger.com). To maintain consistent biological relevance, high-energy ionization and tautomers were omitted during the epik tool preparation phase. The highest-scoring ligands are presented in fig. 3. The prioritized ligands were assessed using Lipinski's rule of five via the qikprop module to predict their potential properties [27].
Protein preparation
The preparation wizard tool was used to prepare the protein. The RCSB protein data bank (https://www.rcsb.org) provided the glucoamylase enzyme's three-dimensional structure. In particular, the 2.88 Å resolution Crystal Structure of Human Maltase-Glucoamylase in complex with acarbose (PDB ID: 3TOP) was used. Missing hydrogen atoms were added, appropriate bonding configurations were assigned, possible metal interactions were addressed, and water molecules within 5 Å of heterogeneous groups were removed as part of the preparation process. All polar group hydrogen atoms were visible after hydrogen bonds were optimized through sample orientation [28, 29].
Protein-ligand docking
The receptor grid indicates the precise region of the target protein where ligand interactions are investigated during the molecular docking process. The receptor grid generation interface in the Maestro Glide tool was used to create this grid [30]. The OPLS3e force field was used in the grid configuration. ligprep was used to define the receptor folders, and the Glide module was used to select them for docking with the flexibility set to extra precision (XP) mode. Based on calculated scores that included the grid score, proprietary glide score, and internal energy strain, the resulting binding interactions were ranked. The pose-viewer was used to visualize these results in a variety of structural output formats. Prioritizing ligands and predicting binding affinity were done using the glide score. Specifically, the molecular docking study was carried out using the α-glucosidase inhibitors docking program's xp mode [31].
Free energy calculation (-ΔG) by using prime molecular mechanics-generalized born surface area approach
The application of schrödinger 2020-4 prime molecular mechanics-generalized born surface area (MM-GBSA) was employed to compute the binding free energy of a protein-ligand complex. Additionally, post-docking energy minimization studies were performed to deepen the analysis of the complex. The minimized docking pose of the protein-ligand complex, achieved through the XP docking process, underwent assessment using the OPLS3e force field and the generalized-born surface area (GBSA) continuum VSGB 2.0 solvent model [32].
Molecular dynamics simulation
MD simulations were performed to evaluate the stability of α-glucosidase (PDB ID: 3TOP; resolution: 2.88 Å; R-free: 0.284) in complex with the phytoconstituents A15, A17, and A18 using desmond v12.6 (DE Shaw Research, New York, USA). The predominant receptor–ligand conformations obtained from molecular docking were selected as starting structures for the simulation study. Each complex was subjected to a 100 ns MD simulation employing the OPLS_2005 force field in a TIP3P water environment. A cubic solvated box was generated with a buffer distance of 10 Å in all directions from the protein surface, resulting in final system dimensions of approximately 92 × 92 × 92 ų. System neutrality and physiological ionic strength were ensured by adding 0.15 M NaCl along with appropriate Na⁺/Cl⁻ counterions. Prior to production runs, each system was energy-minimized using the steepest descent algorithm to eliminate steric clashes and stabilize initial coordinates. The minimized system was then gradually heated to 300 K under an NVT ensemble with heavy-atom restraints for 50 ps, followed by an NPT equilibration phase of 100 ps to stabilize temperature, pressure, and density. The native-state stability of α-glucosidase throughout the simulation was monitored using root mean square deviation (RMSD) analysis. To ensure reproducibility and robustness of the findings, MD simulations for each ligand-bound complex (A15, A17, and A18) were carried out independently for 100 ns, and trajectory data were recorded every 20 ps, yielding 1000 frames per simulation [33].
Absorption, distribution, metabolism, excretion, and toxicity prediction
Pharmacokinetic properties encompassing absorption, distribution, metabolism, excretion parameters, and bioavailability of Phytoconstituents were assessed utilizing the swissadme online web tool (http://www.swissadme.ch/h), [34] and admetsar (https://lmmd.ecust.edu.cn/admetsar2) [35]. All selected ligands' different SMILES were employed as input files. Drug absorption hinges on factors such as being a P-glycoprotein substrate (P-gp substrate), water solubility, skin permeability (log Kp) levels, membrane permeability, and gastrointestinal absorption (GSI). Notably, the blood-brain barrier (BBB) significantly influences drug distribution. Various CYP models facilitate assessing metabolism and volume of distribution, including types like CYP2C19 inhibitors, CYP1A2 inhibitors, and CYP3A4 inhibitors. Excretion primarily relies on total clearance, particularly of renal OCT2 substrate. For in silico toxicity prediction of proposed phytoconstituents, the freely accessible web server protox-iii was utilized (http://tox.charite.de/protox-III).
RESULTS AND DISCUSSION
The physicochemical properties and Lipinski’s rule of five parameters of the selected bioactive phytoconstituents (A15, A17, and A18) of C. prostratus were analyzed and are summarized in table 2. Lipinski’s rule is a widely accepted guideline used to predict oral bioavailability, suggesting that a compound is likely to exhibit favorable absorption and permeability if it does not violate more than one of the following criteria: molecular weight (≤500 Da), Log P (≤5), hydrogen bond donors (≤5), and hydrogen bond acceptors (≤10). As shown in table 1, compounds A15 and A18 fully complied with all the Lipinski parameters, indicating strong potential for good oral bioavailability. Compounds A15 (MW 277.319, Log P 1.6) and A18 (MW 302.24, Log P 0.352) had balanced lipophilicity and acceptable molecular weights, which promoted effective permeability and absorption. Both had the right proportion of donors and acceptors of hydrogen bonds, so there were no rule infractions.
On the other hand, as shown in table 1, A17 (MW 608.73, Log P 5.013) violated two Lipinski parameters: molecular weight and hydrogen bond acceptor count, indicating limited oral bioavailability because of its larger molecular size and polarity. These substances may still have pharmacological activity in spite of these infractions, though, and can be improved by structural alteration or different delivery methods. In accordance with its well-known poor gastrointestinal absorption but efficient local action in the intestinal lumen as an α-glucosidase inhibitor, the standard drug acarbose, used as a positive control, violated three parameters (molecular weight, donor, and acceptor counts), as indicated in table 2. This comparison demonstrates that while departures from Lipinski's criteria may result in decreased bioavailability, they do not always rule out pharmacological efficacy.
Table 2: Lipinski rule of 5 and drug likeness score of bioactive phytoconstituents
| Code | MW | Log p | Donor HB | Accept HB | Rule of five |
| Acceptable range | ≤500 | ≤5 | ≤5 | ≤10 | <5 |
| A15 | 277.319 | 1.6 | 2 | 5 | 0 |
| A17 | 608.73 | 5.013 | 0 | 11 | 2 |
| A18 | 302.24 | 0.352 | 4 | 5.25 | 0 |
| Acarbose (STD) | 645.611 | -7.002 | 14 | 19 | 3 |
STD = Standard; MW = Molecular Weight; Log P = Partition Coefficient (logarithm of octanol/water partition coefficient); Donor HB = Hydrogen Bond Donor; Accept HB = Hydrogen Bond Acceptor.
Molecular docking studies
The binding affinities and interaction profiles of C. prostratus bioactive phytoconstituents (A15, A17, and A18) against the target protein (PDB ID: 3TOP) were assessed using molecular docking studies. Table 3 summarizes the docking scores and detailed molecular interactions, and fig. 2 (A–D) shows the corresponding 2D and 3D interaction visualizations. Greater binding affinity and greater ability of interactions implies larger negative values of the binding energy (s 0 G), which is the stability of the ligand receptor complex.
All three compounds had binding energies of-8.473-9.549 kcal/mol and were therefore shown to have considerable binding affinities with the target enzyme as is reflected in table 3. The affinity of all of them was greatest to A18 (-9.549 kcal/mol), then A15 (-9.511 kcal/mol) and A17 (-8.473 kcal/mol) had relatively low affinity. The compound acarbose that is used as the reference has shown the highest negative binding energy (–11.432 kcal/mol), which is comparable to its commonly known capacity to inhibit the α-glucosidase enzymes. The validation redocking of acarbose into 3TOP resulted in an RMSD of 1.26 Å between the co-crystallized and redocked poses.
The interaction profile of A18 indicates that its high affinity can be attributed to the combined contribution of hydrogen bonding and π–π stacking interactions. Five hydrogen bonds were observed with Hie1584, Asp1279, Asp1526, Asp1157, and Trp1355, together with aromatic stacking interactions involving Trp1355 and Phe1427, and additional hydrophobic contacts with Ile1280, Trp1418, Met1421, and Phe1560. This mode of binding closely mirrors previously published docking studies on quercetin, which report that stabilization through hydrogen bonding with Asp1157/Asp1279 and π–π stacking with Trp1418 and Phe1427 is a major determinant of high inhibitory potential toward α-glucosidase [36]. Similarly, A15 displayed strong anchoring within the catalytic pocket, forming four hydrogen bonds with Asp1157, Arg1510, Asp1279, and Hie1584, along with a salt bridge with Asp1157 and multiple hydrophobic contacts with Pro1159, Phe1427, Trp1418, and Phe1559. This interaction network resembles the well-established binding pattern of potent α-glucosidase inhibitors in which hydrogen-bonding with catalytic residues and simultaneous aromatic interactions significantly enhance affinity.
Table 3: Molecular docking analysis: Convolvulus prostrates derivatives against PDB ID: 3TOP
Bioactive |
Binding energy (kcal/mol) |
H-bond residues |
No of H-bond |
π–π Interaction |
Salt bridge |
Other bond residues(Hydrophobic/Pi-cation/van der Waals) |
| A15 | -9.511 | Asp1157, Arg1510, Asp1279, Hie1584 | 4 | - | Asp1157 | Pro1159, Phe1427, Ser1425, Tyr1167, Phe1559, Phe1560, Met1421, Trp1418, Trp1523, Asp1420, Asp1526, Ile1280, Ile1315, Trp1355, Trp1369, Tyr1251 |
| A17 | -8.473 | Trp1369, Hie1584, Thr1586 |
3 |
- |
- |
Pro1159, Asp1157, Asp1420, Met1421, Trp1418, Phe1559, Phe1560, Lys1460, Ile1587, Phe1427, Ser1425, Val1363, Gly1365, Gln1372, Gln1286, Arg1377, Trp1355, Asp1279, Ile1280, Tyr1251, Ile1315 |
| A18 | -9.549 | Hie1584, Asp1279, Asp1526, Asp1157, Trp1355 | 5 | Trp1355, Phe1427 | - | Ile1280, Ile1315, Ile1587, Tyr1251, Thr1586, Trp1369, Trp1418, Trp1523, Met1421, Phe1559, Phe1560, Ser1425, Lys1460, Arg1510 |
| Acarbose (STD) | -11.432 | Arg1510, Asp1526, Arg1582, Asp1279, Gln1158, Asp1157 | 6 | - | - | Trp1523, Ile1280, Lys1480Trp1418, Thr1586, Hie1584 |
H-bond residues = amino acid residues involved in hydrogen bonding with the ligand; No. of H-bond = total number of hydrogen bonds formed between the ligand and protein; π–π interaction = aromatic ring stacking interaction between ligand and protein residues; STD=Standard.
A17, in contrast, formed only three hydrogen bonds without any aromatic stacking or salt-bridge interactions, relying mainly on weaker van der Waals and hydrophobic contacts, which likely accounts for its comparatively lower affinity. The binding mode of the standard inhibitor acarbose further supports this interpretation; it formed six hydrogen bonds with Arg1510, Asp1526, Arg1582, Asp1279, Gln1158, and Asp1157, consistent with the highly stabilized transition-state mimicry described in crystallographic studies of maltase-glucoamylase [37]. These observations indicate that the strong and persistent binding of A18 and A15 mimics the characteristic multi-interaction mode of acarbose, while the fewer stabilizing interactions of A17 correspond to its moderate binding score.
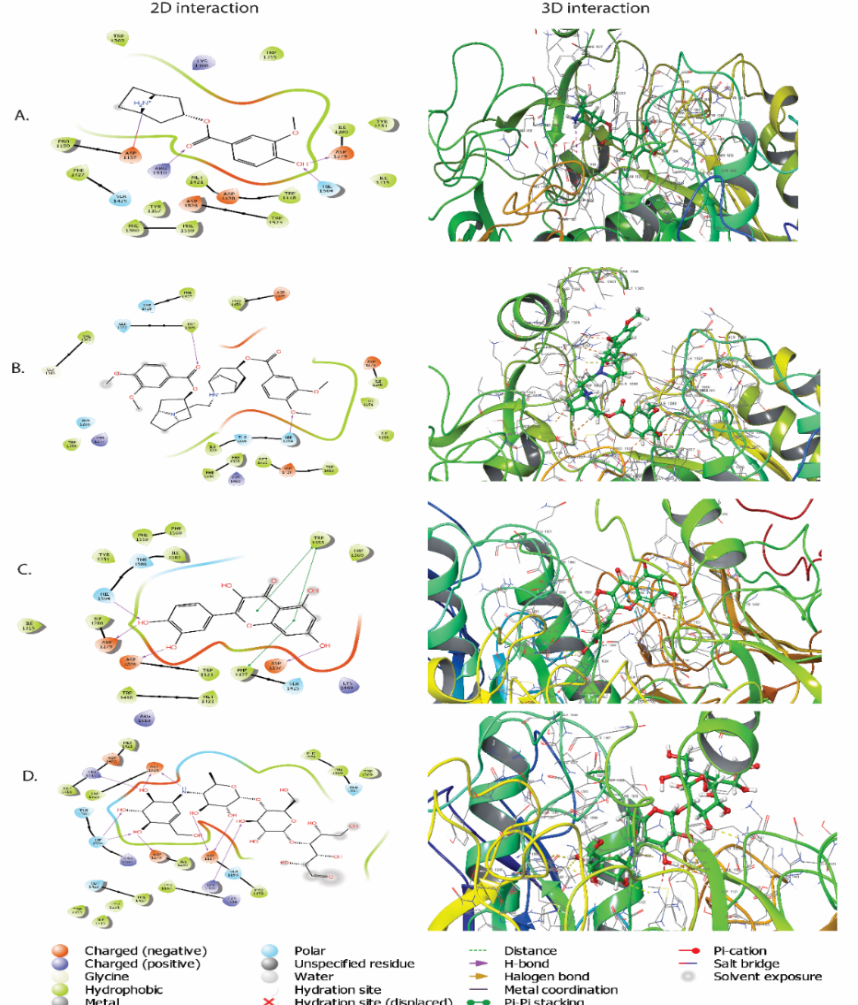
Fig. 2: (A) 2D and 3D interaction of compound A15 with 3TOP (B) 2D and 3D interaction of compound A17 with 3TOP (C) 2D and 3D interaction of compound A18 with 3TOP. (D) 2D and 3D interaction of standard acarbose with 3TOP
MM-GBSA approach
The binding free energies (ΔG bind) of the phytoconstituents A15, A17, and A18 with human α-glucosidase (PDB ID: 3TOP) were evaluated using the Prime/MM-GBSA method to complement the docking results and provide a refined assessment of ligand–protein stability. The calculated ΔG bind values ranged from –20.87 to –31.44 kcal/mol, indicating favorable binding for all three compounds, with A18 showing the strongest predicted affinity. A detailed analysis of the individual energy components further clarifies the driving forces underlying these interactions. Among the screened ligands, A18 (quercetin) demonstrated the most favorable ΔG bind (–31.44 kcal/mol), largely influenced by substantial van der Waals (–34.47 kcal/mol) and electrostatic (–39.54 kcal/mol) contributions, together with a moderately stabilizing hydrogen-bonding component (–3.43 kcal/mol). Its planar flavonoid structure allows efficient packing within the hydrophobic pocket of α-glucosidase, while multiple hydroxyl groups form persistent hydrogen bonds with key catalytic residues, including Asp1157, Asp1279, and Asp1526, thereby explaining its superior MM-GBSA performance.
In contrast, A17 displayed a moderate binding affinity (ΔG_bind = –20.91 kcal/mol), with stabilization primarily arising from van der Waals interactions (–48.92 kcal/mol) and lipophilic contacts (–27.11 kcal/mol). The comparatively weaker electrostatic contribution (–46.70 kcal/mol) and limited hydrogen bonding (–1.26 kcal/mol) suggest that the ligand interacts mainly through shape complementarity rather than strong directional interactions. This is consistent with its bulkier structure, which may hinder optimal positioning for hydrogen-bond formation despite maintaining hydrophobic interactions within the active site. A15 exhibited a similar overall binding energy (ΔG_bind = –20.87 kcal/mol), driven by balanced van der Waals (–38.84 kcal/mol) and lipophilic (–20.04 kcal/mol) contributions. Its electrostatic term (–48.65 kcal/mol) was comparable to that of A17, although fewer and less stable hydrogen-bonding interactions (–3.15 kcal/mol) were observed relative to A18. These characteristics indicate that while A15 occupies the binding pocket effectively, its interactions are not as strongly reinforced by persistent hydrogen bonds with catalytic residues.
The comparative MM-GBSA analysis reveals that van der Waals and lipophilic interactions are the dominant contributors to ligand stabilization within the predominantly hydrophobic α-glucosidase active site. A18 stands out because it uniquely combines favorable hydrophobic packing with stable hydrogen bonding to catalytically important acidic residues, resulting in the lowest and most favorable ΔG bind value among the tested compounds. The reference inhibitor acarbose displayed the strongest electrostatic interactions (–109.51 kcal/mol), consistent with its highly polar, carbohydrate-like structure. Collectively, these results validate the docking observations and strongly support A18 as the most stable and thermodynamically favorable ligand, while A15 and A17 also exhibit meaningful but comparatively weaker interaction profiles.
Table 4: Binding free energy calculation using MM-GBSA approach
| Bioactive | ∆G bind (kcal/mol) | ∆G bind coulomb | ∆G bind covalent | ∆G bind vander | ∆G bind H bond | ∆G bind lipophilic |
| A15 | -20.87 | -48.65 | 7.33 | -38.84 | -3.15 | -20.04 |
| A17 | -20.91 | -46.70 | 6.53 | -48.92 | -1.26 | -27.11 |
| A18 | -31.44 | -39.54 | 3.56 | -34.47 | -3.43 | -13.28 |
| Acarbose | -26.58 | -109.51 | 12.27 | -34.27 | -9.25 | -30.03 |
ΔG bind (kcal/mol) = Binding Free Energy; ΔG bind coulomb = Coulombic Contribution to Binding Free Energy; ΔG bind covalent = Covalent Interaction Contribution to Binding Free Energy; ΔG bind Vander = Van der Waals Contribution to Binding Free Energy; ΔG bind H-bond = Hydrogen Bonding Contribution to Binding Free Energy; ΔG bind lipophilic = Lipophilic Interaction Contribution to Binding Free Energy.
Molecular dynamics simulation
To evaluate the stability and dynamic behavior of α-glucosidase (PDB ID: 3TOP) in complex with bioactive compounds A15, A17, A18, and the conventional inhibitor acarbose, MD simulations lasting 100ns were conducted. Structural stability and residue flexibility were assessed using the RMSD and root mean square fluctuation (RMSF) analyses, respectively (fig. 3).
Root mean square fluctuation analysis
The conformational stability of the ligand and protein backbone during the simulation is shown by the RMSD plots (left panels). In the first 40ns before reaching stability, complexes A15 (fig. 3A) and A17 (fig. 3B) displayed moderate fluctuations in the protein RMSD (2.0–3.0 Å), suggesting that the ligands gradually adapted within the binding pocket. During the trajectory (100ns) the A18 complex (fig. 3C) was highly and steady bonded and with relatively consistent RMSD(~2.2 Å) with few variations. Acarbose complex (fig. 3D), in turn, displayed a slightly higher variation in RMSD (~2.5–3.2 Å), therefore, signifying that the protein experienced moderate alterations in its conformations on ligand binding. The same trends were observed in ligand RMSD trend where A18 had minimum number of fluctuations and this indicates that it is stable at the active site.
Root mean square fluctuation analysis
The flexibility of amino acid residues upon ligand binding is demonstrated by the RMSF plots (right panels). Higher fluctuations were seen in all complexes, mostly in the loop regions surrounding residues 350–450 and 700–750, which are typically linked to flexible or surface-exposed enzyme segments. While A15 and A17 displayed somewhat higher RMSF values close to the catalytic region, suggesting slight structural rearrangements, A18 caused the least amount of residue fluctuation among the tested ligands. According to its RMSD profile, the acarbose complex showed a moderate degree of flexibility. In comparison to A15, A17, and acarbose, the A18 complex shows better structural stability and fewer backbone fluctuations, according to the MD simulation results. This confirms the MM-GBSA results and implies that A18 interacts with α-glucosidase in a more stable and advantageous way, enhancing its potential as a promising inhibitor.
Protein–ligand contacts during 100 ns MD simulation
The stability and binding behavior of the synthesized compounds A15, A17, and A18 within the target protein's active site were assessed using 100 ns MD simulations, in conjunction with the standard drug acarbose. The residues involved and the types of interactions, including ionic interactions, water bridges, hydrophobic contacts, and hydrogen bonds, are highlighted in the protein–ligand contact plots and matching 2D interaction maps (fig. 4).
Fig. 4A, Compound A15, was demonstrated to have stable binding in the active site as demonstrated by hydrophobic interactions and continuous hydrogen bonds among key catalytic residues. The occupancy of the residues such as Tyr63, Asp197 and His201 in the trajectory was high, so they play an important role in maintaining the stability of the complex.
In the case of A17, the formations of strong hydrogen bonds with Asp300 and water-bridges with Tyr151 and Arg315 were observed (fig. 4B). Interactions remained fixed during the simulation, which implies moderately steady stability and a particular affinity of residues despite the number of contacts being relatively smaller than in A15.
A18 (fig. 4C) showed high binding affinity and positive orientation as it established many hydrophobic contacts and hydrogen bonds and in particular with Asp197, Glu233 and Tyr63. During the simulation, there were always hydrophilic and hydrophobic interactions, which implied enhanced complex stabilization of the ligand and protein.
Under its discrete binding model, the standard inhibitor acarbose (fig. 4D) exhibited a huge halogen bonding network with residues Asp197, Asp300 and Glu233. The percentage of hydrogen bond contacts and additional water bridges helped it to give it an overall strong binding stability. Among the phytoconstituents, A18 was shown to be a promising lead compound, and it had a similar interaction profile as acarbose. The stability and variety of non-covalent interactions suggest that such ligands maintain a favorable conformation in the active site, resulting in increased selectivity of binding the target protein.
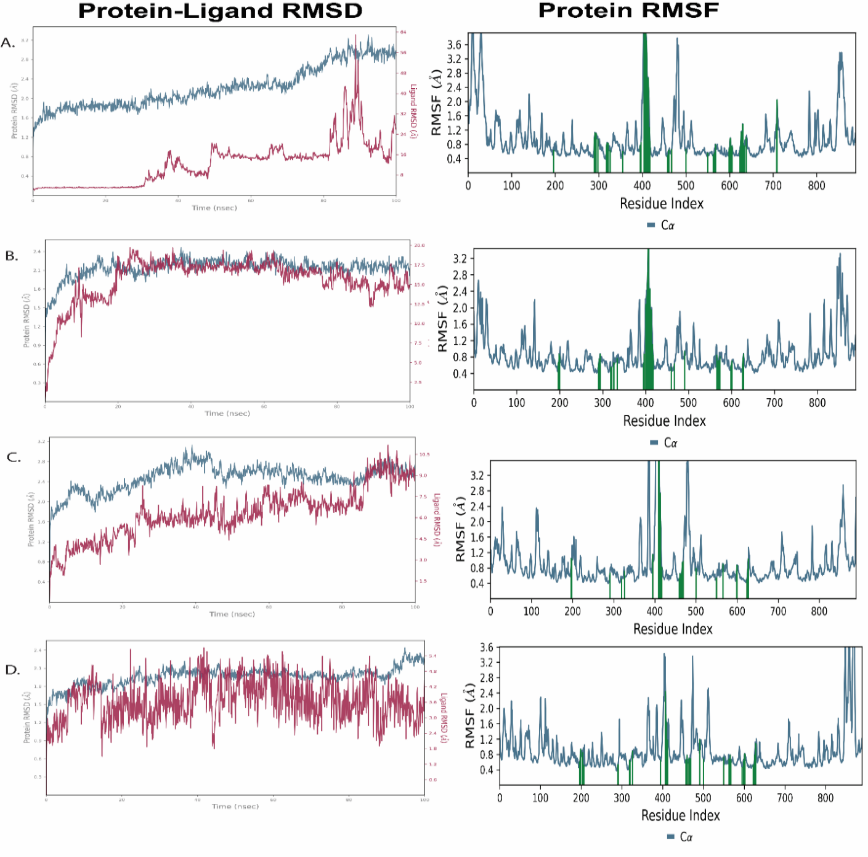
Fig. 3: RMSD assesses the overall structural stability of the protein–ligand complex, while RMSF measures residue-level flexibility to identify rigid and dynamic regions during the simulation of 3TOP docked with compounds A15, A17, A18 and acarbose retrieved in 100 ns time frame. A) RMSD and RMSF plots of 3TOP with A15, B) RMSD and RMSF plots of 3TOP with A17, C) RMSD and RMSF plots of 3TOP with A18, D) RMSD and RMSF plots of 3TOP with acarbose
Table 5: Predicted ADME parameters of the lead compounds
| Property | Model | Unit | Compound | ||
| A15 | A17 | A18 | |||
| Absorption | Water solubility | (Log mol/l) | 2.93 | 5.57 | 1.63 |
| Caco2 permeability | (Log Papp in 10-6 cm/s) | 0.5211 | 0.8957 | 0.8957 | |
| Intestinal absorption (human) | (% Absorbed) | 0.7978 | 0.9650 | 0.9650 | |
| Skin Permeability | (Log Kp) | -6.22 | -6.02 | -7.05 | |
| P-glycoprotein substrate | (Yes/No) | No | No | No | |
| P-glycoprotein I inhibitor | (Yes/No) | No | No | No | |
| P-glycoprotein II inhibitor | (Yes/No) | No | No | No | |
| Distribution | Fraction unbound (human) | (Fu) | 0.53 | 0.59 | 0.3 |
| BBB permeability | (Log BB) | -0.422 | 0.911 | 0.5711 | |
| CNS permeability | (Log PS) | 1 | 1 | -2 | |
| Metabolism | CYP2D6 substrate | (Yes/No) | No | No | No |
| CYP3A4 substrate | (Yes/No) | No | No | Yes | |
| CYP1A2 inhibitor | (Yes/No) | No | No | No | |
| CYP2C19 inhibitor | (Yes/No) | No | No | No | |
| CYP2C9 inhibitor | (Yes/No) | No | No | No | |
| CYP2D6 inhibitor | (Yes/No) | Yes | Yes | Yes | |
| CYP3A4 inhibitor | (Yes/No) | No | No | Yes | |
| Excretion | Renal OCT2 substrate | - | 0.6355 | 0.5979 | 0.9310 |
BBB permeability = ability of the compound to cross the blood–brain barrier; CNS permeability = predicted penetration into the central nervous system; CYP2D6 substrate/CYP3A4 substrate = compound metabolized by the corresponding cytochrome P450 isoenzyme; CYP1A2, CYP2C19, CYP2C9, CYP2D6, CYP3A4 inhibitor = compound predicted to inhibit the respective cytochrome P450 isoenzyme; Renal OCT2 substrate = compound predicted to be transported by organic cation transporter-2 in the kidneys.
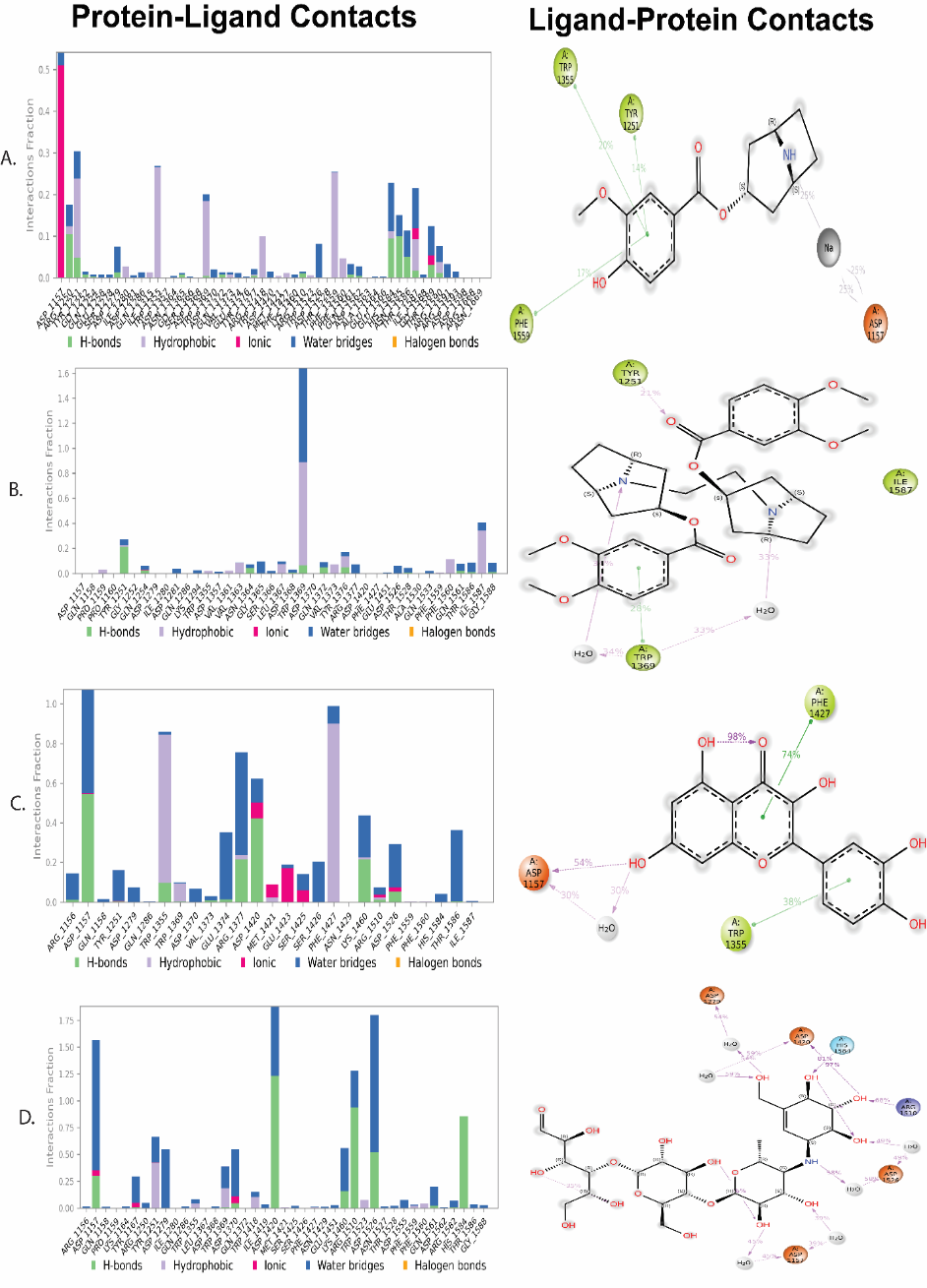
Fig. 4: The plots depict the interaction fractions of protein–ligand contacts between A15, A17, A18 and the standard acarbose with various amino acid residues during the 100 ns MD simulation
Absorption, distribution, metabolism, and excretion (ADME) analysis
In order to evaluate the ADME properties of the lead compounds A15, A17, and A18, their pharmacokinetic behavior was predicted using in silico ADME models (table 5). These findings provide insights into their potential bioavailability and overall drug-likeness.
A17 (Log mol/l = 5.57), the most soluble among the three, exhibited moderate to good water solubility. The predicted Caco-2 permeability values (0.5211–0.8957 × 10⁻⁶ cm/s) indicate favorable intestinal permeability. All compounds showed high human intestinal absorption (>90%), supporting good oral absorption potential. Limited dermal absorption was suggested by acceptable skin permeability values (Log Kp ranging from −6.22 to −7.05). Since none of the compounds were predicted to be P-glycoprotein substrates or inhibitors, the likelihood of efflux-associated absorption concerns is minimal.
Blood–brain barrier (BBB) permeability predictions revealed variable outcomes. A18 (Log BB = 0.5711) may be able to cross the BBB, whereas A15 (–0.422) and A17 (0.911) showed differing permeability levels. CNS permeability (Log PS) values further suggested that A17 and A18 may access the central nervous system (CNS).
Moderate plasma protein binding was indicated by the fraction unbound (Fu = 0.53–0.59), suggesting reasonable systemic circulation. All compounds demonstrated minimal interaction with major CYP enzymes, as none were predicted to be CYP2D6 or CYP3A4 substrates or inhibitors. Interestingly, all three compounds showed inhibitory potential toward CYP1A2 and CYP2C19, suggesting selective metabolic modulation that may influence metabolic stability or drug–drug interaction potential. Additionally, all compounds were predicted as renal OCT2 substrates (0.5979–0.9310), indicating a moderate likelihood of renal excretion.
Based on the overall ADME predictions, A18 appears to possess a balanced pharmacokinetic profile with good solubility, high intestinal absorption, moderate plasma binding, and limited CYP-mediated metabolism. However, its BBB permeability (Log BB = 0.57) must be considered a liability rather than an advantage. The revised discussion now clearly highlights that while quercetin (A18) shows favorable general pharmacokinetic traits, its CNS permeability may necessitate formulation or structural modification strategies to reduce BBB penetration if it is to be developed further as a therapeutic α-glucosidase inhibitor.
Toxicity profile
An in silico toxicity analysis was performed on the lead compounds A15, A17, and A18 produced by C. prostratus, to forecast the potential toxicological risks of the compound (including hepatotoxicity, carcinogenicity, immunotoxicity, mutagenicity, cytotoxicity, and receptor-mediated interactions) via use of the protox-iii server (table 6). All the three compounds, as per findings, had a good safety profile. Most of the parameters were forecasted to be dormant, indicating a minimal potential of toxicity. There was little chance of liver damage or genetic toxicity because none of the compounds had hepatotoxic, mutagenic, or cytotoxic effects. Perhaps as a result of structural characteristics affecting immune receptor interactions, compound A17 displayed a weak alert for carcinogenicity (probability 0.55), whereas compound A15 displayed mild immunotoxic potential (probability 0.52). A18, on the other hand, was inactive across these parameters, indicating a relatively higher level of safety.
A17 and A18 showed a weak activity on estrogen receptor alpha (ER-α), ligand-binding domain with moderate probabilities of activity (0.87 and 0.95, respectively), which in turn signals a low potential of endocrine disruption but most of the compounds were inactive on androgen, aromatase, and PPAR-γ receptors. A18's lack of cytotoxicity and mutagenicity supports good cellular safety, but it also demonstrated activity toward the mitochondrial membrane potential (probability 1.0), indicating a potential interaction at higher concentrations.
While A18 had an LD₅₀ of 159 mg/kg and was classified under class 3 (toxic if swallowed) with 100% prediction accuracy, A15 and A17 had predicted LD₅₀ values of 650 mg/kg, placing them in toxicity class 4 (harmful if swallowed). A18 has a slightly higher predicted toxicity, but given its potent pharmacological potential, its overall safety profile is still acceptable. A18 emerged as a strong yet safe bioactive lead candidate from C. prostratus, deserving of additional experimental validation, while the other two compounds showed low toxicity and good biocompatibility (table 6).
Table 6: Toxicity profile of leads bioactives from Convolvulus prostrates using Protox III
| S. No. | Target feature | Compound-A15 | Compound-A17 | Compound-A18 | |||
| Prediction# | Probability | Prediction | Probability | Prediction | Probability | ||
| Hepatotoxicity | Inactive | 0.80 | Inactive | 0.94 | Inactive | 0.69 | |
| Carcinogenicity | Inactive | 0.60 | Active | 0.55 | Inactive | 0.99 | |
| Immunotoxicity | Active | 0.52 | Inactive | 0.69 | Inactive | 0.87 | |
| Mutagenicity | Inactive | 0.75 | Inactive | 0.80 | Inactive | 0.51 | |
| Cytotoxicity | Inactive | 0.53 | Active | 0.50 | Inactive | 0.99 | |
| AhR | Inactive | 0.90 | Inactive | 0.78 | Active | 0.91 | |
| AR | Inactive | 0.98 | Inactive | 0.92 | Inactive | 0.99 | |
| AR-LBD | Inactive | 0.97 | Inactive | 0.99 | Inactive | 0.97 | |
| Aromatase | Inactive | 0.92 | Inactive | 0.85 | Inactive | 0.91 | |
| ER Alpha | Inactive | 0.90 | Inactive | 0.96 | Active | 0.87 | |
| ER-LBD | Inactive | 0.90 | Inactive | 0.96 | Active | 0.95 | |
| PPAR-Gamma | Inactive | 0.94 | Inactive | 0.96 | Inactive | 0.98 | |
| nrf2/ARE | Inactive | 0.96 | Inactive | 0.96 | Inactive | 0.99 | |
| HSE | Inactive | 0.96 | Inactive | 0.96 | Inactive | 0.99 | |
| MMP | Inactive | 0.75 | Inactive | 0.70 | Active | 1.0 | |
| Phosphoprotein (Tumor suppressor) p53 | Inactive | 0.90 | Inactive | 0.88 | Inactive | 0.97 | |
| ATAD5 | Inactive | 0.96 | Inactive | 0.98 | Inactive | 1.0 | |
| Predicted LD50 | 650 mg/kg | 650 mg/kg | 159 mg/kg | ||||
| Predicted Toxicity Class | 4 | 4 | 3 | ||||
| Prediction accuracy | 70.97% | 69.26% | 100% | ||||
#value in the brackets are probability; AhR = Aryl hydrocarbon Receptor; AR = Androgen Receptor; AR-LBD = Androgen Receptor Ligand Binding Domain; ER = Estrogen Receptor; ER-LBD = Estrogen Receptor Ligand Binding Domain; PPAR = Peroxisome Proliferator-Activated Receptor Gamma; nrf2/ARE = Nuclear factor (erythroid-derived 2)-like 2/antioxidant responsive element; HSE = Heat shock factor response element; MMP = Mitochondrial Membrane Potential; ATAD5 = ATPase family AAA (ATPases Associated with diverse cellular Activities) domain-containing protein 5.
The present study demonstrates that among the screened C. prostratus phytoconstituents, compounds A18 and A15 exhibit strong drug-likeness, favorable binding affinity, and robust interaction stability toward α-glucosidase, in agreement with trends reported for other naturally occurring flavonoids and glycosidase inhibitors. Previous studies on flavonoids such as quercetin, kaempferol, and rutin have established that compliance with Lipinski’s rule—particularly moderate lipophilicity and optimal hydrogen-bonding capability—correlates strongly with oral bioavailability and effective enzyme inhibition. Similar observations have been reported by Proenca et al. (2017) [38] and He et al. (2029) [39], who noted that molecules exhibiting balanced LogP values and limited Lipinski violations tend to show superior gastrointestinal absorption and target engagement. The present findings align with these reports, especially for A15 and A18, both of which showed complete adherence to the rule of five.
The docking analysis revealed that A18 demonstrated a high binding affinity (–9.549 kcal/mol) supported by multiple hydrogen bonds and π–π stacking interactions. This pattern closely matches earlier findings on quercetin and flavonoid analogs, where stabilization via Asp1157, Asp1279, Trp1355, and Phe1427 was shown to be central to strong α-glucosidase inhibition (Tian et al., 2024; Kwon et al., 2022) [40, 41]. The interaction mode of A15, characterized by engagement of Asp1157 and Arg1510, also reflects the binding profiles documented for potent α-glucosidase inhibitors isolated from medicinal plants such as Morus alba and Camellia sinensis, further emphasizing the structural relevance of acidic and aromatic residues in catalytic pocket stabilization.
The MM-GBSA results reinforced the docking outcomes, with A18 showing the highest overall stability (ΔG_bind = –31.44 kcal/mol). Prior computational studies have pointed out that flavonoids with planar aromatic rings and multiple hydroxyl groups exhibit stronger van der Waals and electrostatic contributions, enhancing binding free energy-a pattern observed in both the current analysis and earlier reports by Liu et al. (2020) [42] and Genheden et al. (2015) [43]. The moderate MM-GBSA profiles of A15 and A17 correlate with their reduced hydrogen-bonding capacity and larger steric bulk (in A17), similar to trends documented in bulky glycoside inhibitors.
MD simulation revealed that A18 maintained the lowest RMSD and minimal residue fluctuations, suggesting high conformational stability. Comparable MD findings have been reported for quercetin analogs bound to α-glucosidase and maltase-glucoamylase, where reduced backbone fluctuations and persistent hydrogen bonds were predictive of superior inhibitory activity [44]. Furthermore, the interaction persistence patterns, particularly involving Asp197, Glu233, and Tyr63 were consistent with earlier structural studies that identified these residues as key catalytic elements.
ADME predictions indicated high intestinal absorption and moderate plasma protein binding for all compounds, comparable to reported pharmacokinetic trends for plant-derived flavonoids. The predicted BBB permeability of A18 is in line with previous reports showing that quercetin readily crosses the BBB, which, while beneficial for neuroprotective applications, may pose a concern for peripherally targeted antidiabetic therapy [45]. The favorable CYP450 and OCT2 profiles support the potential of these compounds as orally bioavailable agents, similar to findings from studies evaluating herbal inhibitors of α-glucosidase.
Regarding toxicity, all three compounds exhibited generally safe profiles consistent with published toxicological data for structurally similar phytochemicals. The mild toxicity class associated with A18 reflects typical flavonoid behavior, with previous ProTox and experimental datasets confirming low carcinogenic and mutagenic risk for quercetin-like scaffolds when administered at therapeutic doses.
The present work advances the understanding of C. prostratus–derived phytoconstituents by demonstrating interaction patterns, stability trends, and pharmacokinetic behavior that closely parallel those reported for clinically relevant α-glucosidase inhibitors and natural flavonoids. This comparative evaluation underscores the potential of A18 and A15 as promising lead candidates and highlights the need for further in vitro and in vivo studies to validate their therapeutic efficacy.
CONCLUSION
In this work, phytoconstituents reported from C. prostratus were systematically evaluated for their α-glucosidase inhibitory potential using an integrated in silico approach comprising molecular docking, MM-GBSA binding energy calculations, 100 ns MD simulations, and ADMET/toxicity prediction. Among the analyzed molecules, A18identified as quercetin, a well-known flavonoid naturally present in C. prostrates exhibited the most favorable binding affinity, stable interaction profile with key catalytic residues, and generally acceptable pharmacokinetic characteristics. However, its predicted effect on mitochondrial membrane potential suggests a possible mitochondrial toxicity liability that warrants careful examination in experimental studies.
Rather than representing the identification of a novel inhibitor, this study provides a comprehensive computational characterization of quercetin and related phytoconstituents from C. prostratus in the context of α-glucosidase inhibition. The findings collectively support these compounds as promising candidates for further biological assessment. Future in vitro enzyme assays, cell-based evaluations, and in vivo studies are required to validate their therapeutic relevance and safety for potential use in the management of postprandial hyperglycemia.
ACKNOWLEDGEMENT
We gratefully acknowledge the NGSM Institute of Pharmaceutical Sciences, Nitte (Deemed to be University), Mangalore, Karnataka, and the NGSM-CADD Laboratory for providing essential facilities and resources that enabled the successful completion of this work.
FUNDING
The scientific work received no funding.
AUTHORS CONTRIBUTIONS
MGS: Conceptualization, data collection and analysis, and writing of the original draft. AS: Conceptualization, supervision, data collection and analysis, and manuscript review. SK and PR: Scientific accuracy verification, manuscript review and editing. KYK and PP: Manuscript review and editing.
CONFLICT OF INTERESTS
The authors confirm that there are no known conflicts of interest associated with this publication.
REFERENCES
Ojo OA, Ibrahim HS, Rotimi DE, Ogunlakin AD, Ojo AB. Diabetes mellitus: from molecular mechanism to pathophysiology and pharmacology. Med Novel Technol Devices. 2023 Sep;19:100247. doi: 10.1016/j.medntd.2023.100247.
Alam S, Hasan MdK, Neaz S, Hussain N, Hossain MdF, Rahman T. Diabetes mellitus: insights from epidemiology, biochemistry risk factors diagnosis, complications and comprehensive management. Diabetology. 2021 Apr 16;2(2):36-50. doi: 10.3390/diabetology2020004.
Belhadj M, Malek R, Baghous H, Boukheloua M, Arbouche Z, Nouri N. Perspectives of type 2 diabetes mellitus management in algeria: a comprehensive expert review. Front Clin Diabetes Healthc. 2025 Apr 15;6:1495849. doi: 10.3389/fcdhc.2025.1495849, PMID 40303934.
Mersal FA. Improving self-care practice for adults with type 2 diabetes mellitus. Innov J Med Health Sci. 2013 Oct 14;3(3):93-101.
Savelieff MG, Chen KS, Elzinga SE, Feldman EL. Diabetes and dementia: clinical perspective, innovation knowledge gaps. J Diabetes Complications. 2022 Nov;36(11):108333. doi: 10.1016/j.jdiacomp.2022.108333, PMID 36240668.
Gregg EW, Buckley J, Ali MK, Davies J, Flood D, Mehta R. Improving health outcomes of people with diabetes: target setting for the WHO global diabetes compact. Lancet. 2023 Apr 15;401(10384):1302-12. doi: 10.1016/S0140-6736(23)00001-6, PMID 36931289.
Galicia Garcia U, Benito Vicente A, Jebari S, Larrea Sebal A, Siddiqi H, Uribe KB. Pathophysiology of type 2 diabetes mellitus. Int J Mol Sci. 2020 Aug 30;21(17):6275. doi: 10.3390/ijms21176275, PMID 32872570.
Singdam P, Kamnate A, Somsap OA, Tohkayomatee R. Phytochemical screening, antioxidant potential and α-glucosidase inhibition of causonis trifolia leaf extracts: a solvent-based comparative study. Pharmacogn J. 2025 May 7;17(2):164-70. doi: 10.5530/pj.2025.17.20.
Tannous S, Stellbrinck T, Hoter A, Naim HY. Interaction between the α-glucosidases sucrase-isomaltase and maltase-glucoamylase in human intestinal brush border membranes and its potential impact on disaccharide digestion. Front Mol Biosci. 2023 Mar 8;10:1160860. doi: 10.3389/fmolb.2023.1160860, PMID 36968271.
Garcia CA, Gardner JG. Bacterial α-diglucoside metabolism: perspectives and potential for biotechnology and biomedicine. Appl Microbiol Biotechnol. 2021 May;105(10):4033-52. doi: 10.1007/s00253-021-11322-x, PMID 33961116.
Hiyoshi T, Fujiwara M, Yao Z. Postprandial hyperglycemia and postprandial hypertriglyceridemia in type 2 diabetes. J Biomed Res. 2017 Nov 1;33(1):1-16. doi: 10.7555/jbr.31.20160164, PMID 29089472.
Bhatnagar A, Mishra A. α-glucosidase inhibitors for diabetes/blood sugar regulation. In: Maheshwari VL, Patil RH, editors. Natural products as enzyme inhibitors. Singapore: Springer Nature; 2022. p. 269-83. doi: 10.1007/978-981-19-0932-0_12.
Aoki K, Sato H, Terauchi Y. Usefulness of antidiabetic alpha-glucosidase inhibitors: a review on the timing of administration and effects on gut hormones. Endocr J. 2019 May 28;66(5):395-401. doi: 10.1507/endocrj.EJ19-0041, PMID 31019154.
Dash RP, Babu RJ, Srinivas NR. Reappraisal and perspectives of clinical drug-drug interaction potential of α-glucosidase inhibitors such as acarbose voglibose and miglitol in the treatment of type 2 diabetes mellitus. Xenobiotica. 2018 Jan;48(1):89-108. doi: 10.1080/00498254.2016.1275063, PMID 28010166.
Elsebai MF, Mocan A, Atanasov AG. Cynaropicrin: a comprehensive research review and therapeutic potential as an anti-hepatitis C virus agent. Front Pharmacol. 2016 Dec 8;7:472. doi: 10.3389/fphar.2016.00472, PMID 28008316.
Xie S, Zhan F, Zhu J, Xu S, Xu J. The latest advances with natural products in drug discovery and opportunities for the future: a 2025 update. Expert Opin Drug Discov. 2025 Jul;20(7):827-43. doi: 10.1080/17460441.2025.2507382, PMID 40391763.
Najmi A, Javed SA, Al Bratty M, Alhazmi HA. Modern approaches in the discovery and development of plant-based natural products and their analogues as potential therapeutic agents. Molecules. 2022 Jan 6;27(2):349. doi: 10.3390/molecules27020349, PMID 35056662.
Tripathi R. In vitro antidiabetic free radical quenching effect and phytochemical profiling of Shankhpushpi with special reference to Chitrakoot region. J Emerg Technol Innovatives Res. 2018;5(7):549-57.
Ho CC, Tan HM. Rise of herbal and traditional medicine in erectile dysfunction management. Curr Urol Rep. 2011 Dec;12(6):470-8. doi: 10.1007/s11934-011-0217-x, PMID 21948222.
Mukherjee U, Sehar U, Brownell M, Reddy PH. Sleep deprivation in dementia comorbidities: focus on cardiovascular disease diabetes anxiety/depression and thyroid disorders. Aging (Albany, NY). 2024 Nov 20;16(21):13409-29. doi: 10.18632/aging.206157, PMID 39571101.
Romano JD, Tatonetti NP. Informatics and computational methods in natural product drug discovery: a review and perspectives. Front Genet. 2019 Apr 30;10:368. doi: 10.3389/fgene.2019.00368, PMID 31114606.
Chaachouay N, Zidane L. Plant-derived natural products: a source for drug discovery and development. Drugs Drug Candidates. 2024 Feb 19;3(1):184-207. doi: 10.3390/ddc3010011.
Ajayi II, Fatoki TH, Alonge AS, Balogun TC, Nwagwe OR, Moge GM. In silico ADME, molecular targets docking and molecular dynamics simulation of key phytoconstituents of lobelia inflata. J Comput Biophys Chem. 2024 Dec 20;23(10):1359-73. doi: 10.1142/S2737416524500480.
Mohanraj K, Karthikeyan BS, Vivek Ananth RP, Chand RP, Aparna SR, Mangalapandi P. IMPPAT: a curated database of Indian medicinal plants phytochemistry and therapeutics. Sci Rep. 2018 Mar 12;8(1):4329. doi: 10.1038/s41598-018-22631-z, PMID 29531263.
Al Khzem AH, Shoaib TH, Mukhtar RM, Alturki MS, Gomaa MS, Hussein D. Repurposing FDA-approved agents to develop a prototype Helicobacter pylori shikimate kinase (HPSK) inhibitor: a computational approach using virtual screening MM-GBSA calculations MD simulations and DFT analysis. Pharmaceuticals (Basel). 2025 Jan 27;18(2):174. doi: 10.3390/ph18020174, PMID 40005988.
Srinivasa MG, Shivakumar KDU, Kumar DU, Mehta CH, Nayak UY, Revanasiddappa BC. In silico studies of (Z)-3-(2-chloro-4-nitrophenyl)-5-(4-nitrobenzylidene)-2-Thioxothiazolidin-4-One derivatives as PPAR-γ agonist: design molecular docking MM-GBSA assay toxicity predictions DFT calculations and MD simulation studies. J Comput Biophys Chem. 2024 Feb 19;23(1):117-36. doi: 10.1142/S2737416523500540.
Srinivasa MG, Pujar KG, Prabhakaran P. Design synthesis and antidiabetic evaluation of thiazolidinedione derivatives as potent PPAR-γ agonists: insights from molecular docking MD simulations and in vitro studies. J Mol Struct. 2025 Apr;1328:141324. doi: 10.1016/j.molstruc.2025.141324.
Mutiah R, Sukardiman MA, Milliana A, Rahmawati E, Fitrianingsih AA, Yueniwati Y. Network pharmacology apoptosis and cell cycle inhibition of sesquiterpene compounds from qusthul Hindi root extract (Saussurea lappa) in breast cancer: an in silico and in vitro approach. Int J Appl Pharm. 2023 Nov 7;15(6):132-41. doi: 10.22159/ijap.2023v15i6.48798.
Baqi MA, Jayanthi K, Rajeshkumar R. Molecular docking insights into probiotics sakacin P and sakacin a as potential inhibitors of the Cox-2 pathway for colon cancer therapy. Int J App Pharm. 2025;17(1):153-60. doi: 10.22159/ijap.2025v17i1.52476.
Sahayarayan JJ, Rajan KS, Vidhyavathi R, Nachiappan M, Prabhu D, Alfarraj S. In-silico protein-ligand docking studies against the estrogen protein of breast cancer using pharmacophore-based virtual screening approaches. Saudi J Biol Sci. 2021 Jan;28(1):400-7. doi: 10.1016/j.sjbs.2020.10.023, PMID 33424323.
Shah BM, Modi P, Trivedi P. Pharmacophore-based virtual screening 3D-QSAR, molecular docking approach for identification of potential dipeptidyl peptidase IV inhibitors. J Biomol Struct Dyn. 2021 Apr;39(6):2021-43. doi: 10.1080/07391102.2020.1750485, PMID 32242496.
Yu Z, Jacobson MP, Friesner RA. What role do surfaces play in GB models? A new-generation of surface-generalized born model based on a novel gaussian surface for biomolecules. J Comput Chem. 2006 Jan 15;27(1):72-89. doi: 10.1002/jcc.20307, PMID 16261581.
Kumar GS, Moustafa M, Sahoo AK, Maly P, Bharadwaj S. Computational investigations on the natural small molecule as an inhibitor of programmed death ligand 1 for cancer immunotherapy. Life (Basel). 2022 Apr 29;12(5):659. doi: 10.3390/life12050659, PMID 35629327.
Daina A, Michielin O, Zoete V. SwissADME: a free web tool to evaluate pharmacokinetics drug-likeness and medicinal chemistry friendliness of small molecules. Sci Rep. 2017 Mar 3;7:42717. doi: 10.1038/srep42717, PMID 28256516.
Yang H, Lou C, Sun L, Li J, Cai Y, Wang Z. admetSAR 2.0: web-service for prediction and optimization of chemical ADMET properties. Bioinformatics. 2019 Mar 15;35(6):1067-9. doi: 10.1093/bioinformatics/bty707, PMID 30165565.
Riyaphan J, Pham DC, Leong MK, Weng CF. In silico approaches to identify polyphenol compounds as α-glucosidase and α-amylase inhibitors against type-II diabetes. Biomolecules. 2021 Dec 14;11(12):1877. doi: 10.3390/biom11121877, PMID 34944521.
Sim L, Quezada Calvillo R, Sterchi EE, Nichols BL, Rose DR. Human intestinal maltase-glucoamylase: crystal structure of the N-terminal catalytic subunit and basis of inhibition and substrate specificity. J Mol Biol. 2008 Jan 18;375(3):782-92. doi: 10.1016/j.jmb.2007.10.069, PMID 18036614.
Proenca C, Freitas M, Ribeiro D, Oliveira EF, Sousa JL, Tome SM. α-glucosidase inhibition by flavonoids: an in vitro and in silico structure-activity relationship study. J Enzyme Inhib Med Chem. 2017 Dec;32(1):1216-28. doi: 10.1080/14756366.2017.1368503, PMID 28933564.
He C, Liu X, Jiang Z, Geng S, Ma H, Liu B. Interaction mechanism of flavonoids and α-glucosidase: experimental and molecular modelling studies. Foods. 2019 Aug 21;8(9):355. doi: 10.3390/foods8090355, PMID 31438605.
Tian LL, Bi YX, Wang C, Zhu K, Xu DF, Zhang H. Bioassay-guided discovery and identification of new potent α-glucosidase inhibitors from Morus alba L. and the interaction mechanism. J Ethnopharmacol. 2024 Mar 25;322:117645. doi: 10.1016/j.jep.2023.117645, PMID 38147942.
Kwon RH, Thaku N, Timalsina B, Park SE, Choi JS, Jung HA. Inhibition mechanism of components isolated from Morus alba branches on diabetes and diabetic complications via experimental and molecular docking analyses. Antioxidants (Basel). 2022 Feb 14;11(2):383. doi: 10.3390/antiox11020383, PMID 35204264.
Liu Y, Zhan L, Xu C, Jiang H, Zhu C, Sun L. α-glucosidase inhibitors from Chinese bayberry (Morella rubra Sieb. et Zucc.) fruit: molecular docking and interaction mechanism of flavonols with different B-ring hydroxylations [α-Glucosidase inhibitors from Chinese bayberry (Morella rubra Sieb. et Zucc.) fruit: molecular docking and interaction mechanism of flavonols with different B-ring hydroxylations]. RSC Adv. 2020 Aug 10;10(49):29347-61. doi: 10.1039/d0ra05015f, PMID 35521141.
Genheden S, Ryde U. The MM/PBSA and MM/GBSA methods to estimate ligand-binding affinities. Expert Opin Drug Discov. 2015 May;10(5):449-61. doi: 10.1517/17460441.2015.1032936, PMID 25835573.
Kikiowo B, Ahmad I, Alade AA, T Ijatuyi T, Iwaloye O, Patel HM. Molecular dynamics simulation and pharmacokinetics studies of ombuin and quercetin against human pancreatic α-amylase. J Biomol Struct Dyn. 2023 Dec;41(20):10388-95. doi: 10.1080/07391102.2022.2155699, PMID 36524470.
Youdim KA, Qaiser MZ, Begley DJ, Rice Evans CA, Abbott NJ. Flavonoid permeability across an in situ model of the blood-brain barrier. Free Radic Biol Med. 2004 Mar 1;36(5):592-604. doi: 10.1016/j.freeradbiomed.2003.11.023, PMID 14980703.