Int J Curr Pharm Res, Vol 18, Issue 1, 116-120Original Article
BACTERIOLOGICAL ISOLATION AND THEIR ANTIBIOGRAM AND MYCOLOGICAL ANALYSIS OF PIGEON DROPPINGS IN A TERTIARY CARE HOSPITAL
SAILAJA M., PURIMITLA USHARANI, SOUPARNIKA PR*
Department of Microbiology, Dr Patnam Mahender ReddyInstitute of Medical sciences, Chevella, Ranga Reddy Dist, Telangana
*Corresponding author: Souparnika PR; *Email: souparnika5.pr@gmail.com
Received: 06 Oct 2025, Revised and Accepted: 26 Nov 2025
ABSTRACT
Objective: Pigeon feces are a major source of infectious pathogens spreading to the surrounding environment in yards and live-bird markets. The aim of this study is to isolate, identify, and characterize the bacterial and fungal species present in pigeon droppings, and to evaluate the antibiotic susceptibility patterns (antibiogram) of the bacterial isolates.
Methods: The present study is a cross-sectional, descriptive, laboratory-based study conducted for a period of eight months from Jan 2025 to Aug 2025 in a Tertiary care teaching hospital, Chevella. 110 pigeon dropping samples were taken using sterile cotton swabs from various locations, including temples, historic places, and public parks. Pigeon droppings were cultured on Nutrient, MacConkey and Blood agar and incubated at 37 °C for 24–48 h to isolate bacteria. Identification was based on colony morphology, Gram staining, and standard biochemical tests. Antimicrobial susceptibility was determined using the Kirby–Bauer disc diffusion method on Mueller–Hinton agar following CLSI guidelines. Fungal isolates were obtained on Sabouraud Dextrose Agar at 25–28 °C and identified by their macroscopic and microscopic features.
Results: In the present study, the maximum bacterial count was identified in 10-4 dilution which ranged from 41 x 105 cfu/ml. The major isolates obtained were CONS 38.45%, Pseudomonas aeruginosa 27.6%, E. coli 16.9%. Among the fungal isolates, Candida species 33.3% followed by Aspergllus niger 29.16% from pigeon droppings. Most of the CONS were resistant to penicillin antibiotic 80% and Pseudomonas isolates were resistant to CXM and COT 77.8% followed by 72.3% to AMC and third-generation cephalosporins. Most of the bacterial isolates obtained in the study were multidrug-resistant isolates.
Conclusion: Overall, pigeon droppings represent a potential public health hazard, contributing to environmental contamination, source of MDR isolates and possible zoonotic transmission and hence preventive measures should be followed to prevent to prevent environmental contamination and Zoonotic transmission.
Keywords: Pigeon droppings, Bacteria, Fungi, MDR, Antibiotics
© 2026 The Authors. Published by Innovare Academic Sciences Pvt Ltd. This is an open access article under the CC BY license (https://creativecommons.org/licenses/by/4.0/)
DOI: https://dx.doi.org/10.22159/ijcpr.2026v18i1.8048 Journal homepage: https://innovareacademics.in/journals/index.php/ijcpr
INTRODUCTION
Humans who have frequent contact with pigeons at home, live bird markets, and on farms are at serious risk of contracting diseases, including Salmonella spp., Campylobacter, Escherichia coli, and Chlamydia, while they are around them. Healthy pigeons are a significant human salmonellosis source [1]. Pigeon feces are a major source of infectious pathogens spreading to the surrounding environment in yards and live-bird markets. Pigeons are a source of E. coli and are very good at spreading the bacteria to animals, birds, and people. It has been proposed that E. coli poses a potential zoonotic risk to humans, causing urinary tract infections [2]. In domestic animal species, and poultry specifically, antimicrobial resistance (AMR) restricts the therapeutic options for treating bacterial illnesses [3]. AMR disease strains may be hosted and spread by birds, endangering human health [4]. With respect to the main bacterial infections, the prevalence of AMR has been increasing [5]. A growing problem worldwide is the presence of multidrug-resistant (MDR) Salmonella spp. in several animal species that are bred for food. Through the food chain, antibiotic-resistant zoonotic agents from an animal host may infect humans [6].
Pigeons are found all throughout the world, in both rural and urban areas, and they have intimate interactions with both people and other animals. Because pigeon feces may carry a variety of microorganisms, they may aid in the spread of infectious diseases in the environment. Strong human pathogens, such strains of diarrheagenic Escherichia coli may be among them, increasing the risk of exposure and infection in humans [7, 8]. Clinically and economically, the development of antibiotic resistance in Gram-negative bacteria poses a serious threat to the treatment of humans and animals. Pseudomonas, Stenotrophomonas, Aeromonas, and Enterobacteriaceae are examples of environmental microbes that are multi-resistant opportunistic pathogens that have been connected to severe illnesses in both humans and animals [8].
Pigeons can act as potential carriers of antimicrobial resistance (AMR) due to their frequent exposure to environments contaminated with antibiotic residues and resistant bacteria, such as urban areas, hospitals, farms, and landfills. These birds may harbor resistant bacteria like Escherichia coli or Salmonella in their digestive systems without showing any symptoms, making them silent reservoirs. Studies have identified resistance genes such as bla_CTX-M and mcr-1 in bacteria isolated from pigeons, indicating their role in the environmental spread of AMR. Although pigeons are not a primary source of human infection, close or indirect contact with them, particularly through their droppings, can contribute to the transmission of resistant bacteria. This highlights the importance of including urban wildlife in AMR surveillance programs, as part of the One Health approach that connects human, animal, and environmental health. Pigeon droppings can cause several human infections. Histoplasmosis and Cryptococcus’s are fungal infections caused by inhaling spores from dried droppings, primarily affecting the lungs and posing a risk to immunocompromised individuals. Psittacosis, a bacterial infection caused by Chlamydia psittaci, can lead to flu-like symptoms and pneumonia after inhaling dust from droppings. Salmonellosis and E. coli infections may occur through contact with contaminated surfaces or ingestion, leading to gastrointestinal illness. Less commonly, Campylobacteriosis can also result from exposure to contaminated areas. Fungi commonly isolated from pigeon droppings include Cryptococcus neoformans, Aspergillus species, Candida species, Cladosporium, and Penicillium. These fungi thrive in the nutrient-rich droppings and can pose health risks, especially Cryptococcus neoformans, which is a known pathogen causing cryptococcosis in immunocompromised individuals. The pigeon population in Hyderabad is high due to its warm climate, abundant food sources from human leftovers and temple offerings, plenty of nesting spots in old buildings, and general human tolerance and feeding habits. The aim of this study is to isolate, identify, and characterize the bacterial and fungal species present in pigeon droppings, and to evaluate the antibiotic susceptibility patterns (antibiogram) of the bacterial isolates. This investigation seeks to assess the public health risks associated with microbial contaminants found in pigeon excreta and to provide scientific data for developing effective control and preventive measures against possible zoonotic infections and antimicrobial resistance dissemination.
MATERIALS AND METHODS
Study design and study period
The present study is a cross-sectional, descriptive, laboratory-based study conducted for a period of eight months from Jan 2025 to Aug 2025 in a tertiary care teaching hospital.
Sampling
In Chevella, Ranga Reddy District, Telangana, 110 pigeon feces samples were taken using sterile cotton swabs from various locations, including temples, historic places, and public parks. After being properly sealed and labeled with the date and location of collection, pigeon droppings were gathered in sterile, clean plastic bags (plastic packets with disposable hand gloves). Nearly 100 g of sample was taken from each site. After being finely powdered by a grinding machine, the pigeon droppings were gathered in sterile, airtight containers and sent straight to the microbiology lab for examination.
Preparation of pigeon dropping suspension
In 9 ml of saline, 1g of the dry dropping powder was completely vortexed. After being moved into the next screw-capped tubes, the aliquot (1.0 ml) was serially diluted in tenth to tenth-of-a-dilution increments. A sterile spreader was used to spread a 0.1 ml aliquot from each dilution onto newly made nutritional agar plates after it had been transferred aseptically. For twenty-four hours, the inoculation plates were kept in a bacteriological incubator at 37 °C. The number of colonies that formed on the plate was measured and reported as cfu/ml. The streak plate method was used to create pure cultures of bacteria. Based on the macro-morphological colony characteristics, discrete bacteria colonies that developed were subcultured on nutrient agar and incubated at 37 °C for 24 h.
Isolation and identification of microorganisms
The bacterial isolates were identified using morphological characterization, Gram staining, cultural and biochemical characteristics by using standard microbiological procedures. MacConkey agar, Nutrient agar and Blood agar plates were used for the bacteria isolation from pigeon droppings and Sabouraud Dextrose Agar media plates for fungal isolation.
The identification of fungi was based on macro and microscopic characteristics. The surface and the reverse of the colonies were observed, as well as diameter, colony morphology, texture and pigmentation were identified. For isolation and identification, of the bacterial and fungal culture on the surface of the agar plate are incubated and inoculated in petri plates and kept at 37 °C for bacteria and 28 °C for fungi. Bacterial morphology characteristics, such as colony diameter, growth, colour, form, were observed on culture medium. Morphological features such as shape and arrangement were observed after Gram’s staining according to standard protocol. The fungal morphology was studied macroscopically by observing the colony features (colour, shape, size and hyphae) and microscopically by a compound microscope using a lactophenol cotton blue stained slide mounted with a small portion of the mycelium. For the identification of bacterial isolates routine standard biochemical tests such as Indole test, Triple sugar test, Citrate utilization test, Urease test, Nitrite reduction test, Mannitol motility, carbohydrate fermentation, catalase test, coagulase test etc. were performed. Isolates were inoculated in these medium and incubated at 37 °C for 24 h.
Antimicrobial susceptibility test
All bacteria isolates were tested for antimicrobial susceptibility against routinely used antibiotics, meropenem (10 µg), ceftazidime (30 µg), cefuroxime (30 µg), cefotaxime (30 µg), ceftriaxone (30 µg), trimethoprim/sulfamethoxazole (1.25/23.75 µg), amoxicillin+clavulanic acid (25/10 µg), ciprofloxacin (5 µg), Amikacin (10 µg), gentamicin (10 µg) etc., using disk diffusion technique developed by Bauer et al. To determine whether the isolates were resistant or sensitive to the tested antimicrobial agents, the zone of growth inhibition was measured and compared to standards given by the Clinical and Laboratory Standards Institute. Resistance to three or more antimicrobial groups is considered multidrug resistance or MDR.
RESULTS
Table 1 showed the bacterial count for serial dilutions 10-1 to 10-9of pigeon droppings. The highest bacterial count was reported from 10-4to 10-7serial dilutions. Out of 110 pigeon samples, 65 samples showed bacterial growth and 24 samples showed fungal growth. Among the 65 samples showed bacterial growth, the highest number of samples 25 (38.4%) showed Coagulase-negative Staphylococci followed by 18 samples showed Staphylococci 27.6% (table 2). Out 24 fungal isolates screened from the pigeon dropping samples, the major isolates were Aspergillus niger in 07 samples 29.16% followed by Aspergillus fumigatus 20.83% (table 3). Table 4 depicted the antimicrobial susceptibility findings of bacterial isolates. Most of the E. coli isolates were highly susceptible to meropenem 72.7% and showed resistance to most of the tested antibiotics like AMC, CXM, CIP81.1% followed by resistance to all third-generation cephalosporins 72.7%.50% of the Klebsiella species were sensitive to MRP, AK, GEN and COT. All the 4 strains were resistant to AMC100%, and 75% strains were resistant to second and third-generation cephalosporins and ciprofloxacin. Out of 18 isolates of Pseudomonas aeruginosa 72.2% strains were highly susceptible to meropenem followed by 55.5% to Amikacin. Most of the Pseudomonas isolates were resistant to CXM and COT 77.8% followed by 72.3% to AMC and third-generation cephalosporins. Among the 7 isolates of Acinetobacter species 71.4% isolates showed maximum susceptibility to meropenem and 57.2% isolates to gentamycin and amikacin and most of the isolates were resistant to CIP 85.8% and COT 71.4%. All the 25 CONS isolated were sensitive to linezolid and teicoplanin 100% and88% strains were sensitive to clindamycin and erythromycin, 72% to tetracycline. Most of the strains were resistant to penicillin antibiotic 80% (table 5).
Table 1: Bacterial count by serial dilution technique
| Serial dilution | Bacterial count CFU/ml |
| 10-1 | 19 cfu/ml |
| 10-2 | 23 cfu/ml |
| 10-3 | 35 cfu/ml |
| 10-4 | 41 x 105 cfu/ml |
| 10-5 | 20 x 106cfu/ml |
| 10-6 | 8 x 107cfu/ml |
| 10-7 | 3 x 108 cfu/ml |
| 10-8 | 5 cfu/ml |
| 10-9 | 3 cfu/ml |
Table 2: Bacterial isolates from pigeon samples
| Bacteria | Number of isolates from pigeon samples | Percentage % |
| E. coli | 11 | 16.9 |
| Klebsiella sps. | 04 | 6.15 |
| Pseudomonas sps. | 18 | 27.6 |
| CONS | 25 | 38.4 |
| Acinetobacter sps. | 07 | 10.76 |
| Total | 65 |
Table 3: Fungal isolates from pigeon samples
| Fungi | Number of isolates from pigeon samples | Percentages |
| Aspergillus niger | 07 | 29.16% |
| Aspergillus fumigatus | 05 | 20.83% |
| Mucor | 04 | 16.6% |
| Candida | 08 | 33.3% |
| Total | 24 |
Table 4: Antimicrobial susceptibility of bacterial isolates
| Bacterial Isolate | Number of Isolates | Antimicrobial susceptibility | AMC | CXM | CTR | CTX | GEN | AK | MRP | COT | CIP | CAZ |
| E. coli | 11 | Number of sensitive isolates | 02 (18.9%) | 02 (18.9%) | 03 (27.3%) |
03 (27.3%) |
05 (45.5%) | 05 (45.5%) | 08 (72.7%) | 03 (27.3%) |
02 (18.9%) | 03 (27.3%) |
| Number of resistance isolates | 09 (81.1%) |
09 (81.1%) |
08 (72.7%) |
08 (72.7%) |
06 (54.5%) | 06 (54.5%) | 03 (27.3%) |
08 (72.7%) |
09 (81.1%) |
08 (72.7%) |
||
| Klebsiella sps. | 04 | Number of sensitive isolates | 00 (0.00%) | 01 (25%) |
01 (25%) |
01 (25%) |
02 (50%) |
02 (50%) |
02 (50%) |
02 (50%) |
01 (25%) |
01 (25%) |
| Number of resistance isolates | 04 (100%) |
03 (75%) | 03 (75%) | 03 (75%) | 02 (50%) |
02 (50%) |
02 (50%) |
02 (50%) |
03 (75%) | 03 (75%) | ||
| Pseudomonas sps. | 18 | Number of sensitive isolates | 05 (27.7%) | 04 (22.2%) | 05 (27.7%) | 05 (27.7%) | 09 (50%) | 10 (55.5%) | 13 (72.2%) | 04 (22.2%) | 05 (27.7%) | 05 (27.7%) |
| Number of resistance isolates | 13 (72.3%) |
14 (77.8%) | 13 (72.3%) |
13 (72.3%) |
09 (50%) | 08 (44.5%) | 05 (27.8%) | 14 (77.8%) | 13 (72.3%) |
13 (72.3%) |
||
| Acinetobacter sps. | 07 | Number of sensitive isolates | 03 (42.8%) | 03 (42.8%) | 03 (42.8%) | 03 (42.8%) | 04 (57.2%) | 04 (57.2%) | 05 (71.4%) | 02 (28.6%) | 01 (14.2%) | 03 (42.8%) |
| Number of resistance isolates | 04 (57.2%) | 04 (57.2%) | 04 (57.2%) | 04 (57.2%) | 03 (42.8%) | 03 (42.8%) | 02 (28.6%) | 05 (71.4%) | 06 (85.8%) | 04 (57.2%) |
Table 5: Coagulase-negative staphylococci antibiotic susceptibility pattern
| Antibiotics | CONS (n=25) | |
| Number and percentage of sensitive strains | Number and percentage of resistance strains | |
| Gentamycin | 15 (60%) | 10 (40%) |
| Tetracycline | 18 (72%) | 07 (28%) |
| Clindamycin | 22 (88%) | 03 (12%) |
| Linezolid | 25 (100%) | 00 (0.00%) |
| Cotrimoxazole | 13 (52%) | 12 (48%) |
| Erythromycin | 22 (88%) | 03 (12%) |
| Cefoxitin | 10 (40%) | 15 (60%) |
| Penicillin | 05 (20%) | 20 (80%) |
| Ciprofloxacin | 10 (40%) | 15 (60%) |
| Teicoplanin | 25 (100%) | 00 (0.00%) |
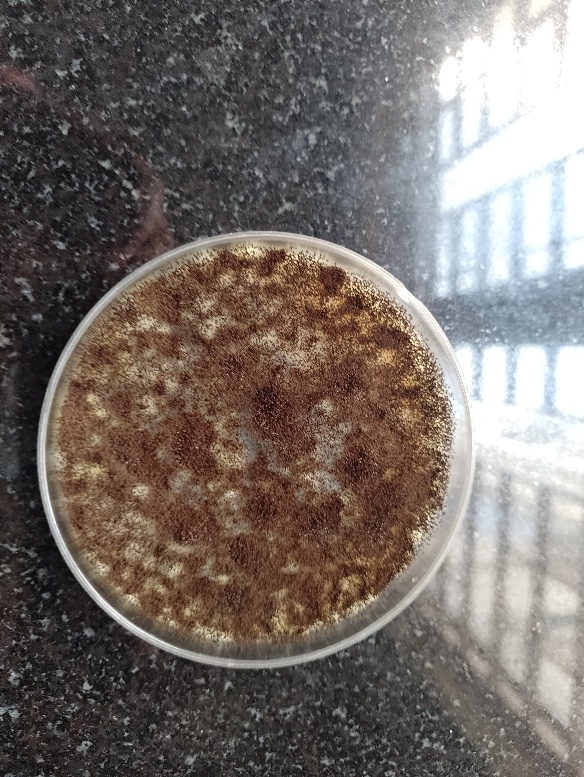
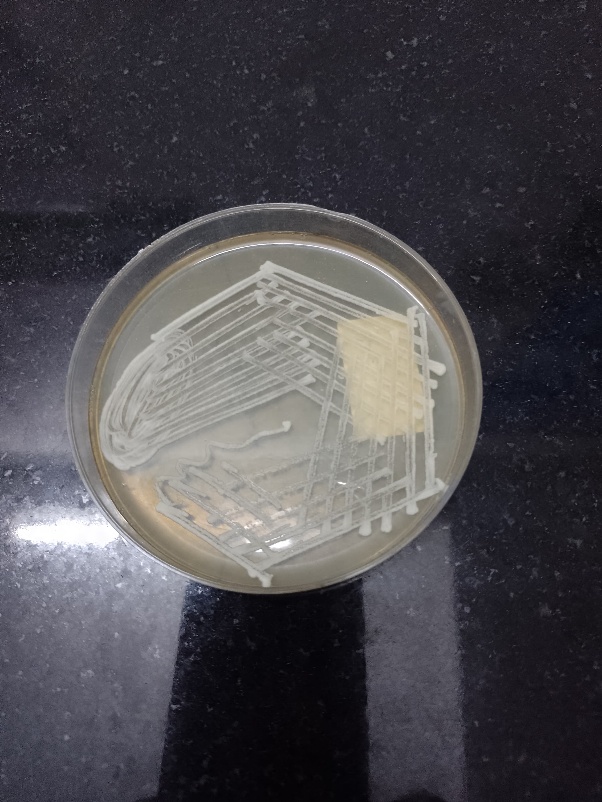
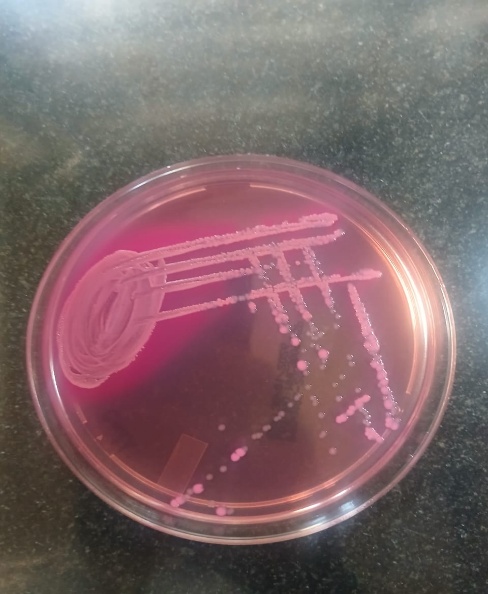
Fig. 1a: Growth of aspergillus species |
Fig. 1b: Growth of candida on subculture from plate count method |
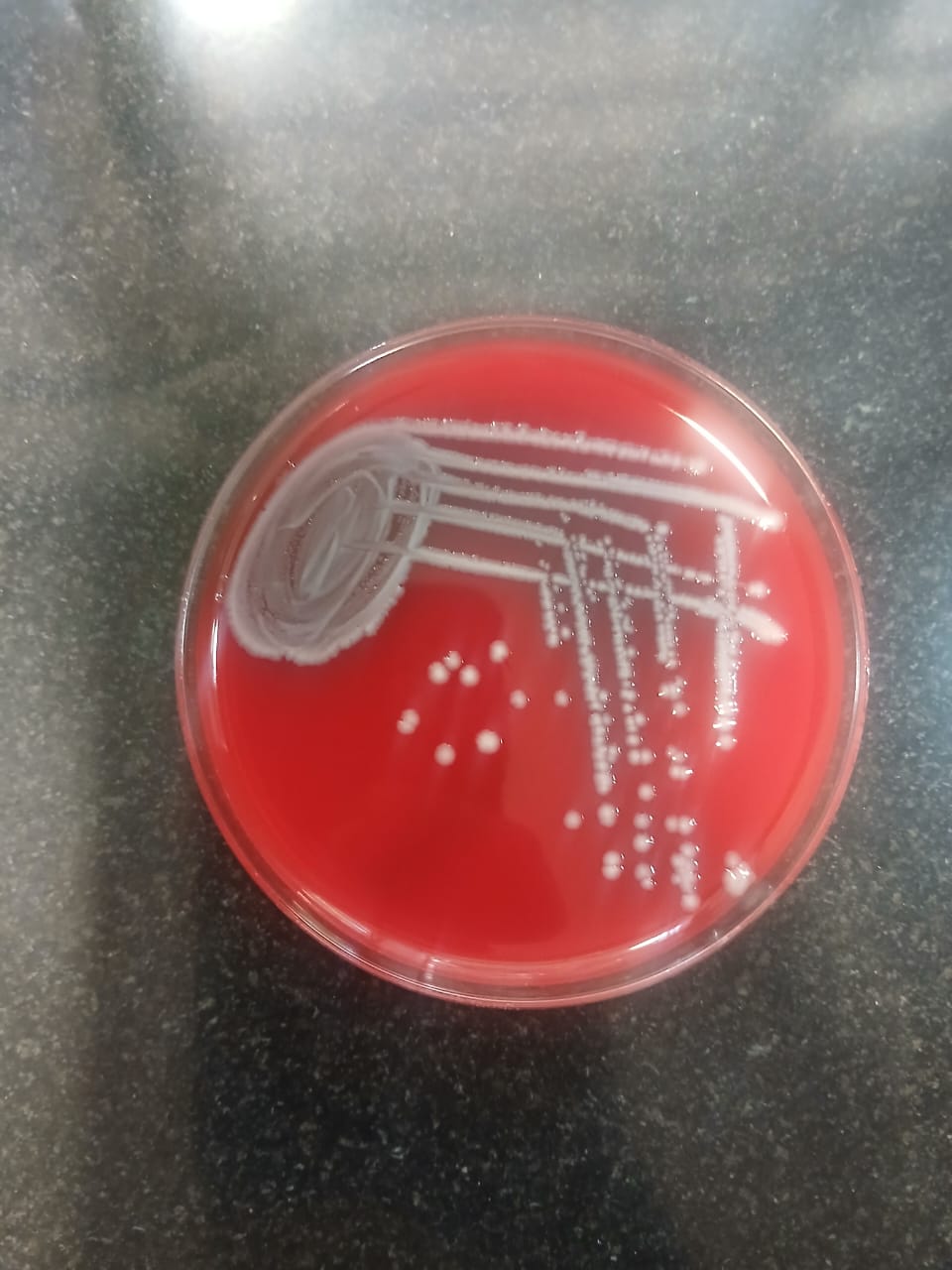
Fig. 1c: Growth of E. coli on subculture from plate count method |
Fig. 1d: Growth of klebsiella sps. on subculture from plate count method |
Fig. 1 a-d: Microbial isolates from pigeon droppings
DISCUSSION
Esakkiammal and Sheeba Rajakumari in 2024 [9] performed similar study and found that, maximum number of bacterial population was exhibited in dilution 10-4 which ranged from 76×105 cfu/m. In the present study, the maximum bacterial count was identified in10-4dilutionwhich ranged from 41 x 105 cfu/ml. Esakkiammal and Sheeba Rajakumari (2024) [9] isolated bacteria isolate of Campylobacter sp., Salmonella sp., and Micrococcus species. However, in the present study, most of the isolates were CONS38.45%, Pseudomonas aeruginosa 27.6%, E. coli16.9%, Acinetobacter species 10.76% and Klebsiella species. They also isolated different fungal isolates Fusarium sp., Aspergillus sp., and Candida sp. In the present study, we got major fungal isolates Candida species 33.3%, followed by Aspergillus niger 29.16%from pigeon droppings. Dutta et al. (2013) studied on similar work and revealed that E. coli 60.7% was the major isolate; however, in the present study we isolated 16.9% E. coli isolates [10]. The use of antibiotics is crucial for lowering mortality and illness rates associated with communicable diseases in both humans and animals, according to Tadesse et al. and Hur et al. [11-12] Intestinal disease is one of the most prevalent infectious diseases in animals that are treated with antibiotics. However, widespread use of antibiotics in both humans and animals has led to the emergence of antibiotic-resistant bacteria [11-12]. The emergence of resistant and multidrug-resistant diseases among animal isolates is a medical concern, even though few antibiotics used in animal husbandry treat human infections and many animal pathogens are zoonotic [13]. As a result, we looked at the sensitivity profile of K. pneumoniae, E. coli, and E. cloacae that were found in pigeon excrement in the present investigation. Elsheikh et al. [8] in 2024 studied about the presence of enteric bacteria from pigeon feces in Jeddah and their antimicrobial susceptibility and they found that, 50 antibiotic-resistant isolates, 28% (14/50) were Klebsiella pneumoniae, 24% (12/50) were Enterobacter cloacae, and 48% (24/50) were Escherichia coli. Ninety percent (90%) of the isolates showed resistance to cefuroxime, 56% to gentamicin, 52% to amoxicillin/clavulanic acid, and 100% to meropenem. The results were quite differed from the present study. In the present study, we obtained highest number of CONS 38.45% and Pseudomonas aeruginosa 27.6% in the pigeon droppings. All the 25 CONS isolated were sensitive to linezolid and teicoplanin 100% and88% strains were sensitive to clindamycin and erythromycin, 72% to tetracycline. Most of the strains were resistant to penicillin antibiotic 80%. Out of 18 isolates of Pseudomonas aeruginosa 72.2% strains were highly susceptible to meropenem, followed by 55.5% to Amikacin. Most of the Pseudomonas isolates were resistant to CXM and COT 77.8% followed by 72.3% to AMC and third-generation cephalosporins. It was concluded from the study that, pigeon droppings pose a significant human health risk and act as a potential source for MDR isolates. The accumulation of pigeon droppings constitutes a public health danger and causes diseases.
CONCLUSION
The bacteriological and mycological examination of pigeon droppings revealed that these excreta serve as potential reservoirs of various pathogenic microorganisms, including both bacteria and fungi of public health significance. Common bacterial isolates may include Escherichia coli, Klebsiella spp., Pseudomonas aeruginosa, Staphylococcus aureus etc. while fungal isolates often include Aspergillus spp., Candida spp., Mucor. The antibiogram typically indicates that many of these bacterial isolates exhibit resistance to multiple antibiotics, underscoring the risk of environmental dissemination of antimicrobial-resistant strains. Similarly, the presence of opportunistic and pathogenic fungi highlights a risk of respiratory or systemic infections, especially in immunocompromised individuals. Overall, pigeon droppings represent a potential public health hazard, contributing to environmental contamination and possible zoonotic transmission and hence preventive measures should be followed to prevent to prevent environmental contamination and Zoonotic transmission like regular cleaning and disinfection at areas contaminated with pigeon droppings. Bird deterrents like nets and spikes can be installed to reduce pigeon roosting near human dwellings. Collected droppings should be properly disposed of to prevent environmental contamination. Public education on the health risks associated with pigeon droppings and routine monitoring for pathogenic microbes and antimicrobial resistance are also essential preventive measures.
FUNDING
Nil
AUTHORS CONTRIBUTIONS
All authors have contributed equally
CONFLICT OF INTERESTS
Declared none
REFERENCES
Tanaka C, Miyazawa T, Watarai M, Ishiguro N. Bacteriological survey of feces from feral pigeons in Japan. J Vet Med Sci. 2005;67(9):951-3. doi: 10.1292/jvms.67.951, PMID 16210811.
Ewers C, Li G, Wilking H, Kießling S, Alt K, Antao EM, Laturnus C, Diehl I, Glodde S, Homeier T, Bohnke U, Steinruck H, Philipp HC, Wieler LH. Avian pathogenic uropathogenic and newborn meningitis-causing Escherichia coli: how closely related are they? Int J Med Microbiol. 2007;297(3):163–76. doi: 10.1016/j.ijmm.2007.01.003.
Williams RJ, Heymann DL. Containment of antibiotic resistance. Science. 1998;279(5354):1153-4. doi: 10.1126/science.279.5354.1153, PMID 9508688.
Gutierrez LM, Garcia Lopez ML, Otero A, Garciafernandez MC, Moreno B. Incidence of staphylococci in ovine mastitic milk and antibiotic susceptibility of the strains. Milchwissenschaf. 1990;45(12):778-81.
Parry CM, Threlfall EJ. Antimicrobial resistance in typhoidal and nontyphoidal salmonellae. Curr Opin Infect Dis. 2008;21(5):531-8. doi: 10.1097/QCO.0b013e32830f453a, PMID 18725804.
Molbak K, Gerner Smidt P, Wegener HC. Increasing quinolone resistance in Salmonella enterica serotype Enteritidis. Emerg Infect Dis. 2002;8(5):514-5. doi: 10.3201/eid0805.010288, PMID 11996688.
Wani SA, Samanta I, Bhat MA, Nishikawa Y. Investigation of shiga toxin-producing Escherichia coli in avian species in India. Lett Appl Microbiol. 2004;39(5):389-94. doi: 10.1111/j.1472-765X.2004.01586.x, PMID 15482427.
Elsheikh ME, Alandiyjany M, El Said M, Abouelmagd F, Ikram N, Awais M. Antimicrobial susceptibility and molecular identification of antibiotic-resistant enteric bacteria isolated from pigeon feces in the city of Jeddah, Saudi Arabia. Cureus. 2024 Aug 21;16(8):e67371. doi: 10.7759/cureus.67371, PMID 39310621, PMCID PMC11413833.
Esakkiammal M, Sheeba Rajakumari DV. Isolation and identification of microorganisms from homing pigeon droppings (Columba livia domestica). Rev Electron Vet. 2024;25(2):1402-11. doi: 10.69980/redvet.v25i2.1822.
Dutta P, Borah MK, Sarmah R, Gangil R. Isolation, histopathology and antibiogram of Escherichia coli from pigeons Columba livia. Vet World. 2013;6(2):91-4. doi: 10.5455/vetworld.2013.91-94.
Tadesse DA, Zhao S, Tong E, Ayers S, Singh A, Bartholomew MJ. Antimicrobial drug resistance in Escherichia coli from humans and food animals United States, 1950-2002. Emerg Infect Dis. 2012 May;18(5):741-9. doi: 10.3201/eid1805.111153, PMID 22515968, PMCID PMC3358085.
Hur J, Jawale C, Lee JH. Antimicrobial resistance of Salmonella isolated from food animals: a review. Food Res Int. 2012;45(2):819-30. doi: 10.1016/j.foodres.2011.05.014.
Landers TF, Cohen B, Wittum TE, Larson EL. A review of antibiotic use in food animals: perspective policy and potential. Public Health Rep. 2012 Jan-Feb;127(1):4-22. doi: 10.1177/003335491212700103, PMID 22298919, PMCID PMC3234384.