
Int J App Pharm, Vol 18, Issue 2, 2026, 338-344Original Article
EVALUATION OF SERUM LEVELS OF CYP2D6, CYP3A4, GSH, AND MDA AS PREDICTIVE BIOMARKERS FOR METHAMPHETAMINE-INDUCED LIVER DYSFUNCTION USING ADVANCED STATISTICAL MODELING
ALI FARIED SALMAN, GHID HASSAN ABDULHADI*, MUSHTAQ TALIB HASHIM
Department of biochemistry, College of Medicine-University of Baghdad, Iraq
*Corresponding author: Ghid Hassan Abdulhadi; *Email: ghid.h.ah@comed.uobaghdad.edu.iq
Received: 29 Oct 2025, Revised and Accepted: 05 Jan 2026
ABSTRACT
Objective: This study aimed to evaluate the predictive potential of serum protein levels of CYP2D6, CYP3A4, glutathione (GSH), and malondialdehyde (MDA) as early biomarkers of METH-induced liver dysfunction using multivariate statistical approaches, compared to standard LFTs.
Methods: This case–control study included 90 METH users and 45 healthy control individuals. Serum CYP2D6, CYP3A4, GSH, and MDA levels, and routine LFTs were determined. Analysis was performed using principal component analysis (PCA), receiver operating characteristic (ROC) curves, and decision tree modeling.
Results: METH users showed significantly decreased serum levels of CYP2D6, CYP3A4, and GSH and higher MDA levels (p<0.0001), suggestive of oxidative metabolic disequilibrium. While most routine LFTs were normal, other markers, including AST, AST/ALT ratio, and albumin-to-globulin ratio, increased significantly. PCA demonstrated that early injury components could be divided into separate groups according to the markers of their metabolic and oxidative components. CYP2D6 ≤ 1.98 ng/ml was the best discriminating predictor according to the decision tree with 93.3% accuracy (AUC = 94).
Conclusion: Serum protein levels of CYP2D6, CYP3A4, GSH, and MDA demonstrated high sensitivity for detecting subclinical liver injury in METH users before conventional LFTs became abnormal. Their integration into diagnostic models may facilitate early interventions in high-risk populations.
Keywords: Methamphetamine, Cytochrome P-450 CYP2D6, Cytochrome P-450 CYP3A4, Glutathione, Malondialdehyde, Principal component analysis
© 2026 The Authors.Published by Innovare Academic Sciences Pvt Ltd. This is an open access article under the CC BY license (https://creativecommons.org/licenses/by/4.0/)
DOI: https://dx.doi.org/10.22159/ijap.2026v18i2.57318 Journal homepage: https://innovareacademics.in/journals/index.php/ijap
INTRODUCTION
Cytochrome P450 enzymes (CYPs) are a group of heme-containing enzymes that play an important role in the metabolism of many drugs and other xenobiotics. CYP enzymes are located in the endoplasmic reticulum of cells throughout the body but are most abundant in the liver. CYP enzymes can catalyze an extensive range of reactions including oxidation, reduction, hydrolysis, and isomerization [1]. Of the 57 potentially functional human CYP enzymes, members of the CYP1, CYP2, and CYP3 families account for nearly 80% of clinically used drugs [2]. CYP-dependent metabolism not only transforms lipophilic compounds into water-soluble forms for easier elimination but also influences therapeutic outcomes by modulating drug efficacy, safety, bioavailability, and resistance, both in major metabolic organs and at local sites of drug action, with CYP3A4 and CYP2D6 contributing to over 50% of CYP-related drug metabolism [3, 4].
Among the CYP isoforms, CYP2D6 and CYP3A4 have received a great deal of clinical attention mainly because they are two important enzymatic members in phase I studies (75% drugs) with unique regulatory pathways [5]. 2D6 mediates the oxidative metabolism of 20-25% of drugs used in a clinical setting. Most individuals have highly variable enzymatic activity owing to genetic polymorphisms, which can lead to phenotypes of poor or ultrarapid metabolizers [6, 7]. In contrast, the most abundant hepatic isoform, CYP3A4, is involved in metabolism in more than 30% of clinically recommended drugs. In contrast to CYP2D6, factors other than genotype influence their activity more substantially [5, 8, 9].
Methamphetamine (METH), a highly addictive psychostimulant, was originally synthesized from its related compound amphetamine [10]. The metabolism of METH is primarily mediated by hepatic CYPs, particularly CYP2D6 and CYP3A4, which convert it to its main metabolites amphetamine and 4-hydroxyamphetamine [10]. This metabolic process, which is essential for detoxification, is also a source of cellular stress. Enzymatic conversion can lead to the production of reactive oxygen species (ROS), a process known as bioactivation. This contributes to a state of oxidative stress that, in conjunction with other pathways, such as the activation of intracellular nitric oxide (NO), can overwhelm cellular antioxidant defenses and lead to hepatocyte injury and apoptotic cell death [11].
Exposure to METH is associated with oxidative stress, characterized by increased malondialdehyde (MDA) levels, a marker of lipid peroxidation, and decreased glutathione (GSH) levels, a vital intracellular antioxidant [12, 13]. The imbalance between MDA and GSH levels reflects oxidative damage, which plays a central role in METH-induced hepatotoxicity pathogenesis [13].
Principal component analysis (PCA) is a powerful multivariate statistical technique that reduces data dimensionality and explores latent patterns within complex datasets. PCA is widely used, especially in medical informatics, to discover underlying pathological axes grouping related biomarkers, thus providing exposure points for critical biochemical pathways [14, 15]. Concurrently, decision tree models present a method of supervised machine learning that classifies samples according to the most discriminative variable splitting, thereby providing interpretable prediction criteria [14]. Combining both methods would result in a better conception and categorization of METH-induced hepatic changes by biochemical parameters [14-16].
Despite growing recognition of the impact of METH on liver function and CYPs regulation, integrative studies that simultaneously examine its biochemical, enzymatic, and toxicological consequences within a single experimental framework are lacking [17]. To the best of our knowledge, multivariate statistical approaches, such as PCA and decision tree modeling, have rarely been used to identify predictive biomarkers or classification patterns in METH-induced hepatic dysfunction [18, 19]. Therefore, this study aimed to evaluate the detection and classification potential of serum levels of the metabolic enzymes CYP2D6 and CYP3A4, the oxidative stress markers GSH and MDA, and conventional liver function tests (LFTs) in METH-induced hepatic dysfunction using PCA and decision tree classification as advanced statistical tools.
MATERIALS AND METHODS
Subject
This case-control study was conducted between January and June 2025. A total of 135 participants were enrolled, comprising 90 patients diagnosed with METH, and 45 age-and BMI-matched healthy individuals who served as controls. The participants were aged between 18 and 40 y old.
The inclusion criteria for the METH group were a confirmed diagnosis of METH use disorder by a psychiatrist according to the DSM-5 criteria, a history of chronic METH use (≥3 times per week for at least one year), and a positive urine screening test for amphetamines at the time of recruitment. The route of administration (smoking, injection, or oral administration) was not systematically recorded for all the participants. Exclusion criteria for all participants included known pre-existing liver diseases (e. g., hepatitis B/C and cirrhosis), comorbid substance use disorders (except nicotine), HIV infection, current use of medications known to affect CYPs activity or liver function, and any chronic medical conditions (e. g., diabetes and cardiovascular disease).
Ethical approval and informed consent
The study protocol was reviewed and approved by the Ethical Committee of the Al-Qanat Center for Social Rehabilitation in Baghdad, Iraq. This study was conducted in accordance with the ethical principles of the Declaration of Helsinki. Written informed consent was obtained from all the participants after the nature of the study procedure was fully explained. (IRB 1068 12/01/2025)
Blood sample collection and biochemical analysis
Venous peripheral blood samples (5 ml) were collected from all participants. The samples were allowed to clot at room temperature for 30 min before being centrifuged at 1000 × g for 20 min to separate the serum. The collected serum was promptly aliquoted into sterile Eppendorf tubes and stored at − 80 °C until analysis. This procedure was implemented to minimize the potential proteolytic degradation of enzyme proteins and prevent the oxidation of sensitive analytes, such as glutathione [20].
Biochemical assays
Liver function tests
LFTs, such as alanine aminotransferase (ALT), aspartate aminotransferase (AST), alkaline phosphatase, and total serum bilirubin (TSB), were performed on the same day of blood collection using a Beckman Coulter 480 AU Chemistry Analyzer (Beckman Coulter, Germany).
ELISA kit validation
To ensure the reliability of ELISA data, the performance of each kit was validated in our laboratory. The intra-and inter-assay CVs values were<8% and<10%, respectively, for all analytes. Spike-and-recovery experiments demonstrated recovery rates between 90 and 110%. The assay also showed excellent linearity over the tested concentration range (R²>0.99).
ELISA-based measurements
The serum protein concentrations of CYP2D6 (catalog no. EH0783; Fine Test, China), and CYP3A4 (Catalog No. EH0777; Fine Test, China) were measured using commercial sandwich ELISA kits. The serum levels of reduced glutathione (GSH) (catalog no. EH3524; Fine Test, China) and malondialdehyde (MDA) (Catalog No. ELK1135; ELK Biotechnology, China) was determined using a competitive inhibition ELISA kit. The ELISA measurements were conducted using a BioTek ELISA microplate reader (BioTek Instruments, USA). All assays were performed strictly in accordance with the manufacturer’s protocols. To ensure the reliability of the ELISA data, the performance of each kit was validated using pooled serum samples. Intra-assay precision was determined by running 10 replicates of the same sample in a single assay, while inter-assay precision was determined by running the same sample in duplicate across five independent assay runs. The coefficients of variation (CVs) were within acceptable limits for all biomarkers.
Statistical analysis
The data were subjected to statistical analysis using the IBM SPSS Statistics software (version 27, IBM Corp., Armonk, NY, USA). Data are expressed as mean±SD. Comparisons between patients and normal controls were carried out by t-test, and p<0.001 and<0.05 were considered to be highly significant or significant differences, respectively. The Pearson's coefficient (r) was used to express the correlation between the studied parameters. The sensitivity and specificity of the tests were evaluated, and the cut-off value was established using receiver operating characteristic (ROC) curve analysis (performed using the ROC Curve module in SPSS). The interrelationships among biochemical variables were estimated using principal component analysis (PCA) with the Factor Analysis module in SPSS, and the potential interaction patterns among them were determined by a decision tree using the Chi-squared Automatic Interaction Detection (CHAID) method (performed using the Decision Tree module in SPSS) [21]. All advanced statistical analyses (ROC, PCA, and CHAID) were performed using SPSS.
RESULTS
There was no statistically significant difference in age and body mass index (BMI) between the METH use disorder and control groups (table 1). Patients had significantly lower serum protein levels of CYP2D6 and CYP3A4 and lower serum GSH levels than controls (p<0.0001). They also had significantly higher levels of MDA, AST/ALT, albumin, globulin, and A/G ratio.
Table 1: Comparison of demographic, biochemical and enzymatic markers between METH users and healthy controls
| Parameters | Control group (n=45) mean±SD | METH patients (n=90) mean±SD | P value |
| Age (years) | 27.75±7.49 | 30.25±6.15 | 0.096 |
| BMI (kg/m²) | 22.64±2.30 | 23.44±2.47 | 0.064 |
| CYP2D6 (ng/ml) | 4.17±0.92 | 1.145±0.69 | <0.0001** |
| CYP3A4 (pg/ml) | 1539±448.65 | 483.43±180.5 | <0.0001** |
| MDA (ng/ml) | 21.63±1.511 | 27.70±2.80 | <0.0001** |
| GSH (ng/ml) | 101.43±1.74 | 59.01±9.34 | <0.0001** |
| ALT (U/l) | 28.06±8.57 | 31.2±14.82 | 0.123 |
| AST (U/l) | 21.95±8.00 | 26.5±13.20 | 0.01* |
| AST/ALT | 0.810±0.26 | 1.008±0.61 | 0.009* |
| ALP (U/l) | 82.53±14.30 | 88.34±31.41 | 0.142 |
| TSB (mg/dl) | 0.86±0.193 | 0.84±0.378 | 0.807 |
| A/G | 1.35±0.08 | 1.54±0.466 | 0.00022* |
Data are presented as mean±SD. *P<0.05 is considered significant. **P<0.001 is considered highly significant. Abbreviations: BMI, body mass index; CYP2D6, cytochrome P450 2D6; CYP3A4, cytochrome P450 3A4; MDA, malondialdehyde; GSH, glutathione; ALT, alanine aminotransferase; AST, aspartate aminotransferase; ALP, alkaline phosphatase; TSB, total serum bilirubin; A/G, albumin-to-globulin ratio.
CYP2D6 demonstrated high diagnostic performance (AUC = 0.995, sensitivity and specificity = 95.6%). MDA had excellent predictive ability, with an AUC of 0.967, sensitivity of 95.6%, and moderate specificity (77.8%). Notably, CYP3A4 and GSH showed perfect classification performance (AUC = 1.000) in the training dataset. Although these results are promising, perfect classifiers are rare in clinical practice and may reflect the homogeneity of our study cohorts or potential overfitting. All AUCs were highly significant (p<0.001), as shown in table 2.
Table 2: Diagnostic performance of biomarkers for differentiating METH users from controls
| Marker | AUC | Cut-off | Sensitivity (%) | Specificity (%) | Accuracy (%) | P-Value | 95% CL |
| CYP2D6 | 0.995 | 2.6100 | 95.6% | 95.6% | 95.6% | <0.001 | 0.989-1.000 |
| CYP3A4 | 1.000 | 849.37 | 100% | 100% | 100% | <0.001 | 1.000-1.000 |
| GSH | 1.000 | 88.145 | 100% | 100% | 100% | <0.001 | 1.000-1.000 |
| MDA | 0.967 | 23.85 | 95.6% | 77.8% | 89.6% | <0.001 | 0.941-0.994 |
Abbreviations: AUC, area under the curve; CL, confidence limit; CYP2D6, cytochrome P450 2D6; CYP3A4, cytochrome P450 3A4; GSH, glutathione; MDA, malondialdehyde. PCA was conducted on ten biochemical variables, including metabolic enzymes, oxidative stress indicators, and liver function markers (table 3). The Kaiser-Meyer-Olkin (KMO) measure of sampling adequacy was 0.591, indicating mediocre suitability for factor analysis.
Table 3: Suitability of data for principal component analysis (KMO and bartlett's test)
| Kaiser-meyer-olkin measure of sampling adequacy | .591 |
| Bartlett’s test of sphericity | Approx. Chi-Square |
| Df | |
| Sig. |
Abbreviations: KMO, Kaiser-Meyer-Olkin; Df, degrees of freedom; Sig., significance. Given the median KMO value (0.591), three factors with eigenvalues greater than 1 were retained, contributing to 71.305% of the total variance (table 4). The first, second, and third components explained 33.965%, 23.107%, and 14.233% of the variance, respectively.
Table 4: Total variance explained by the extracted principal components
| Component | Initial eigenvalues | Extraction sums of squared loadings | ||||
| Total | % of variance | Cumulative % | Total | % of variance | Cumulative % | |
| 1 | 3.396 | 33.965 | 33.965 | 3.396 | 33.965 | 33.965 |
| 2 | 2.311 | 23.107 | 57.072 | 2.311 | 23.107 | 57.072 |
| 3 | 1.423 | 14.233 | 71.305 | 1.423 | 14.233 | 71.305 |
| 4 | .917 | 9.165 | 80.470 | |||
| 5 | .598 | 5.983 | 86.453 | |||
| 6 | .522 | 5.219 | 91.672 | |||
| 7 | .459 | 4.589 | 96.261 | |||
| 8 | .198 | 1.979 | 98.240 | |||
| 9 | .124 | 1.240 | 99.480 | |||
| 10 | .052 | .520 | 100.000 | |||
Extraction method: principal component analysis.
The rotated component matrix (varimax rotation) is listed in table 5. While the KMO value suggests only mediocre sampling adequacy, the variables were grouped as follows for exploratory purposes: component 1 (33.97%) was associated with oxidative stress and serum cytochrome protein levels, with strong positive loadings for CYP2D6 (0.917), CYP3A4 (0.913), and GSH (0.900), and a strong negative loading for MDA (-0.768). Component 2 (23.11%): Traditional automatic liver function parameters such as ALT (0.732), AST (0.782), ALP (0.762), and TSB (0.766). Component 3 (14.23%) was primarily driven by the AST/ALT ratio (0.923) and ALT (0.544), whereas it was enriched with AST (0.391) and A/G ratio (0.436)}.
Table 5: Rotated component matrix showing factor loadings of biochemical variables
| Biochemical variables | Component | ||
| 1 | 2 | 3 | |
| ALT (U/l) | -.080 | .732 | -.544 |
| AST (U/l) | -.186 | .782 | .391 |
| AST/ALT | -.216 | .007 | .923 |
| ALP (U/l) | -.077 | .762 | -.024 |
| TSB (mg/dl) | .073 | .766 | -.175 |
| MDA (ng/ml) | -.768 | .139 | .008 |
| GSH (ng/ml) | .900 | -.004 | .030 |
| CYP2D6 (ng/ml) | .917 | -.086 | -.051 |
| CYP3A4 (pg/ml) | .913 | .034 | .048 |
| A/G Ratio | -.281 | .089 | -.436 |
Rotation Method: Varimax with Kaiser Normalization. a. Rotation converged in 5 iterations.
To visualize the distinction between METH users and healthy controls, a two-dimensional PCA plot was generated using the first two principal components, which collectively explained the largest proportion of total variance. Although the KMO value indicated only mediocre data suitability for PCA, as depicted in fig. 1, the score plot of the first two PCs revealed a distinct separation between the two groups. The METH user cohort and the control group formed separate clusters that were primarily distinguished along the axis of first principal component 1.
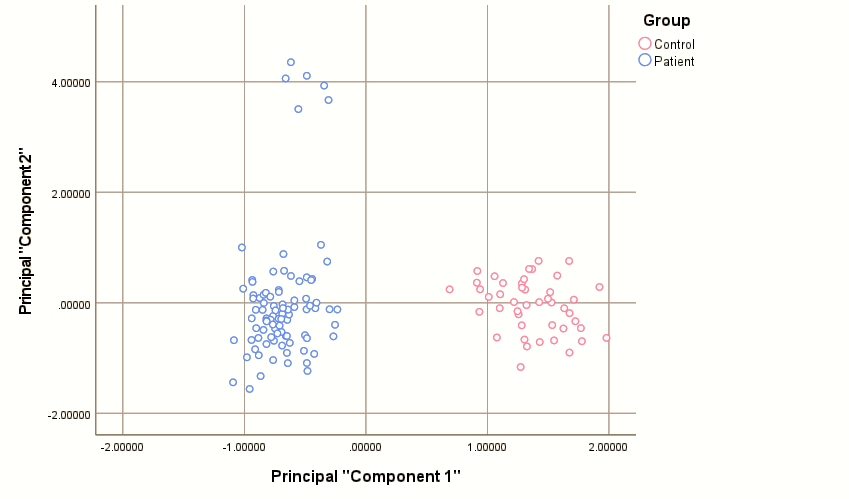
Fig. 1: PCA score plot of the first two principal components. the plot shows a clear separation between METH users (red) and healthy controls (blue), with the distinction occurring primarily along the first principal component (PC1), which represents an axis of oxidative stress and cytochrome P450 protein levels
A decision tree classification model was constructed using the Chi-squared Automatic Interaction Detection (CHAID) algorithm to identify the most significant predictors for differentiating METH users from healthy individuals. The algorithm identified CYP2D6 as the most statistically significant predictor (χ² = 101.25, df=1, p<0.001), with an optimal cutoff value of 1.980 ng/ml (fig. 2).
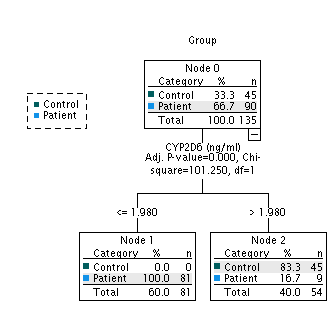
Fig. 2: CHAID decision tree model for classifying participants. The model identifies a single, optimal split point based on the serum level of CYP2D6 (≤ 1.980 ng/ml) to differentiate METH users from healthy controls
As shown in table 7, the model achieved a sensitivity of 90% and specificity of 100% on the training data, with an overall classification accuracy of 93.3%. It is important to note that these performance metrics represent the training set results without an independent test set validation or cross-validation. This suggests that CYP2D6, using a threshold of ≤1.980 ng/ml, may serve as a discriminating biomarker for differentiating METH users from controls within our study population, although external validation is required to confirm its generalizability.
Table 7: Classification performance of the CHAID decision tree model
| Actual \ Predicted | Predicted |
| Predicted control | |
| Actual Control | 45 (TN) |
| Actual Patient | 9(FN) |
| Overall Percentage | 40.0% |
Abbreviations: TN, true negative; FP, false positive; FN, false negative; TP, true positive.
DISCUSSION
The present study introduced a significant advancement in monitoring METH-induced hepatotoxicity by demonstrating that serum levels of CYP2D6 and CYP3A4, in conjunction with markers of oxidative stress, serve as highly sensitive and early predictors of subclinical liver injury. Our central and most novel finding is the profound decrease in circulating concentrations of these key metabolic enzymes in METH users, a change that occurs before conventional LFTs become abnormal. By employing advanced statistical models, we not only confirmed the predictive power of these biomarkers but also elucidated a hierarchical pattern of injury, positioning metabolic and oxidative dysregulation as primary events in METH-induced hepatic pathology.
The significant reduction in serum CYP2D6 and CYP3A4 protein levels was the cornerstone of our findings. Historically, a major limitation in using CYPs as biomarkers is that they are primarily intracellular membrane-bound proteins within the liver, making their serum concentration an unconventional measure of hepatic function. However, our results are strongly supported by an emerging body of evidence demonstrating that CYP enzymes are packaged into extracellular vesicles (EVs) or exosomes and released into circulation [22, 23]. Groundbreaking work has shown that circulating EV-containing CYPs are metabolically active, and their levels can change in response to liver damage [24]. For instance, studies have shown that circulating CYP activity increases following drug-induced liver injury (DILI), suggesting that the release of these enzymes reflects the underlying hepatic stress [25, 26].
Therefore, decreased serum CYP levels observed in METH users may represent a novel pathophysiological mechanism. This could be attributed to several factors, including METH-induced transcriptional downregulation of hepatic synthesis, altered packaging of CYPs into exosomes, or accelerated clearance of circulating vesicles [27, 28]. While the precise mechanism requires further investigation, our study provides compelling evidence that measuring circulating CYP protein levels is a viable and sensitive noninvasive strategy for detecting early hepatic dysregulation, aligning with research that calls for the use of CYP activity to evaluate acute hepatic dysfunction [25, 29, 30].
The present findings of decreased GSH and increased MDA levels corroborate the well-established role of oxidative stress in METH-induced toxicity [31]. Chronic METH exposure generates ROS, which depletes cellular antioxidants such as GSH and causes lipid peroxidation, as indicated by elevated MDA [32]. Turan et al. (2023) investigated oxidative stress biomarkers in 50 METH users versus 36 controls and demonstrated significantly elevated serum thiol/disulfide ratios and ischemia-modified albumin levels [33]. Their research supports the present findings by establishing that multiple oxidative stress parameters are simultaneously altered in METH users, reinforcing the concept of a unified metabolic-redox injury axis. According to Raspopović et al. (FAN, 2024), GSH is considered a sensitive marker of oxidative stress with the ability to reflect initial cellular damage before cell injury becomes readily apparent in organs [34]. Similarly, Cherian et al. [2019] found malondialdehyde to be a good marker of lipid peroxidation under inflammatory conditions, indicating that the strong negative correlation between these two variables in the heatmap indicates that METH-induced oxidative damage involves concurrent antioxidant exhaustion and membrane injury [35]. This bimodal pattern may indicate a more realistic picture of early drug-induced liver injury.
As a secondary observation, we noted modest but statistically significant increases in the AST/ALT and albumin-to-globulin (A/G) ratios, despite most conventional LFTs remaining within the normal range. While an AST/ALT ratio greater than 1 may suggest mitochondrial injury [36, 37] and an elevated A/G ratio is atypical, the clinical significance of these findings in the context of METH use remains unclear and requires further investigation [38, 39]. Given their limited diagnostic relevance compared to primary biomarkers (CYP2D6, CYP3A4, GSH, and MDA), these observations should be considered exploratory [36].
The integrated application of PCA and decision tree modeling provided a deeper interpretation of the biomarker data. PCA stratified the injury process into distinct stages: PC1 represented the initial metabolic-oxidative shift, and PC2 (loaded with ALT, AST, etc.) represented subsequent hepatocellular injury. PC3, which was primarily driven by the AST/ALT ratio, may reflect additional injury patterns, although its interpretation remains exploratory, given the limited diagnostic relevance of this ratio in our cohort.
The decision tree model is more useful for the diagnosis of biomarkers of liver disease. In the present study, CHAID decision tree analysis identified CYP2D6 as the best discriminating variable between METH users and controls in our training dataset, with an optimal cutoff value for prediction of ≤1.98 ng/ml. This enzymatic function as a sensitive marker is also supported by clinical case reports, for example, a syndrome of persistent paroxetine toxicity associated with a phenoconversion event that profoundly decreased CYP2D6 metabolism [40]. This clinical observation is supported by experimental models showing that transcriptional downregulation of CYP2D6 is an early event in DILI [41]. The versatility of tree-based modeling has also been demonstrated in oncological applications, where ensemble models have efficiently identified diverse biomarkers for hepatocellular carcinoma with high accuracy AUC values (up to 0.997) across a number of datasets [42]. This joint evidence further emphasizes the potential of multivariate statistical methodologies that integrate feature optimization and ensemble learning for biomarker mining in hepatology. However, rigorous external validation is essential for clinical translation.
Limitations and future research directions
This study has several limitations that guide future research. The primary limitation is the measurement of serum CYP protein concentrations rather than direct hepatic enzymatic activity. However, by contextualizing our findings within the latest research on circulating exosomal CYPs [22, 23, 43, 44], we reframe this not as a simple surrogate measure but as a novel and potentially more specific indicator of hepatic stress. Future studies should aim to correlate serum protein levels with the direct characterization of hepatic exosomes to validate this mechanism.
Statistically, the perfect AUCs of 1.00 for CYP3A4 and GSH suggest potential overfitting, and the median KMO value (0.591) for our PCA indicates that the results should be interpreted with caution. Furthermore, a decision tree model was developed and evaluated on the same training dataset without independent test set validation or cross-validation procedures. While this model is highly interpretable, its reliance on a single predictor and lack of external validation limit conclusions about its generalizability and predictive performance in independent populations. Future studies should employ robust cross-validation techniques and explore more complex ensemble models, such as Random Forests, to build a more resilient predictive signature for METH-induced hepatotoxicity.
CONCLUSION
In conclusion, this study marks a shift in the understanding of METH-induced hepatic effects, revealing a subtle pattern of subclinical liver dysfunction that cannot be detected by conventional screening. The simultaneous decrease in the serum protein levels of CYP2D6 and CYP3A4, along with changes in oxidative stress markers, revealed an early biological injury occurring at the molecular level before clinical and biochemical abnormalities became apparent. It is important to note that this study measured serum protein levels rather than direct hepatic enzymatic activity. Advanced statistical models, specifically PCA and decision tree modeling, marked a turning point in data interpretation, identifying CYP2D6 as an early pivotal marker to better differentiate abusers from healthy controls. Beyond improving the classification accuracy, these models provide new ways to understand the hidden patterns of liver injury. Accordingly, integrating oxidative stress markers and cytochrome enzymes with predictive analytics tools is a necessary step toward developing early screening strategies capable of intervening before damage progresses to irreversible stages.
ACKNOWLEDGEMENT
The authors extend their sincere gratitude to the participants for their voluntary involvement. We also thank the staff and the ethical committee of the Al-Qanat Center for Social Rehabilitation in Baghdad for their support and facilitation. Special thanks to the laboratory technicians for their meticulous work in sample processing and analysis.
ETHICAL APPROVAL
The study protocol was reviewed and approved by the Ethical Committee of the Al-Qanat Center for Social Rehabilitation in Baghdad, Iraq. This study was conducted in accordance with the ethical principles of the Declaration of Helsinki. Written informed consent was obtained from all the participants after the nature of the study procedure was fully explained. (IRB 1068 12/01/2025).
FUNDING
No funds were received to fulfil this work. We have not used any AI tools or technologies to prepare this manuscript.
AUTHORS CONTRIBUTIONS
Conceptualization, AFS and GHA; methodology, AFS, GHA, and MTH; validation, AFS, GHA, and MTH; formal analysis, MTH; investigation, AFS, GHA, and MTH; resources, AFS; data curation, MTH; writing – original draft preparation, AFS and GHA; writing – review and editing, all authors; visualization, MTH; supervision, AFS; project administration, AFS; funding acquisition, AFS, and GHA. All the authors have read and agreed to the published version of the manuscript.
CONFLICTS OF INTERESTS
The authors have nothing to disclose
REFERENCES
Kondza M, Brizic I, Jokic S. Flavonoids as CYP3A4 inhibitors in vitro. Biomedicines. 2024 Mar;12(3):644. doi: 10.3390/biomedicines12030644, PMID 38540257.
Elfaki I, Almutairi FM, Mir R, Khan R, Abu-duhier F. Cytochrome P450 CYP1B12 gene and its association with T2D in tabuk population, northwestern region of Saudi Arabia. Asian J Pharm Clin Res. 2018 Jan 1;11(1):55-9. doi: 10.22159/ajpcr.2018.v11i1.21657.
Zhao M, Ma J, Li M, Zhang Y, Jiang B, Zhao X. Cytochrome P450 enzymes and drug metabolism in humans. Int J Mol Sci. 2021 Nov;22(23):12808. doi: 10.3390/ijms222312808, PMID 34884615.
Kumar A, Estrada DF. Structural basis of bidirectional allostery across the heme in a cytochrome P450 enzyme. J Biol Chem. 2023;299(8):104977. doi: 10.1016/j.jbc.2023.104977, PMID 37390989.
Loos NH, Beijnen JH, Schinkel AH. The mechanism-based inactivation of CYP3A4 by ritonavir: what mechanism? Int J Mol Sci. 2022 Aug;23(17):9866. doi: 10.3390/ijms23179866, PMID 36077262.
Rohmanika R, Arozal W, Louisa M, Gayatri A, Kekalih A, Marzuki HY JE. Patterns of methamphetamine use and its associations with psychiatric symptoms in patients upon admission at the national rehabilitation center lido bogor Indonesia. Int J Appl Pharm. 2022;14(5):96-100. doi: 10.22159/ijap.2022.v14s5.16.
Frederiksen T. Using population pharmacokinetic analyses of drugs metabolized by CYP2D6 to study the genotype-phenotype translation. Basic Clin Pharmacol Toxicol. 2023 Aug;133(2):113-23. doi: 10.1111/bcpt.13903, PMID 37221697.
Tu DZ, Hu XY, Lei JX, Liu SY, Xiao ZP, Yang L. A patent review of CYP3A4 inhibitors (2018-present). Expert Opin Ther Pat. 2025 May;35(5):503-13. doi: 10.1080/13543776.2025.2470294, PMID 39976548.
Yoon S, Jeong S, Jung E, Kim KS, Jeon I, Lee Y. Effect of CYP3A4 metabolism on sex differences in the pharmacokinetics and pharmacodynamics of zolpidem. Sci Rep. 2021 Sep;11(1):19150. doi: 10.1038/s41598-021-98689-z, PMID 34580385.
Ma Y, Wu H, Wang H, Chen F, Xie Z, Zhang Z. Psychiatric comorbidities and liver injury are associated with unbalanced plasma bile acid profile during methamphetamine withdrawal. Front Endocrinol (Lausanne). 2021;12:801686. doi: 10.3389/fendo.2021.801686, PMID 35046900.
Lewis D, Kenneally M, Van Denheuvel C, Byard RW. Methamphetamine deaths: changing trends and diagnostic issues. Med Sci Law. 2021 Apr;61(2):130-7. doi: 10.1177/0025802420986707, PMID 33423599.
Al-Hakeim HK, Altufaili MF, Almulla AF, Moustafa SR, Maes M. Increased lipid peroxidation and lowered antioxidant defenses predict methamphetamine induced psychosis. Cells. 2022 Nov;11(22):3694. doi: 10.3390/cells11223694, PMID 36429122.
Ramli FF, Rejeki PS, Ibrahim N, Abdullayeva G, Halim S. A mechanistic review on toxicity effects of methamphetamine. Int J Med Sci. 2025;22(3):482-507. doi: 10.7150/ijms.99159, PMID 39898237.
Song YY, Lu Y. Decision tree methods: applications for classification and prediction. Shanghai Arch Psychiatry. 2015 Apr;27(2):130-5. doi: 10.11919/j.issn.1002-0829.215044, PMID 26120265.
Dida N, Kassa Y, Sirak T, Zerga E, Dessalegn T. Substance use and associated factors among preparatory school students in bale zone oromia regional state southeast Ethiopia. Harm Reduct J. 2014 Aug;11(21):21. doi: 10.1186/1477-7517-11-21, PMID 25108629.
Jolliffe IT, Cadima J. Principal component analysis: a review and recent developments. Philos Trans A Math Phys Eng Sci. 2016 Apr;374(2065):20150202. doi: 10.1098/rsta.2015.0202, PMID 26953178.
Dostalek M, Jurica J, Pistovcakova J, Hanesova M, Tomandl J, Linhart I. Effect of methamphetamine on cytochrome P450 activity. Xenobiotica. 2007 Dec;37(12):1355-66. doi: 10.1080/00498250701652877, PMID 17922362.
Yan B, Ye X, Wang J, Han J, Wu L, He S. An algorithm framework for drug-induced liver injury prediction based on genetic algorithm and ensemble learning. Molecules. 2022 May;27(10):3112. doi: 10.3390/molecules27103112, PMID 35630587.
Zhang KK, Wang H, Qu D, Chen LJ, Wang LB, Li JH. Luteolin alleviates methamphetamine-induced hepatotoxicity by suppressing the p53 pathway-mediated apoptosis, autophagy and inflammation in rats. Front Pharmacol. 2021;12:641917. doi: 10.3389/fphar.2021.641917, PMID 33679421.
Rifai N. Tietz textbook of clinical chemistry and molecular diagnostics. 7th ed. St. Louis, MO: Elsevier Health Sciences; 2017. Available from: https://www.nursingbridgetestbank.Com/wpcontent/uploads/2025/12/Download-Sample.pdf.
Elliott AC, Woodward WA. Statistical analysis quick reference guidebook: with SPSS examples. Sage Publications; 2007.
Fu S, Wu D, Jiang W, Li J, Long J, Jia C. Molecular biomarkers in drug-induced liver injury: challenges and future perspectives. Front Pharmacol. 2020;10:1667. doi: 10.3389/fphar.2019.01667, PMID 32082163.
Gerth K, Kodidela S, Mahon M, Haque S, Verma N, Kumar S. Circulating extracellular vesicles containing xenobiotic metabolizing CYP enzymes and their potential roles in extrahepatic cells via cell–cell interactions. Int J Mol Sci. 2019;20(24):6178. doi: 10.3390/ijms20246178, PMID 31817878.
Palomo L, Mleczko JE, Azkargorta M, Conde Vancells J, Gonzalez E, Elortza F. Abundance of cytochromes in hepatic extracellular vesicles is altered by drugs related with drug-induced liver injury. Hepatol Commun. 2018;2(9):1064-79. doi: 10.1002/hep4.1210, PMID 30202821.
Soo JY, Wiese MD, Dyson RM, Gray CL, Clarkson AN, Morrison JL. Methamphetamine administration increases hepatic CYP1A2 but not CYP3A activity in female guinea pigs. PLOS One. 2020;15(5):e0233010. doi: 10.1371/journal.pone.0233010, PMID 32396581.
Loftis JM, Janowsky A. Neuroimmune basis of methamphetamine toxicity. Int Rev Neurobiol. 2014;118:165-97. doi: 10.1016/B978-0-12-801284-0.00007-5, PMID 25175865.
Wang L, Bai M, Jin T, Zheng J, Wang Y, He Y. Effects of CYP3A4 polymorphisms on drug addiction risk among the Chinese han population. Front Public Health. 2019;7:315. doi: 10.3389/fpubh.2019.00315, PMID 31799230.
Zhou S, Cheng K, Peng Y, Liu Y, Hu Q, Zeng S. Regulation mechanism of endoplasmic reticulum stress on metabolic enzymes in liver diseases. Pharmacol Res. 2024 Sep;207:107332. doi: 10.1016/j.phrs.2024.107332, PMID 39089398.
Harbrecht BG, Frye RF, Zenati MS, Branch RA, Peitzman AB. Cytochrome P-450 activity is differentially altered in severely injured patients. Crit Care Med. 2005;33(3):541-6. doi: 10.1097/01.CCM.0000155989.54344.E0, PMID 15753745.
Hossam Abdelmonem B, Abdelaal NM, Anwer EK, Rashwan AA, Hussein MA, Ahmed YF. Decoding the role of CYP450 enzymes in metabolism and disease: a comprehensive review. Biomedicines. 2024 Jul;12(7):1467. doi: 10.3390/biomedicines12071467, PMID 39062040.
McDonnell-Dowling K, Kelly JP. The role of oxidative stress in methamphetamine induced toxicity and sources of variation in the design of animal studies. Curr Neuropharmacol. 2017;15(2):300-14. doi: 10.2174/1570159x14666160428110329, PMID 27121285.
Yamamoto BK, Moszczynska A, Gudelsky GA. Amphetamine toxicities: classical and emerging mechanisms. Ann N Y Acad Sci. 2010 Feb;1187:101-21. doi: 10.1111/j.1749-6632.2009.05141.x, PMID 20201848.
Turan C, Senormancı G, Neselioglu S, Budak Y, Erel O, Senormancı O. Oxidative stress and inflammatory biomarkers in people with methamphetamine use disorder. Clin Psychopharmacol Neurosci. 2023 Aug;21(3):572-82. doi: 10.9758/cpn.22.1047, PMID 37424424.
Bogoje Raspopovic A, Balta V, Vodopic M, Drobac M, Boros A, Dikic D. The possible role of oxidative stress marker glutathione in the assessment of cognitive impairment in multiple sclerosis. Open Med (Wars). 2024;19(1):20240952. doi: 10.1515/med-2024-0952, PMID 38623459.
Cherian DA, Peter T, Narayanan A, Madhavan SS, Achammada S, Vynat GP. Malondialdehyde as a marker of oxidative stress in periodontitis patients. J Pharm Bioallied Sci. 2019 May;11(Suppl 2):S297-300. doi: 10.4103/JPBS.JPBS_17_19, PMID 31198357.
Eskandari MR, Rahmati M, Khajeamiri AR, Kobarfard F, Noubarani M, Heidari H. A new approach on methamphetamine-induced hepatotoxicity: involvement of mitochondrial dysfunction. Xenobiotica. 2014 Jan;44(1):70-6. doi: 10.3109/00498254.2013.807958, PMID 23786375.
Ajoolabady A, Kaplowitz N, Lebeaupin C, Kroemer G, Kaufman RJ, Malhi H. Endoplasmic reticulum stress in liver diseases. Hepatology. 2023 Feb;77(2):619-39. doi: 10.1002/hep.32562, PMID 35524448.
Mohammed NS, Ali ZQ, Mohamed AS, Mirza SA. The impact of methamphetamine on liver injury in Iraqi male addicts. Toxicol Rep. 2024 Dec;13:101806. doi: 10.1016/j.toxrep.2024.101806, PMID 39624223.
Bihari S, Bannard Smith J, Bellomo R. Albumin as a drug: its biological effects beyond volume expansion. Crit Care Resusc. 2020 Sep;22(3):257-65. doi: 10.1016/S1441-2772(23)00394-0, PMID 32900333.
Damborska A, Hanakova L, Pindurova E, Horska K. Case report: therapeutic drug monitoring and CYP2D6 phenoconversion in a protracted paroxetine intoxication. Front Pharmacol. 2024;15:1444857. doi: 10.3389/fphar.2024.1444857, PMID 39295933.
Zhou Z, Chen C, Sun M, Xu X, Liu Y, Liu Q. A decision tree model to predict liver cirrhosis in hepatocellular carcinoma patients: a retrospective study. Peer J. 2023;11:e15950. doi: 10.7717/peerj.15950, PMID 37641600.
Ouyang X, Fan Q, Ling G, Shi Y, Hu F. Identification of diagnostic biomarkers and subtypes of liver hepatocellular carcinoma by multi-omics data analysis. Genes (Basel). 2020 Sep;11(9):1051. doi: 10.3390/genes11091051, PMID 32899915.
Kumar S, Sinha N, Gerth KA, Rahman MA, Yallapu MM, Midde NM. Specific packaging and circulation of cytochromes P450, especially 2E1 isozyme in human plasma exosomes and their implications in cellular communications. Biochem Biophys Res Commun. ScienceDirect. 2017;491(3):675-80. doi: 10.1016/j.bbrc.2017.07.145, PMID 28756226.
Kumar S, Singla B, Singh AK, Thomas Gooch SM, Zhi K, Singh UP. Hepatic extrahepatic and extracellular vesicle cytochrome P450 2E1 in alcohol and acetaminophen-mediated adverse interactions and potential treatment options. Cells. 2022;11(17):2620. doi: 10.3390/cells11172620, PMID 36078027.