
Int J App Pharm, Vol 18, Issue 2, 2026, 464-471Original Article
EVALUATING THE ANTI-INFLAMMATORY POTENTIAL OF GAMMA ORYZANOL FROM ORYZA SATIVA BY TARGETING THE NF-KB PATHWAY: A MOLECULAR DOCKING AND STRUCTURE-BASED PHARMACOPHORE MODELING APPROACH
AASIA KANWAL*, MUHAMMAD HAMDI MAHMOOD , SAIFUL BAHRI TALIP, NORHIDA RAMLI
, SAIFUL BAHRI TALIP, NORHIDA RAMLI
Faculty of Medicine and Health Sciences, Universiti Malaysia Sarawak-94300 Kota Samarahan, Malaysia
*Corresponding author: Aasia Kanwal; *Email: 24010213@siswa.unimas.my
Received: 20 Nov 2025, Revised and Accepted: 07 Feb 2026
ABSTRACT
Objective: The nuclear factor kappa-B (NF-κB) pathway is a key regulator of inflammation observed in polycystic ovary syndrome (PCOS), rendering it a promising target for treatment. Gamma oryzanol (γ-oryzanol) has been reported to display anti-inflammatory properties. However, the particular γ-oryzanol compounds and their specific molecular mechanism by which γ-oryzanol interacts and modulates NF-κB activity have yet to be explained. The study intended to explore the molecular interactions underlying the anti-inflammatory activity of γ-oryzanol against NF-κB through in silico molecular docking and structure-based pharmacophore modeling.
Methods: A receptor-based pharmacophore model was created from the ligand-binding site of the NF-κB through the Molecular Operating Environment 2019 software. The pharmacophore comprised four features: one hydrogen bond donor, one hydrogen bond acceptor, one aromatic, and one hydrophobic feature. The optimized model was used to screen an in-house phytochemical database to find the hit compounds with matching features, followed by molecular docking of hit compounds to evaluate their binding manners and interactions with NF-κB. The docking poses were analyzed for key interactions and ranked based on their docking scores.
Results: Four lead compounds that satisfied the pharmacophore query were 24-methylenecycloartenyl ferulate, cycloartenyl ferulate, campesteryl ferulate, and β-sitosteryl ferulate. The docking results showed that 24-methylenecycloartenyl ferulate had the most potent interaction with NF-kB (-6.9 kcal/mol), followed by cycloartenyl ferulate (-6.7 kcal/mol), campesteryl ferulate (-5.9 kcal/mol), and β-sitosteryl ferulate (-5.1 kcal/mol), indicating their potential to modulate NF-κB.
Conclusion: The present study provides molecular insights into the potential modulatory mechanism of γ-oryzanol against NF-κB. γ-oryzanol, along with structurally related phytochemicals, may serve as a promising scaffold for targeting NF–κB–mediated inflammation, implicated in PCOS. These computational predictions offer a foundation for experimental validation in related inflammatory disease models.
Keywords: Gamma oryzanol, Nuclear factor kappa-B, Polycystic ovary syndrome, Infertility, Pharmacophore modeling
© 2026 The Authors. Published by Innovare Academic Sciences Pvt Ltd. This is an open access article under the CC BY license (https://creativecommons.org/licenses/by/4.0/)
DOI: https://dx.doi.org/10.22159/ijap.2026v18i2.57648 Journal homepage: https://innovareacademics.in/journals/index.php/ijap
INTRODUCTION
Polycystic ovary syndrome (PCOS) is a common endocrine condition observed in women during their reproductive years, marked by disturbances in hormonal balance, abnormalities in metabolic function, structural changes within the ovaries, and persistent low-grade inflammatory activity [1]. The nuclear factor kappa-B (NF-κB) pathway mediates this inflammation, disrupting the reproductive function and resulting in infertility [2]. At present, PCOS treatment remains largely symptom-oriented, highlighting the need for a more systemic and comprehensive therapeutic approach [3]. Emerging PCOS therapies are directed to combat the persistent inflammation, which is now recognized as a central contributor to the disorder. Studies have shown that anti-inflammatory compounds may help correct the major pathological components of PCOS, and therapeutic strategies directed toward NF-κB hold considerable promise for alleviating inflammaging and mitigating the progression of inflammation-driven disorders like PCOS [4, 5]. For this purpose, there is a growing interest in plant-based treatments for PCOS due to their rich bioactive profiles, which influence various biological targets and exhibit comparatively minimal side effects [6, 7]. Among these natural agents, gamma oryzanol (γ-oryzanol) is currently being evaluated for its ability to alleviate various reproductive abnormalities associated with PCOS [8]. It is a bioactive phytosteryl ferulate complex derived from Oryza sativa [9], and has been reported to influence steroid hormone synthesis, exhibit anti-estrogenic, antioxidant, and anti-inflammatory effects [10, 11]. γ-oryzanol exerts its anti-inflammatory effects by attenuating the lipopolysaccharide-induced NF-κB activation. It interferes with the TLR4-mediated signaling pathway and stabilizes IκBα. This stabilization prevents the nuclear translocation of p65, thereby suppressing the transcription of NF-κB–dependent pro-inflammatory cytokines [12, 13]. This pathway modulation makes γ-oryzanol a mechanistically relevant candidate for evaluation as a structure-based NF-κB inhibitor in the present study. This study aimed to investigate the molecular interactions underlying the anti-inflammatory activity of γ-oryzanol against NF-κB through in silico molecular docking and structure-based pharmacophore modeling.
MATERIALS AND METHODS
Pharmacophore study
The 3-dimensional structures of γ-oryzanol compounds, such as 24-methylenecycloartenyl ferulate (24-MCAF), cycloartenyl ferulate (CAF), campesteryl ferulate, and β-sitosteryl ferulate, were downloaded from the PubChem repository and saved as SDF files. The NF-κB protein and ligand files were imported into the Molecular Operating Environment (MOE) 2019 software by Chemical Computing Group for pharmacophore visualization. A receptor structure-based pharmacophore model was created using the pharmacophore query editor tool of MOE. The query editor tool identified key pharmacophore features, such as hydrogen bond acceptors (HBA), hydrogen bond donors (HBD), aromatic (ARO), and hydrophobic (HP) features, from the 3-dimensional structure of NF-κB. The pharmacophore model was validated to ensure its ability to discriminate between known actives and the decoys. 15 known NF-κB inhibitors with documented experimental activity were selected, and 735 decoys were generated through Database of Useful Decoys (DUD. E) for model validation. A database was created in MOE version 2019, containing both actives and decoys. This database was screened against the generated pharmacophore model. Total hits (Ht), active hits (Ha), sensitivity, specificity, enrichment factor (EF), and Güner–Henry score (GH) were calculated as shown in table 1. A GH score ≥0.7 indicates a good pharmacophore model.
Table 1: Pharmacophore model validation
| S. No. | Validation parameters | Validation results |
| Total number of compounds in database (N) | 750 | |
| Total number of actives in database (A) | 15 | |
| Total hits | 22 | |
| Active hits | 14 | |
| Sensitivity | 0.93 | |
| Specificity | 0.98 | |
| Enrichment factor | 31.82 | |
| Güner–Henry score | 0.70 |
After validating the pharmacophore, an in-house database of γ-oryzanol compounds was created in mdb. format. This database represents a targeted set of eight gamma-oryzanol compounds rather than a large virtual library. The optimized pharmacophore was used as a query to virtually screen the in-house database using MOE's pharmacophore search algorithm. The threshold for hit identification was based on rscore of>2, RMSD<1 Å, and successful mapping of ligands to at least 4 pharmacophore features [14]. The compounds that matched the pharmacophoric features were selected as hit compounds and subjected to molecular docking with NF-κB to assess their binding affinities and interactions. The workflow for receptor structure preparation, pharmacophore generation, and molecular docking is shown in fig. 1.
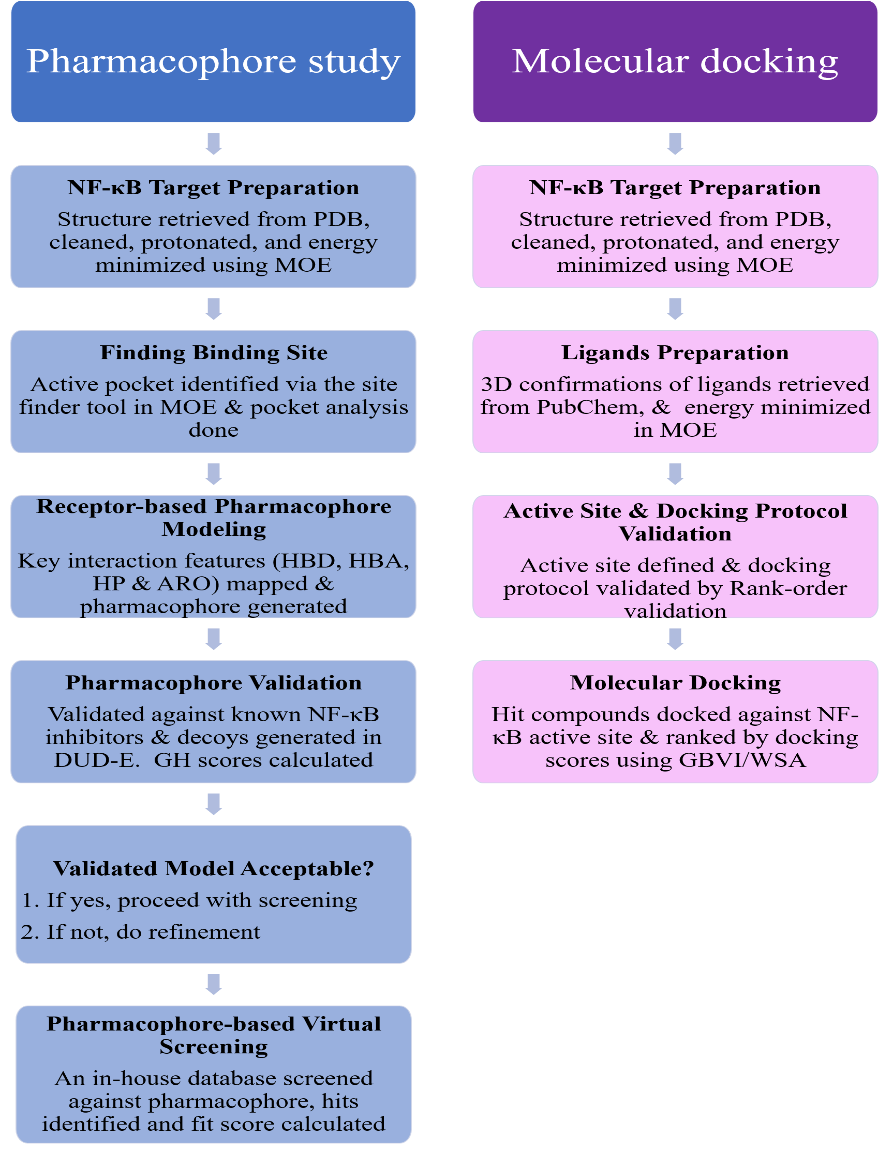
Fig. 1: Workflow of receptor structure preparation, pharmacophore generation, and molecular docking
Preparation of the ligands, target receptor, and identification of the active site
A 3-dimensional structure of NF-κB was downloaded from the Protein Data Bank (PDB ID: 1IKN) in PDB format. The selected protein is a NF-κB p65/p50 heterodimer with a resolution of 2.30 Å. The structure reflects a regulatory, inhibitor-bound state, which preserves the native fold and interdomain architecture of the p65/p50 heterodimer. The choice of this structure was based on its high crystallographic quality, well-resolved heterodimeric interface, and precedence in the literature [15]. The structure was imported into the Molecular Operating Environment (MOE) software and prepared. The duplicate and extra chains were removed. All the ligands and water molecules were removed, hydrogens were added, and energy minimization was done. The active binding site on the prepared NF-κB was identified using the MOE site-finder tool by identifying the key residues on chains of interest.
The three-dimensional structure of the ligands was retrieved from the PubChem database repository, downloaded, and saved in 3-D SDF format. The energy minimization was done for ligands by importing them to the MOE software.
Molecular docking
Molecular docking was performed using the “dock” feature of MOE with default settings. For the placement of the ligand, the triangular matcher method was used to generate ten poses for each ligand, and scored by London dG. For the refinement purpose, the induced fit refinement method was used; five poses were generated and scored by GBVI/WSA dG. For each ligand, the pose with the most negative (lowest) GBVI/WSA dG score was selected. As the NF-κB structure does not have a native/co-crystalized ligand, the RMSD-based redocking validation could not be performed. An alternative rank-order docking validation strategy was adopted [16]. Ten known NF-κB inhibitors with reported experimental activities (IC50) were selected. The IC50 values were converted into negative log (pIC50) values. These inhibitors were docked to the active site of NF-κB, using the same docking protocol, and binding scores were calculated. Their predicted binding scores were compared with the reported experimental activities, and rank-ordered to evaluate whether the docking protocol correctly reproduced their relative potencies. Spearman’s correlation coefficient (ρ) was calculated to quantitatively assess the predicted and experimental rankings. A Spearman’s “ρ”>0.7 indicates acceptable docking protocol validity. Following the docking validation, the γ-oryzanol compounds were docked to the NF-κB active site, using the same docking protocol, and binding scores were calculated. The docking poses were analyzed for key interactions and were ranked based on their GBVI/WSA dG docking scores.
RESULTS
The pharmacophore search identified four hit compounds: 24-methylenecycloartenyl ferulate, cycloartenyl ferulate, campesteryl ferulate, and β-sitosteryl ferulate, as shown in table 2.
Table 2: PubChem CIDs, compound names and chemical structures of hit compounds
| PubChem CIDs | Compounds name | 2-D Structure of compounds |
| CID 9920169 | 24-methylenecycloartenyl ferulate | 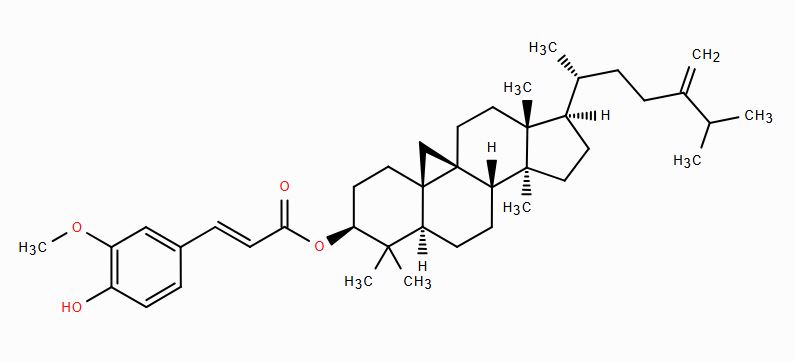 |
| CID 5282164 | Cycloartenyl ferulate | 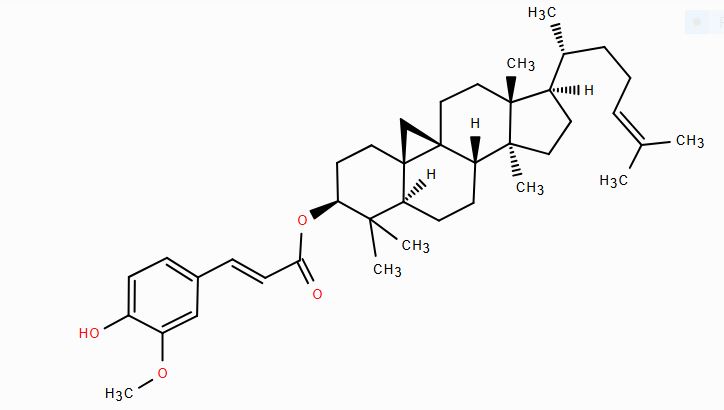 |
| CID 15056832 | Campesteryl ferulate | 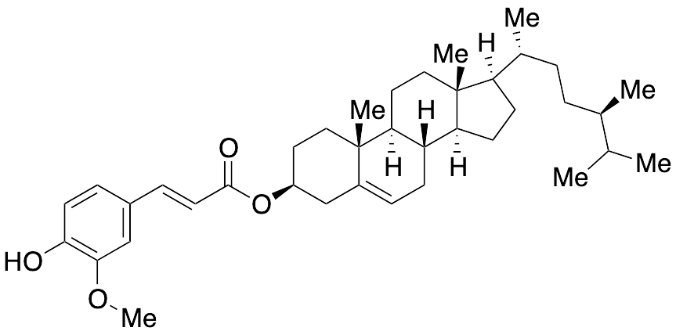 |
| CID 9938436 | β-sitosteryl ferulate | 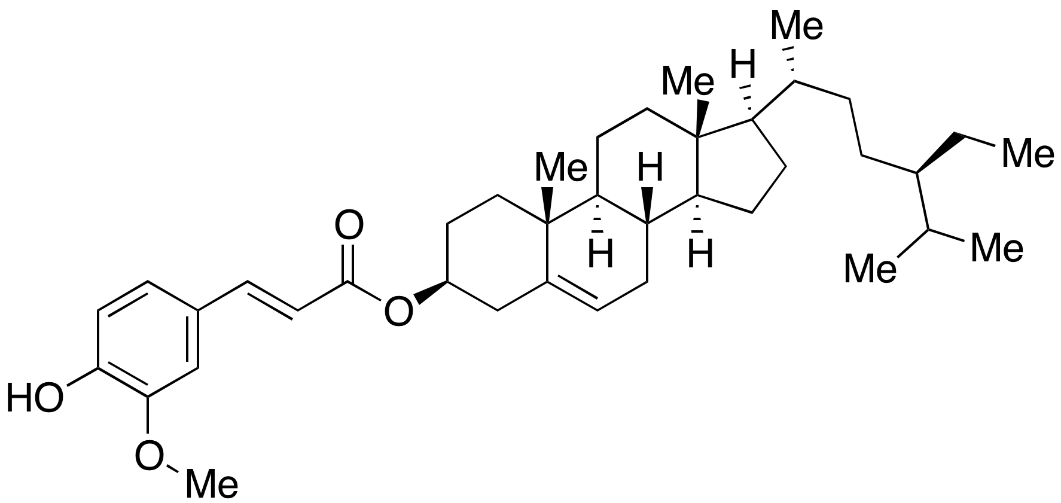 |
The pharmacophore query model generated from the crystal structure of NF-κB comprised four features: one HBD, one HBA, one ARO, and one hydrophobic HP. All γ-oryzanol compounds exhibited 3-4 common pharmacophore features. The 24-methylenecycloartenyl ferulate exhibited one HBD/HBA, two HBA, six HP, and one ARO features. The cycloartenyl ferulate had one HBD, two HBA, five HP, and one ARO. Campesteryl ferulate exhibited one HBD, two HBA, five HP, and one ARO features, while β-sitosteryl ferulate has one HBD, two HBA, eight HP, and one ARO, as shown in fig. 2.
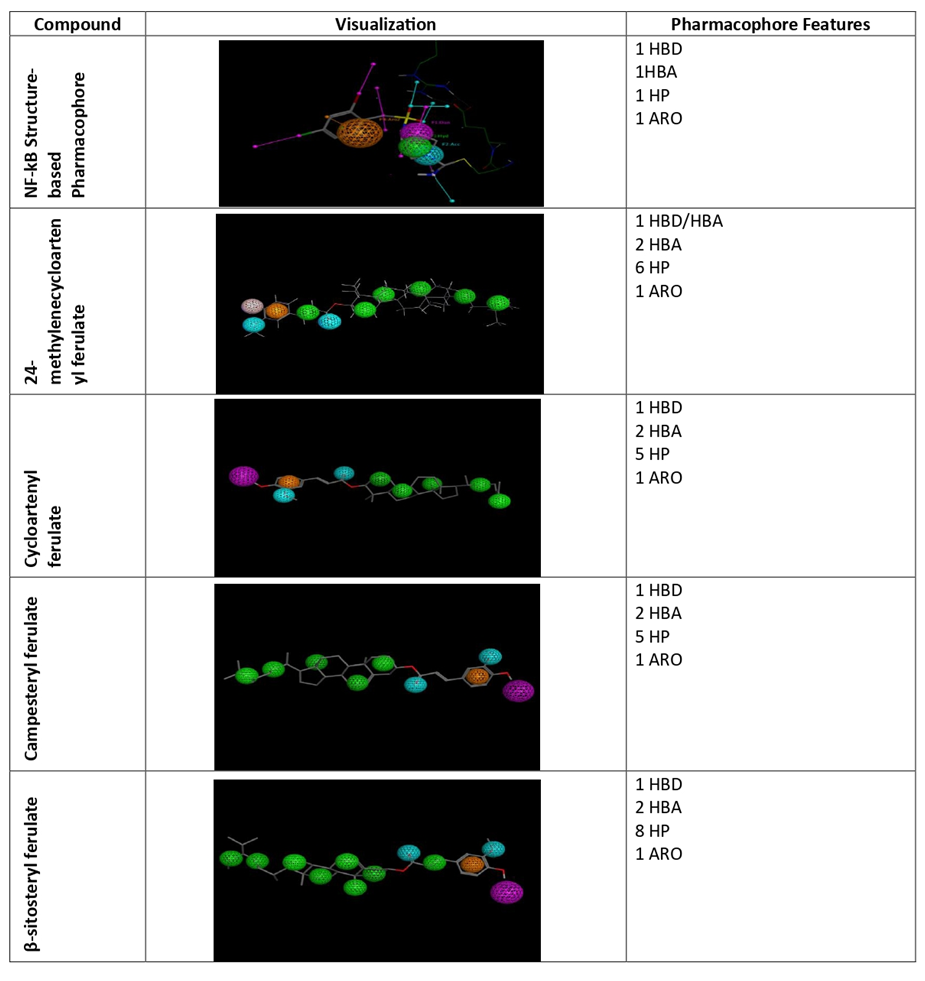
Fig. 2: Receptor structure-based pharmacophore of NF-kB (green: hydrophobic; purple: hydrogen donor; cyan: hydrogen acceptor; orange: aromatic) and pharmacophore features mapping of gamma oryzanol compounds, four γ-oryzanol compounds mapped successfully on the pharmacophore model as shown in fig.3.
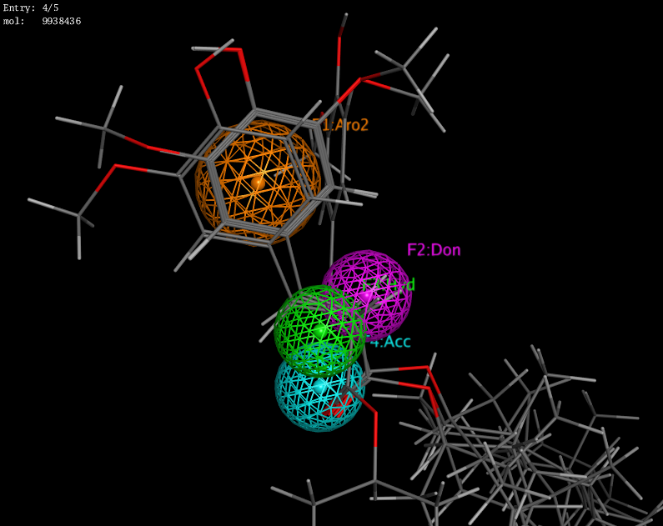
Fig. 3: The pharmacophore mapping of all four compounds on the model, γ-oryzanol compounds i.e. 24-methylenecycloartenyl ferulate, cycloartenyl ferulate, campesteryl ferulate, and β-sitosteryl ferulate exhibited fit score >2 and RMSD <1 as given in Table 3.
Table 3: Scores of pharmacophore screening
| Compounds PubChem CIDs | Fit RMSD | Pharmacophore fit score |
| CID 9920169 | 0.6189 | 3.53 |
| CID 5282164 | 0.6236 | 3.53 |
| CID 15056832 | 0.6194 | 3.53 |
| CID 9938436 | 0.6179 | 3.53 |
Fit RMSD: Fit root mean square deviation refers to the root mean square deviation between the mapped ligand features and the query features after alignment; lower Fit RMSD values indicate better spatial matching.
The rank-order validation of the docking protocol demonstrates that the predicted binding affinities (GBVI/WSA dG) accurately reflected the experimentally reported actual half maximal inhibitory concentration of known NF-κB inhibitors, as shown in table 4, yielding a Spearman’s correlation coefficient of −0.8 and a p-value of 0.01.
Table 4: Rank order docking validation
| Experimental potency rank | pIC50 | Docking score (kcal/mol) |
| 1 | 7.92 | -10.7 |
| 2 | 7.51 | -6.6 |
| 3 | 7.12 | -8.4 |
| 4 | 7.06 | -8.3 |
| 5 | 6.70 | -8.9 |
| 6 | 6.0 | -7.1 |
| 7 | 5.62 | -6.1 |
| 8 | 5.15 | -5.9 |
| 9 | 5.0 | -5.7 |
| 10 | 4.82 | -5.3 |
Spearman’s rank correlation coefficient (ρ) between experimental pIC₅₀ values and predicted docking scores was −0.8 (p = 0.01), indicating a strong and statistically significant inverse correlation; kcal/mol: kilocalorie per mole; pIC₅₀ is the negative base-10 logarithm of the half-maximal inhibitory concentration (IC₅₀) expressed in molar units.
The hit compounds, 24-methylenecycloartenyl ferulate, cycloartenyl ferulate, campesteryl ferulate, and β-sitosteryl ferulate, underwent molecular docking with NF-κB. Among all ligands analyzed, 24-methylenecycloartenyl ferulate had the most potent interaction with NF-κB (-6.9) followed by cycloartenyl ferulate (-6.7), campesteryl ferulate (-5.9), and β-sitosteryl ferulate (-5.1) as shown in fig. 4.
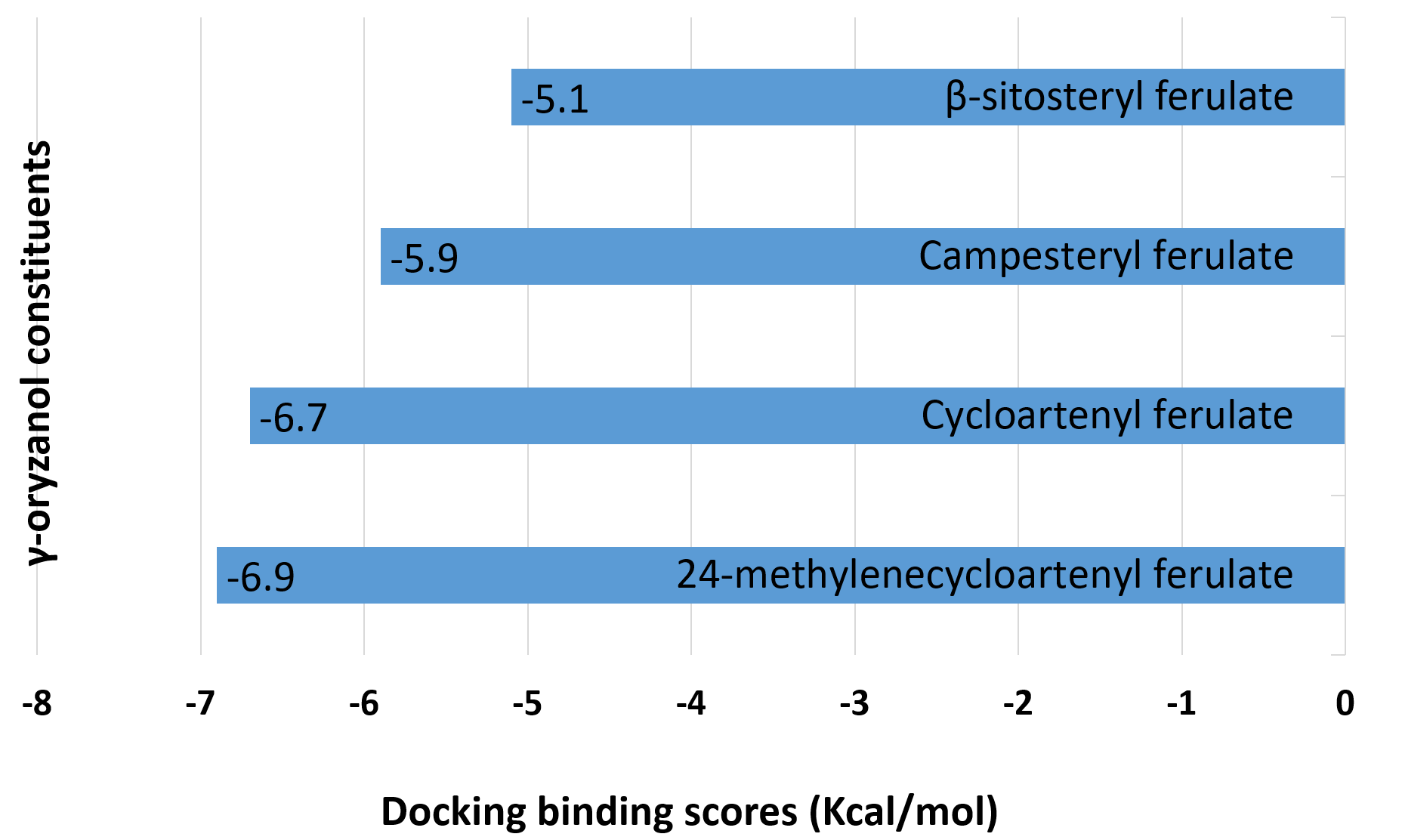

Fig. 4: Molecular docking binding scores (kilocalorie per mole) of gamma oryzanol constituents with NF-κB
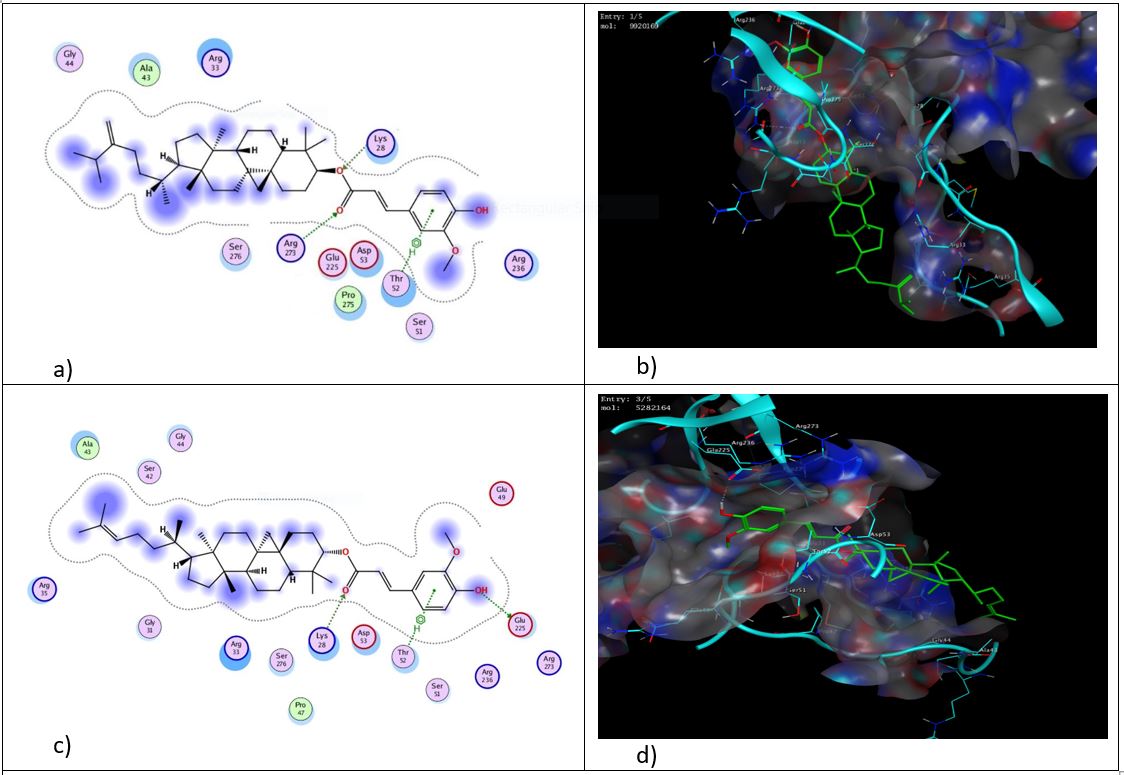
Fig. 5: 2D and 3D representation of binding interactions of 24-methylenecycloartenyl ferulate (a and b) and cycloartenyl ferulate (c and d) with the active site of NF-κB (1IKN)
As shown in fig. 5, among various ligands analyzed, 24-methylenecycloartenyl ferulate had interaction with NF-κB through two hydrogen bonds with key residues Arg 273 and Lys 28 and π-cation interaction with Thr 52, followed by cycloartenyl ferulate through two hydrogen bonds with Lys 28, and Glu 225 and π-cation interaction with Thr 52. Campesteryl ferulate formed two hydrogen bonds with Lys 28 and Gln 241, and β-sitosteryl ferulate interacted by forming two hydrogen bonds with Gly 259 and Arg 236 (Fig.6). Other than the identified hydrogen bonds and π-cation interactions, a major component of the binding affinity of γ-oryzanol compounds results from substantial hydrophobic and van der Waals interactions between the bulky sterol–ferulate backbone and the NF-κB binding pocket. The binding cavity of the Rel homology domain is largely hydrophobic and facilitates the sequestration of large lipophilic ligands. The rigid sterol cores of all γ-oryzanol compounds are deeply buried within this cavity, creating a huge interaction surface and shape complementarity. Whereas hydrogen bonds anchor the ferulate moiety to orient the ligand, cumulative van der Waals forces and hydrophobic enclosure contribute to interaction stability. This effect is particularly evident for 24-methylenecycloartenyl ferulate, whose slightly extended sterol framework enhances steric complementarity and hydrophobic packing within the pocket, providing a possible structural explanation for its more favorable docking score compared with the other compounds.
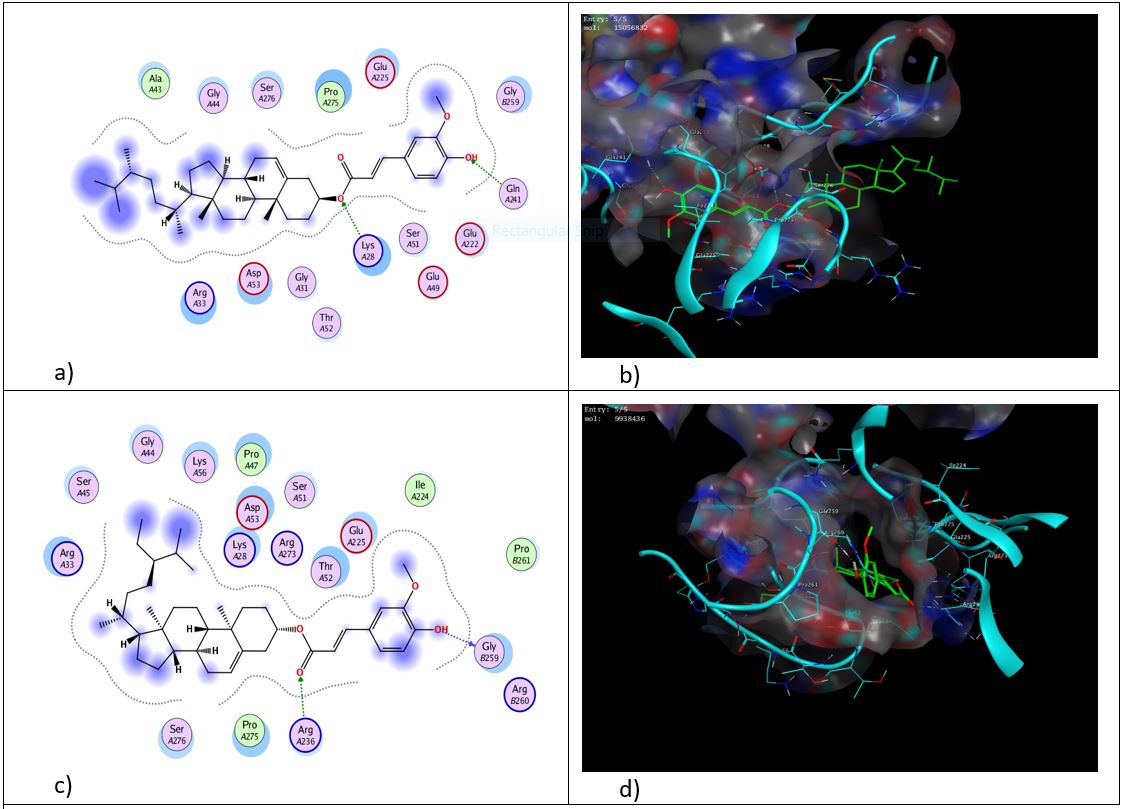
Fig. 6: 2D and 3D representation of binding interactions of campesteryl ferulate (a and b) and β-sitosteryl ferulate (c and d) with the active site of NF-Κb (1IKN)
DISCUSSION
The routine drug development process is resource-intensive and economically burdensome. This has driven the emergence of economical and efficient computer-based approaches for the rapid production and screening of therapeutic compounds for clinical use [17]. In recent years, advances in in silico research have opened new avenues for exploring the phytochemicals and drug discovery [18-21]. The integration of pharmacological therapies with natural products has demonstrated promising potential in the treatment of various diseases [22]. Despite this promising potential, many plant-derived compounds remain unavailable in the commercial markets due to challenges in large-scale production [23, 24]. On the other hand, γ-oryzanol has already been optimized through reliable extraction and purification techniques, allowing its large-scale production as an accessible nutritional and health-promoting supplement [25]. Inflammatory disorders are central research concerns at the moment due to their significant implications for global health [26]. NF-κB primarily controls inflammation and becomes activated upon the secretion of proinflammatory cytokines. Thus, inhibiting NF-κB may be a possible approach to regulating inflammatory cytokines [27, 28]. Contrary to standard therapeutic agents, natural compounds from γ-oryzanol may be promising agents to block and halt NF-κB from producing pro-inflammatory mediators with fewer side effects [29].
The γ-oryzanol compounds, such as 24-MCAF and CAF, hold versatile therapeutic potential that contributes to strengthening the immune system and managing inflammation-related disorders [30]. Different in vivo experimental studies have revealed that γ-oryzanol compounds, predominantly 24-MCAF and CAF, have huge immunoregulatory and antioxidant potential [31, 32].
From the pharmacophore point of view, 24-MCAF and CAF were a better fit to the features, namely one HBD, two HBA, one ARO, and six HP points. These pharmacophoric attributes are more than adequate to meet the pharmacophoric requirements necessary for inhibiting NF-κB. NF-κB pharmacophoric study, conducted by Tsai KC. et al. for screening potent NF-κB inhibitors showed five pharmacophoric characteristics, namely three HP, one HBA, and one ARO point [33]. This pharmacophore demonstrates a robust ability to recognize NF-κB inhibitors and to predict the activities of molecules with structurally diverse profiles. In this regard, γ-oryzanol compounds have shown compatibility with the NF-κB pharmacophore, featuring aromatic and three additional HP and one additional HBA features. The binding modes of the top-scoring γ-oryzanol compounds display a high structural correspondence with established NF-κB inhibitors, particularly in targeting the p65 Rel homology domain. Our findings demonstrate the involvement of Lys 28, Arg 273, and Glu 225, which are critical residues for stabilizing the NF-κB-DNA complex and are involved in its modulation [34, 35]. Interaction with Lys 28 is a hallmark of potent p65 inhibition and particularly mirrors the inhibitory mechanism of quercetin and curcumin, well known to target the p65 subunit and modulate its stability [36,37]. Additionally, the interaction of γ-oryzanol compounds with Arg 273 and Glu 225 aligns with the binding modes of caffeic acid phenethyl ester and andrographolide, which anchor within the binding cavity of the NF-κB to mask these residues and hinder NF-κB interaction and activation. This multi-residue binding at the Rel homology domain matches the interaction mode of curcumin, which nestles within this binding cavity to suppress the transcriptional activity of p65 [38,39]. Combining in silico pharmacophore and docking-based modelling approaches provides significant advantages [40]. This combined approach can overcome some of the limitations inherent to each method, leading to more reliable outcomes [41, 42]. The docking results showed that the 24-MCAF and CAF displayed the strongest binding score (-6.9 kcal/mol) and (-6.7 kcal/mol) as compared to other compounds. In this scenario, 24-MCAF and CAF are anticipated to exhibit a more pronounced anti-inflammatory response against NF-κB due to their favorable binding with key residues. Although 24-MCAF and CAF share the same ferulate group and matching pharmacophore features, 24-MCAF has a 24-methylene group, not present in CAF. This slight difference in structures alters their spatial orientation, binding preference, and yields slightly different docking scores [43].
CONCLUSION
The present study provides molecular insights into the potential modulatory mechanism of γ-oryzanol against NF-κB. It defines the key binding features and specific amino acid residues required for γ-oryzanol to interact with NF-κB, structurally justifying γ-oryzanol as an NF-κB modulator, and generates a validated pharmacophore model. This pharmacophore blueprint may be used for virtual screening and identification of NF-κB inhibitors. γ-oryzanol, along with structurally related phytochemicals, may serve as a promising scaffold for targeting NF–κB–mediated inflammation, implicated in PCOS. These computational predictions offer a foundation for experimental validation in related inflammatory disease models.
FUNDING
Nil
AUTHORS CONTRIBUTIONS
AK: Conceptualization, Data curation, Formal analysis, Investigation, Methodology, Resources, Software, Validation, Visualization, Writing-original draft, review and editing; MHM: Conceptualization, Data curation, Validation, Visualization; SBT: Visualization, Review and editing; NR: Visualization; Review and editing.
CONFLICT OF INTERESTS
Declared none
REFERENCES
Siddiqua A, Malik A, Iqbal U, Sial NT, Mehmood MH, Rehman MF. Antioxidative and anti-inflammatory effects of carvacrol against polycystic ovary syndrome-associated complications using high fat diet and letrozole challenged rat model: a multidisciplinary study cascading in vivo, in vitro, in silico and network pharmacology approaches. Inflammopharmacology. 2025 Jun 20;33(8):4703-24. doi: 10.1007/s10787-025-01809-8, PMID 40540113.
Hu M, Zhang Y, Li X, Cui P, Sferruzzi Perri AN, Brannstrom M. TLR4-associated IRF-7 and NFκB signaling act as a molecular link between androgen and metformin activities and cytokine synthesis in the PCOS endometrium. J Clin Endocrinol Metab. 2021 Apr 1;106(4):1022-40. doi: 10.1210/clinem/dgaa951, PMID 33382900.
Pachiappan S, Arul Balasubramanian MG, Ramalingam K. Pharmacoinformatics-based in silico molecular dynamics simulation for screening phytochemicals as AMPK and INSR modulators for polycystic ovarian syndrome from medicinal plants. International Journal of Pharmaceutical Sciences and Research. 2023;15:1087-92.
Rostamtabar M, Esmaeilzadeh S, Tourani M, Rahmani A, Baee M, Shirafkan F. Pathophysiological roles of chronic low-grade inflammation mediators in polycystic ovary syndrome. J Cell Physiol. 2021;236(2):824-38. doi: 10.1002/jcp.29912, PMID 32617971.
Tan W, Zhang J, Dai F, Yang D, Gu R, Tang L. Insights on the NF-κB system in polycystic ovary syndrome: attractive therapeutic targets. Mol Cell Biochem. 2024;479(3):467-86. doi: 10.1007/s11010-023-04736-w, PMID 37097332.
Lakshmi JN, Babu AN, Kiran SS, Nori LP, Hassan N, Ashames A. Herbs as a source for the treatment of polycystic ovarian syndrome: a systematic review. BioTech (Basel). 2023;12(1):4. doi: 10.3390/biotech12010004, PMID 36648830.
Pavithra L, Ilango K. Identification of phytoconstituents for combating polycystic ovarian syndrome through in silico techniques. Indian J Biochem Biophys. 2023 Jun 27;60(2):99-107. doi: 10.56042/ijbb.v60i2.70682.
Chakraborty S, Khurana N, Kaur J, Mehta M, Sharma N. Gamma oryzanol: a natural compound with potential for treating polycystic ovary syndrome. Pharmacological Research Modern Chinese Medicine. 2024;13:100506. doi: 10.1016/j.prmcm.2024.100506.
Filho AC, Guedes MI, Duarte LS, Lima Neto AB, Cameron LC, Bassini A. Gamma-oryzanol has an equivalent efficacy as a lipid-lowering agent compared to fibrate and statin in two dyslipidemia mice models. Int J Pharm Pharm Sci. 2014;6(11):61-4.
Alizadeh M, Moshtagh S, Abdolalizadeh Amir S, Jeddi M, Tahmasebzadeh S, Radman G. Anti-oxidant and anti-apoptotic effects of gamma-oryzanol on male reproductive function in chronic restraint stress in rats. Avicenna J Phytomed. 2025;15(1):890-906. doi: 10.22038/AJP.2024.24819, PMID 40271500.
Radda MI, Omar N, Yusof SF, Ahmad R, Rohana AJ, Ishak WR. Effectiveness of gamma-oryzanol in glycaemic control and managing oxidative stress, inflammation and dyslipidaemia in diabetes: a systematic review of preclinical studies. Peer J. 2025 Sep 23;13:e20062. doi: 10.7717/peerj.20062, PMID 41018904.
Panyathep A, Punturee K, Chewonarin T. Gamma-oryzanol-rich fraction from purple rice extract attenuates lipopolysaccharide-stimulated inflammatory responses, migration and VEGFA production in SW480 cells via modulation of TLR4 and NF-κB pathways. Nutr Cancer. 2022 Jul 3;74(6):2254-64. doi: 10.1080/01635581.2021.2002921, PMID 34766845.
Chauhan A, Kamal R, Kumar M, Singh TG, Vishwas S, Awasthi A. Exploring the mechanisms and therapeutic role of γ-oryzanol in neuropathic pain: a systematic review. Inflammopharmacology. 2025 Dec 24:1-29. doi: 10.1007/s10787-025-02100-6, PMID 41442076.
Pratap Reddy Gajulapalli V, Lee J, Sohn I. Ligand-based pharmacophore modelling in search of novel anaplastic lymphoma kinase inhibitors. Results Chem. 2023;5:100752. doi: 10.1016/j.rechem.2022.100752.
Lin CT, Lin LW, Chang LC, Wu LY, Tsai FS. Combining network pharmacology, molecular docking and integrative studies to explore the mechanism of helminthostachys zeylanica in alleviating ulcerative colitis. Food Sci Nutr. 2025 Nov;13(11):e71139. doi: 10.1002/fsn3.71139, PMID 41164267.
Martis EA, Teletchea S. Ten quick tips to perform meaningful and reproducible molecular docking calculations. PLOS Comput Biol. 2025 May 9;21(5):e1013030. doi: 10.1371/journal.pcbi.1013030, PMID 40344147.
Muhammed MT, Aki-Yalcin E. Pharmacophore modeling in drug discovery: methodology and current status. Journal of the Turkish Chemical Society Section A. 2021;8(3):749-62. doi: 10.18596/jotcsa.927426.
Dayrit FM, Guidote AM, Gloriani NG, De Paz-Silava SL, Villasenor IM, Macahig RA. Philippine medicinal plants with potential immunomodulatory and anti-SARS-CoV-2 activities. Philipp J Sci. 2021;150(5):999-1015. doi: 10.56899/150.05.12.
Wu C, Liu Y, Yang Y, Zhang P, Zhong W, Wang Y. Analysis of therapeutic targets for SARS-CoV-2 and discovery of potential drugs by computational methods. Acta Pharm Sin B. 2020;10(5):766-88. doi: 10.1016/j.apsb.2020.02.008, PMID 32292689.
Elfiky AA. Natural products may interfere with SARS-CoV-2 attachment to the host cell. J Biomol Struct Dyn. 2021;39(9):3194-203. doi: 10.1080/07391102.2020.1761881, PMID 32340551.
Kori MA, Panera BH, Buwa VA. Molecular docking of phytochemicals as inhibitors of EGFR in non-small cell lung cancer. Innov J Med Sci. 2022;10(4):28-30. doi: 10.22159/ijms.2022v10i4.45085.
Gentile D, Patamia V, Scala A, Sciortino MT, Piperno A, Rescifina A. Putative inhibitors of SARS-COV-2 main protease from a library of marine natural products: a virtual screening and molecular modeling study. Mar Drugs. 2020;18(4):225. doi: 10.3390/md18040225, PMID 32340389.
Marchev AS, Yordanova ZP, Georgiev MIG. Green (cell) factories for advanced production of plant secondary metabolites. Crit Rev Biotechnol. 2020;40(4):443-58. doi: 10.1080/07388551.2020.1731414, PMID 32178548.
Han T, Miao G. Strategies, achievements and potential challenges of plant and microbial chassis in the biosynthesis of plant secondary metabolites. Molecules. 2024 May 2;29(9):2106. doi: 10.3390/molecules29092106, PMID 38731602.
Baixinho JP, Cardeira M, Bento Silva A, Partidario AM, Serra AT, Bronze MD. Optimization of supercritical fluid extraction for the recovery of γ-oryzanol-rich extracts with improved bioactivity from rice bran. Antioxidants (Basel). 2025 Feb 11;14(2):206. doi: 10.3390/antiox14020206, PMID 40002392.
Fioranelli M, Roccia MG, Flavin D, Cota L. Regulation of inflammatory reaction in health and disease. Int J Mol Sci. 2021 May 17;22(10):5277. doi: 10.3390/ijms22105277, PMID 34067872.
Kannan G, Paul BM, Thangaraj P. Stimulation regulation and inflammaging interventions of natural compounds on nuclear factor kappa B (NF-kB) pathway: a comprehensive review. Inflammopharmacology. 2025 Jan;33(1):145-62. doi: 10.1007/s10787-024-01635-4, PMID 39776026.
Mao H, Zhao X, Sun SC. NF-κB in inflammation and cancer. Cell Mol Immunol. 2025;22(8):811-39. doi: 10.1038/s41423-025-01310-w, PMID 40562870.
Nikooyeh B, Zargaraan A, Ebrahimof S, Kalayi A, Zahedirad M, Yazdani H. Added γ-oryzanol boosted anti-inflammatory effects of canola oil in adult subjects with type 2 diabetes: a randomized controlled clinical trial. Eur J Nutr. 2024 Mar;63(2):425-33. doi: 10.1007/s00394-023-03275-w, PMID 37971692.
Sari Aslani K, Davoodian N, Zeini S, Mousavi SA, Eftekhar E. γ-oryzanol ameliorates the oxidative stress and inflammatory response in a mice model of LPS-induced liver injury. Dis Diagn. 2024 Sep 5;13(3):107-13. doi: 10.34172/ddj.1585.
Ali MA, Chew SC, Lin Nyam K, Aryusuk K. Green extraction technologies antioxidant mechanism and therapeutic potentials of gamma-oryzanol. Food Rev Int. 2025 Jun 20:1-37. doi: 10.1080/87559129.2025.2521362.
Lisnawati L, Poeranto S, Endharti AT, Santoso MI. Antioxidant and anti-inflammatory activity of γ-oryzanol compared to rice bran oil to repair ovarian histological structure from one push transfluthrin exposure effect. Open Access Maced J Med Sci. 2022 Jan 1;10(B):1-12. doi: 10.3889/oamjms.2022.7543.
Tsai KC, Teng LW, Shao YM, Chen YC, Lee YC, Li M. The first pharmacophore model for potent NF-κB inhibitors. Bioorg Med Chem Lett. 2009 Oct 1;19(19):5665-9. doi: 10.1016/j.bmcl.2009.08.021, PMID 19726185.
Al Mousa AA, Abouelela ME, Mansour A, Nasr M, Ali YH, Al Ghamidi NS. Wound healing metabolite profiling and in silico studies of Aspergillus terreus. Curr Issues Mol Biol. 2024 Oct 19;46(10):11681-99. doi: 10.3390/cimb46100694, PMID 39451574.
Wang P, Zhang R, Liu M, Gao J, Zhang Y, Liu X. Exploring the effect and mechanism of baicalin on sepsis-induced acute lung injury based on network pharmacology and experimental verification. Sci Rep. 2025 Nov 25;15(1):41959. doi: 10.1038/s41598-025-25943-z, PMID 41290918.
Chen FE, Huang DB, Chen YQ, Ghosh G. Crystal structure of p50/p65 heterodimer of transcription factor NF-κB bound to DNA. Nature. 1998 Jan 22;391(6665):410-3. doi: 10.1038/34956, PMID 9450761.
Natarajan K, Singh S, Burke Jr TR, Grunberger D, Aggarwal BB. Caffeic acid phenethyl ester is a potent and specific inhibitor of activation of nuclear transcription factor NF-kappa B. Proc Natl Acad Sci USA. 1996 Aug 20;93(17):9090-5. doi: 10.1073/pnas.93.17.9090, PMID 8799159.
Xia YF, Ye BQ, Li YD, Wang JG, He XJ, Lin X. Andrographolide attenuates inflammation by inhibition of NF-κB activation through covalent modification of reduced cysteine 62 of p50. J Immunol. 2004 Sep 15;173(6):4207-17. doi: 10.4049/jimmunol.173.6.4207, PMID 15356172.
Gupta SC, Sundaram C, Reuter S, Aggarwal BB. Inhibiting NF-κB activation by small molecules as a therapeutic strategy. Biochim Biophys Acta. 2010;1799(10-12):775-87. doi: 10.1016/j.bbagrm.2010.05.004, PMID 20493977.
Islam A, Jena D, Mondal NS, Teli A, Mondal S, Gautam MK. In silico approaches for drug designing technology: bridging discovery and development. Curr Drug Discov Technol. 2025 Sep;22(5):706-16. doi: 10.2174/0115701638326869250207060616, PMID 39966365.
Vazquez J, Lopez M, Gibert E, Herrero E, Luque FJ. Merging ligand-based and structure-based methods in drug discovery: an overview of combined virtual screening approaches. Molecules. 2020 Oct 15;25(20):4723. doi: 10.3390/molecules25204723, PMID 33076254.
Ancuceanu R, Lascu BE, Draganescu D, Dinu M. In silico ADME methods used in the evaluation of natural products. Pharmaceutics. 2025 Jul 31;17(8):1002. doi: 10.3390/pharmaceutics17081002, PMID 40871023.
Ito J, Sawada K, Ogura Y, Xinyi F, Rahmania H, Mohri T. Definitive evidence of the presence of 24-methylenecycloartanyl ferulate and 24-methylenecycloartanyl caffeate in barley. Sci Rep. 2019 Aug 29;9(1):12572. doi: 10.1038/s41598-019-48985-6, PMID 31467350.